8GZO
 
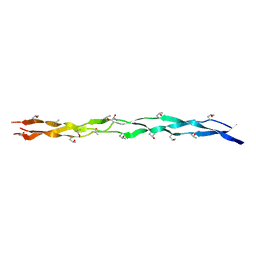 | |
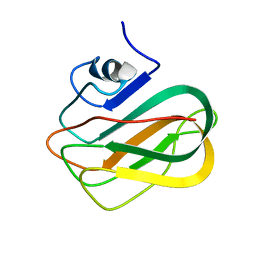
2RVA
 
 | |
1J1F
 
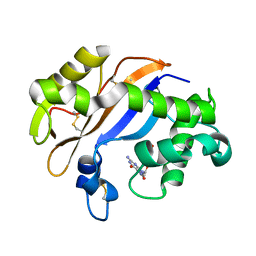 | | Crystal structure of the RNase MC1 mutant N71T in complex with 5'-GMP | | Descriptor: | GUANOSINE-5'-MONOPHOSPHATE, RIBONUCLEASE MC1 | | Authors: | Numata, T, Suzuki, A, Kakuta, Y, Kimura, K, Yao, M, Tanaka, I, Yoshida, Y, Ueda, T, Kimura, M. | | Deposit date: | 2002-12-03 | | Release date: | 2003-05-20 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal Structures of the Ribonuclease MC1 Mutants N71T and N71S in Complex with 5'-GMP: Structural Basis for Alterations in Substrate Specificity
Biochemistry, 42, 2003
|
|
6K26
 
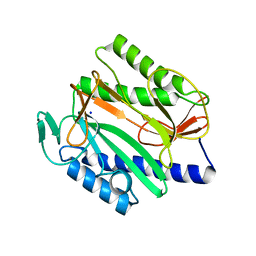 | | Crystal structure of Vibrio cholerae methionine aminopeptidase | | Descriptor: | Methionine aminopeptidase, SODIUM ION | | Authors: | Pillalamarri, V, Addlagatta, A. | | Deposit date: | 2019-05-13 | | Release date: | 2020-05-20 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Methionine aminopeptidases with short sequence inserts within the catalytic domain are differentially inhibited: Structural and biochemical studies of three proteins from Vibrio spp.
Eur.J.Med.Chem., 209, 2020
|
|
6K5S
 
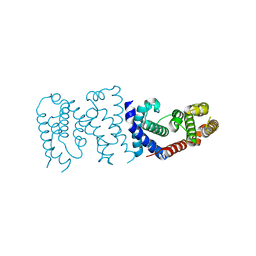 | |
2RMS
 
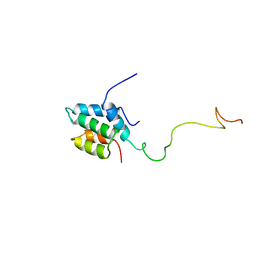 | | Solution structure of the mSin3A PAH1-SAP25 SID complex | | Descriptor: | MSin3A-binding protein, Paired amphipathic helix protein Sin3a | | Authors: | Sahu, S.C, Swanson, K.A, Kang, R.S, Huang, K, Brubaker, K, Ratcliff, K, Radhakrishnan, I. | | Deposit date: | 2007-11-14 | | Release date: | 2008-01-22 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Conserved Themes in Target Recognition by the PAH1 and PAH2 Domains of the Sin3 Transcriptional Corepressor
J.Mol.Biol., 375, 2007
|
|
2RNM
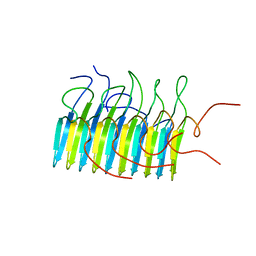 
 | | Structure of The HET-s(218-289) prion in its amyloid form obtained by solid-state NMR | | Descriptor: | Small s protein | | Authors: | Wasmer, C, Lange, A, Van Melckebeke, H, Siemer, A, Riek, R, Meier, B.H. | | Deposit date: | 2008-01-24 | | Release date: | 2008-04-01 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Amyloid fibrils of the HET-s(218-289) prion form a beta solenoid with a triangular hydrophobic core
Science, 319, 2008
|
|
6K6N
 
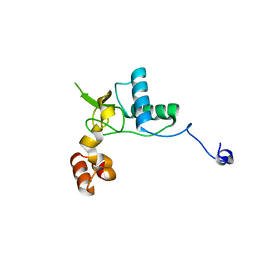 | | Crystal structure of SIVmac239 Nef protein | | Descriptor: | Protein Nef | | Authors: | Hirao, K, Andrews, S, Kuroki, K, Kusaka, H, Tadokoro, T, Kita, S, Ose, T, Rowland-Jones, S, Maenaka, K. | | Deposit date: | 2019-06-04 | | Release date: | 2020-03-25 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.0002 Å) | | Cite: | Structure of HIV-2 Nef Reveals Features Distinct from HIV-1 Involved in Immune Regulation.
Iscience, 23, 2020
|
|
2RQ8
 
 | | Solution NMR structure of titin I27 domain mutant | | Descriptor: | Titin | | Authors: | Yagawa, K, Oguro, T, Momose, T, Kawano, S, Sato, T, Endo, T. | | Deposit date: | 2009-03-05 | | Release date: | 2010-02-02 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structural basis for unfolding pathway-dependent stability of proteins: Vectorial unfolding vs. global unfolding
Protein Sci., 2010
|
|
2RG5
 
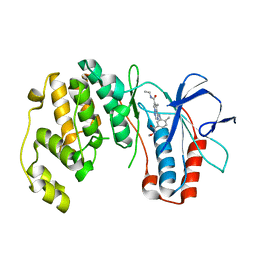 | | Phenylalanine pyrrolotriazine p38 alpha map kinase inhibitor compound 11B | | Descriptor: | Mitogen-activated protein kinase 14, N-ethyl-4-{[5-(methoxycarbamoyl)-2-methylphenyl]amino}-5-methylpyrrolo[2,1-f][1,2,4]triazine-6-carboxamide | | Authors: | Sack, J.S. | | Deposit date: | 2007-10-02 | | Release date: | 2008-01-15 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Design, Synthesis, and Anti-inflammatory Properties of Orally Active 4-(Phenylamino)-pyrrolo[2,1-f][1,2,4]triazine p38alpha Mitogen-Activated Protein Kinase Inhibitors
J.Med.Chem., 51, 2008
|
|
1IDD
 
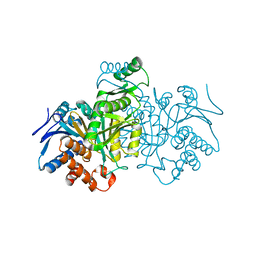 | | ISOCITRATE DEHYDROGENASE Y160F MUTANT APO ENZYME | | Descriptor: | ISOCITRATE DEHYDROGENASE | | Authors: | Lee, M.E, Dyer, D.H, Klein, O.D, Bolduc, J.M, Stoddard, B.L, Koshland Junior, D.E. | | Deposit date: | 1995-01-18 | | Release date: | 1996-03-08 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Mutagenesis and Laue structures of enzyme intermediates: isocitrate dehydrogenase.
Science, 268, 1995
|
|
2RQL
 
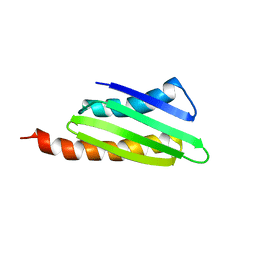 | |
2RRH
 
 | | NMR structure of vasoactive intestinal peptide in Methanol | | Descriptor: | VIP peptides | | Authors: | Umetsu, Y, Tenno, T, Goda, N, Ikegami, T, Hiroaki, H. | | Deposit date: | 2010-11-12 | | Release date: | 2011-04-06 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structural difference of vasoactive intestinal peptide in two distinct membrane-mimicking environments.
Biochim.Biophys.Acta, 1814, 2011
|
|
1IEM
 
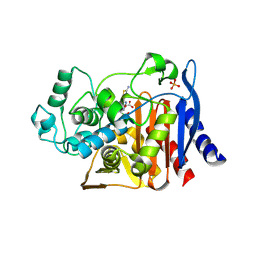 | | Crystal Structure of AmpC beta-lactamase from E. coli in Complex with a Boronic Acid Inhibitor (1, CefB4) | | Descriptor: | PHOSPHATE ION, PINACOL[[2-AMINO-ALPHA-(1-CARBOXY-1-METHYLETHOXYIMINO)-4-THIAZOLEACETYL]AMINO]METHANEBORONATE, beta-lactamase | | Authors: | Powers, R.A, Caselli, E, Focia, P.J, Prati, F, Shoichet, B.K. | | Deposit date: | 2001-04-10 | | Release date: | 2001-08-15 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structures of ceftazidime and its transition-state analogue in complex with AmpC beta-lactamase: implications for resistance mutations and inhibitor design.
Biochemistry, 40, 2001
|
|
2RT5
 
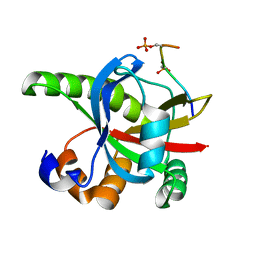 | |
1IFX
 
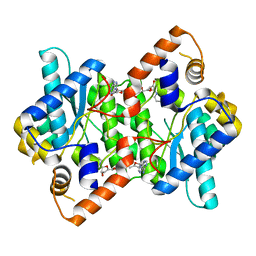 | | CRYSTAL STRUCTURE OF NH3-DEPENDENT NAD+ SYNTHETASE FROM BACILLUS SUBTILIS COMPLEXED WITH TWO MOLECULES DEAMIDO-NAD | | Descriptor: | NH(3)-DEPENDENT NAD(+) SYNTHETASE, NICOTINIC ACID ADENINE DINUCLEOTIDE | | Authors: | Devedjiev, Y, Symersky, J, Singh, R, Brouillette, W, Muccio, D, Jedrzejas, M, Brouillette, C, DeLucas, L. | | Deposit date: | 2001-04-13 | | Release date: | 2001-06-06 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Stabilization of active-site loops in NH3-dependent NAD+ synthetase from Bacillus subtilis.
Acta Crystallogr.,Sect.D, 57, 2001
|
|
2RHE
 
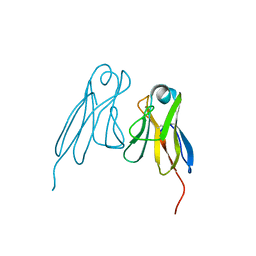 | |
2RV9
 
 | |
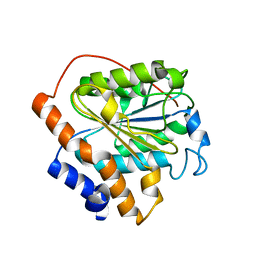
6K87
 
 | |
2SAK
 
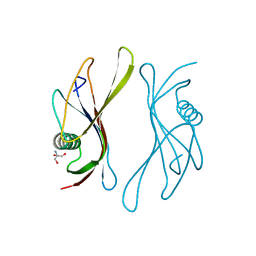 | | STAPHYLOKINASE (SAKSTAR VARIANT) | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, STAPHYLOKINASE | | Authors: | Rabijns, A, De Bondt, H.L, De Maeyer, M, Lasters, I, De Ranter, C. | | Deposit date: | 1997-02-20 | | Release date: | 1998-02-25 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Three-dimensional structure of staphylokinase, a plasminogen activator with therapeutic potential.
Nat.Struct.Biol., 4, 1997
|
|
2SCU
 
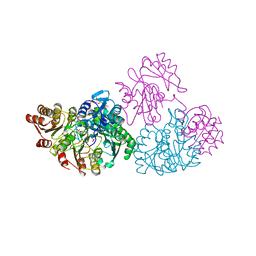 | | A detailed description of the structure of Succinyl-COA synthetase from Escherichia coli | | Descriptor: | COENZYME A, PROTEIN (SUCCINYL-COA LIGASE), SULFATE ION | | Authors: | Fraser, M.E, Wolodko, W.T, James, M.N.G, Bridger, W.A. | | Deposit date: | 1998-09-24 | | Release date: | 1999-08-02 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | A detailed structural description of Escherichia coli succinyl-CoA synthetase.
J.Mol.Biol., 285, 1999
|
|
2SH1
 
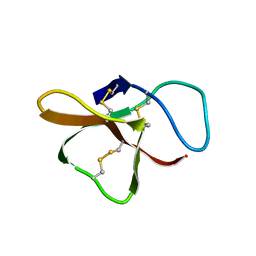 | |
2THI
 
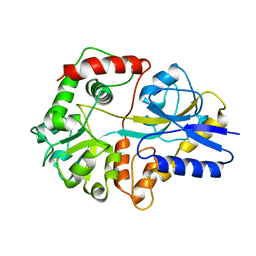 | |
6KA0
 
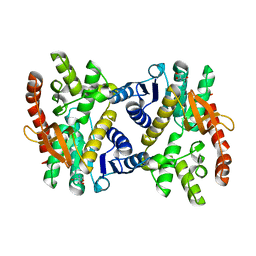 | | Silver-bound E.coli Malate dehydrogenase | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Malate dehydrogenase, SILVER ION | | Authors: | Wang, H, Wang, M, Sun, H. | | Deposit date: | 2019-06-19 | | Release date: | 2020-06-17 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.22 Å) | | Cite: | MDH is a major silver target in E. coli
To Be Published
|
|
8HLM
 
 | | Crystal structure of p53/BCL2 fusion complex (complex 2) | | Descriptor: | Apoptosis regulator Bcl-2, Cellular tumor antigen p53, ZINC ION | | Authors: | Guo, M, Wang, H, Wei, H, Chen, Y. | | Deposit date: | 2022-11-30 | | Release date: | 2023-07-26 | | Last modified: | 2024-05-29 | | Method: | X-RAY DIFFRACTION (2.522 Å) | | Cite: | Structures of p53/BCL-2 complex suggest a mechanism for p53 to antagonize BCL-2 activity.
Nat Commun, 14, 2023
|
|
