2FY7
 
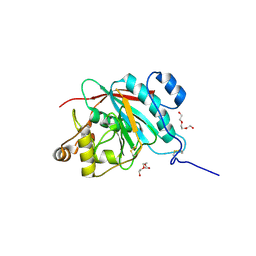 | | Crystal structure of the catalytic domain of the human beta1,4-galactosyltransferase mutant M339H in apo form | | Descriptor: | Beta-1,4-galactosyltransferase 1, TRIETHYLENE GLYCOL | | Authors: | Ramakrishnan, B, Ramasamy, V, Qasba, P.K. | | Deposit date: | 2006-02-07 | | Release date: | 2006-03-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural Snapshots of beta-1,4-Galactosyltransferase-I Along the Kinetic Pathway.
J.Mol.Biol., 357, 2006
|
|
2FY8
 
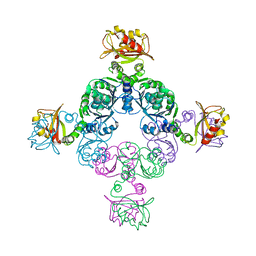 | |
2FY9
 
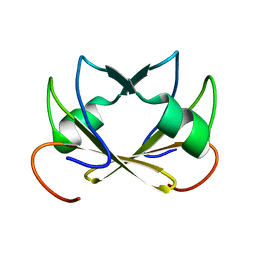 | |
2FYA
 
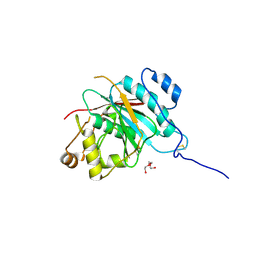 | | Crystal structure of the catalytic domain of the human beta1, 4-galactosyltransferase mutant M339H complex with manganese | | Descriptor: | Beta-1,4-galactosyltransferase 1, MANGANESE (II) ION, TRIETHYLENE GLYCOL | | Authors: | Ramakrishnan, B, Ramasamy, V, Qasba, P.K. | | Deposit date: | 2006-02-07 | | Release date: | 2006-03-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural Snapshots of beta-1,4-Galactosyltransferase-I Along the Kinetic Pathway.
J.Mol.Biol., 357, 2006
|
|
2FYB
 | | Crystal structure of the catalytic domain of the human beta1,4-galactosyltransferase mutant M339H in complex with Mn and UDP-galactose in open conformation | | Descriptor: | Beta-1,4-galactosyltransferase 1, MANGANESE (II) ION, URIDINE-5'-DIPHOSPHATE | | Authors: | Ramakrishnan, B, Ramasamy, V, Qasba, P.K. | | Deposit date: | 2006-02-07 | | Release date: | 2006-03-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural Snapshots of beta-1,4-Galactosyltransferase-I Along the Kinetic Pathway.
J.Mol.Biol., 357, 2006
|
|
2FYC
 
 | | Crystal structure of the catalytic domain of bovine beta1,4-galactosyltransferase-I in complex with alpha-lactalbumin, Ca and UDP-galactose | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Alpha-lactalbumin, CALCIUM ION, ... | | Authors: | Ramakrishnan, B, Ramasamy, V, Qasba, P.K. | | Deposit date: | 2006-02-07 | | Release date: | 2006-03-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Snapshots of beta-1,4-Galactosyltransferase-I Along the Kinetic Pathway.
J.Mol.Biol., 357, 2006
|
|
2FYD
 
 | | catalytic domain of bovine beta 1, 4-galactosyltransferase in complex with alpha-lactalbumin, glucose, Mn, and UDP-N-acetylgalactosamine | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-acetamido-2-deoxy-beta-D-galactopyranose, Alpha-lactalbumin, ... | | Authors: | Ramakrishnan, B, Ramasamy, V, Qasba, P.K. | | Deposit date: | 2006-02-07 | | Release date: | 2006-03-14 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural Snapshots of beta-1,4-Galactosyltransferase-I Along the Kinetic Pathway.
J.Mol.Biol., 357, 2006
|
|
2FYE
 
 | | Mutant Human Cathepsin S with irreversible inhibitor CRA-14013 | | Descriptor: | N-[(1R)-1-[({[(5S)-3,5-DIMETHYL-2,5-DIHYDROISOXAZOL-4-YL]METHYL}SULFONYL)METHYL]-2-OXO-2-({(1S)-3-PHENYL-1-[2-(PHENYLSULFONYL)ETHYL]PROPYL}AMINO)ETHYL]MORPHOLINE-4-CARBOXAMIDE, cathepsin S preproprotein | | Authors: | Somoza, J.R. | | Deposit date: | 2006-02-07 | | Release date: | 2006-08-08 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Mutant Human Cathepsin S with irreversible inhibitor CRA-14013
To be Published
|
|
2FYF
 
 | | Structure of a putative phosphoserine aminotransferase from Mycobacterium Tuberculosis | | Descriptor: | GLYCEROL, PYRIDOXAL-5'-PHOSPHATE, SULFATE ION, ... | | Authors: | Coulibaly, F, Lassalle, E, Baker, E.N, Mycobacterium Tuberculosis Structural Proteomics Project (XMTB) | | Deposit date: | 2006-02-07 | | Release date: | 2007-01-16 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structure of phosphoserine aminotransferase from Mycobacterium tuberculosis.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
2FYG
 
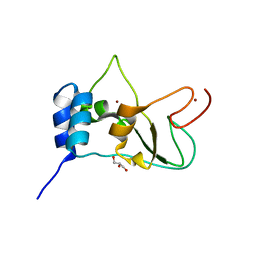 | | Crystal structure of NSP10 from Sars coronavirus | | Descriptor: | GLYCEROL, Replicase polyprotein 1ab, ZINC ION | | Authors: | Joseph, J.S, Saikatendu, K.S, Subramanian, V, Neuman, B.W, Brooun, A, Griffith, M, Moy, K, Yadav, M.K, Velasquez, J, Buchmeier, M.J, Stevens, R.C, Kuhn, P. | | Deposit date: | 2006-02-07 | | Release date: | 2006-08-08 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of nonstructural protein 10 from the severe acute respiratory syndrome coronavirus reveals a novel fold with two zinc-binding motifs.
J.Virol., 80, 2006
|
|
2FYH
 
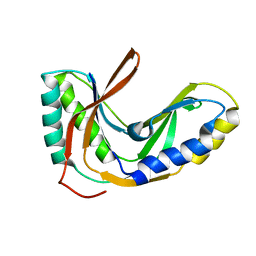 | | Solution structure of the 2'-5' RNA ligase-like protein from Pyrococcus furiosus | | Descriptor: | putative integral membrane transport protein | | Authors: | Okada, K, Matsuda, T, Sakamoto, T, Muto, Y, Yokoyama, S, Kanai, A, Kawai, G, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2006-02-08 | | Release date: | 2007-02-20 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Characterization of a heat-stable enzyme possessing GTP-dependent RNA ligase activity from a hyperthermophilic archaeon, Pyrococcus furiosus
Rna, 15, 2009
|
|
2FYI
 
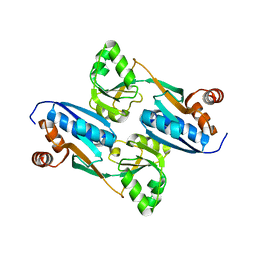 | | Crystal Structure of the Cofactor-Binding Domain of the Cbl Transcriptional Regulator | | Descriptor: | HTH-type transcriptional regulator cbl | | Authors: | Stec, E, Neumann, P, Wilkinson, A.J, Brzozowski, A.M, Bujacz, G.D. | | Deposit date: | 2006-02-08 | | Release date: | 2006-02-21 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural Basis of the Sulphate Starvation Response in E. coli: Crystal Structure and Mutational Analysis of the Cofactor-binding Domain of the Cbl Transcriptional Regulator.
J.Mol.Biol., 364, 2006
|
|
2FYJ
 
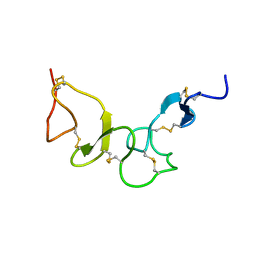 | | NMR Solution structure of calcium-loaded LRP double module | | Descriptor: | Low-density lipoprotein receptor-related protein 1 | | Authors: | Jensen, G.A, Andersen, O.M, Bonvin, A.M, Bjerrum-Bohr, I, Etzerodt, M, O'shea, C, Poulsen, F.M, Kragelund, B.B. | | Deposit date: | 2006-02-08 | | Release date: | 2006-10-10 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Binding Site Structure of One LRP-RAP Complex:Implications for a Common Ligand-Receptor Binding Motif.
J.Mol.Biol., 362, 2006
|
|
2FYK
 
 | |
2FYL
 
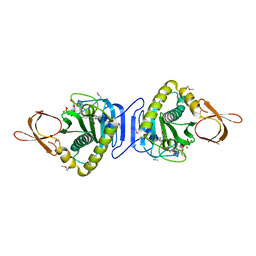
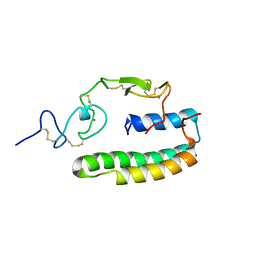 | | Haddock model of the complex between double module of LRP, CR56, and first domain of receptor associated protein, RAP-d1. | | Descriptor: | Alpha-2-macroglobulin receptor-associated protein, CALCIUM ION, Low-density lipoprotein receptor-related protein 1 | | Authors: | Jensen, G.A, Andersen, O.M, Bonvin, A.M, Bjerrum-Bohr, I, Etzerodt, M, O'shea, C, Poulsen, F.M, Kragelund, B.B. | | Deposit date: | 2006-02-08 | | Release date: | 2006-10-10 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Binding Site Structure of One LRP-RAP Complex:Implications for a Common Ligand-Receptor Binding Motif.
J.Mol.Biol., 362, 2006
|
|
2FYM
 
 | |
2FYN
 
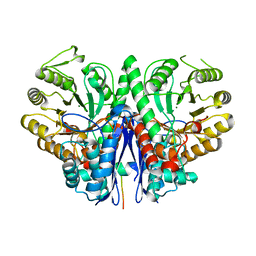
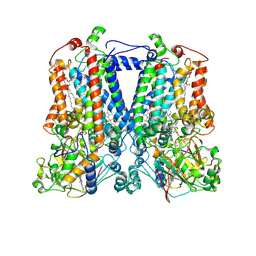 | | Crystal Structure Analysis of the double mutant Rhodobacter Sphaeroides bc1 complex | | Descriptor: | (1R)-2-{[(R)-(2-AMINOETHOXY)(HYDROXY)PHOSPHORYL]OXY}-1-[(DODECANOYLOXY)METHYL]ETHYL (9Z)-OCTADEC-9-ENOATE, Cytochrome b, Cytochrome c1, ... | | Authors: | Esser, L, Xia, D. | | Deposit date: | 2006-02-08 | | Release date: | 2006-08-29 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Surface-modulated motion switch: Capture and release of iron-sulfur protein in the cytochrome bc1 complex.
Proc.Natl.Acad.Sci.Usa, 103, 2006
|
|
2FYO
 
 | | Crystal structure of rat carnitine palmitoyltransferase 2 in space group P43212 | | Descriptor: | Carnitine O-palmitoyltransferase II, mitochondrial | | Authors: | Rufer, A.C, Thoma, R, Benz, J, Stihle, M, Gsell, B, De Roo, E, Banner, D.W, Mueller, F, Chomienne, O, Hennig, M. | | Deposit date: | 2006-02-08 | | Release date: | 2007-02-08 | | Last modified: | 2017-10-18 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The crystal structure of carnitine palmitoyltransferase 2 and implications for diabetes treatment
Structure, 14, 2006
|
|
2FYP
 
 | | GRP94 in complex with the novel HSP90 Inhibitor Radester amine | | Descriptor: | 2-(3-AMINO-2,5,6-TRIMETHOXYPHENYL)ETHYL 5-CHLORO-2,4-DIHYDROXYBENZOATE, Endoplasmin, PENTAETHYLENE GLYCOL, ... | | Authors: | Immormino, R.M, Gewirth, D.T. | | Deposit date: | 2006-02-08 | | Release date: | 2007-02-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Inhibittory Ligands Adopt Different Conformations When Bound to Hsp90 or GRP94: Implications for Paralog-specific Drug Design
To be Published
|
|
2FYQ
 
 | |
2FYR
 
 | |
2FYS
 
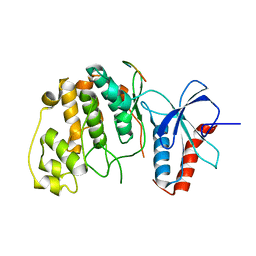 | | Crystal structure of Erk2 complex with KIM peptide derived from MKP3 | | Descriptor: | Dual specificity protein phosphatase 6, Mitogen-activated protein kinase 1 | | Authors: | Liu, S, Sun, J.P, Zhou, B, Zhang, Z.Y. | | Deposit date: | 2006-02-08 | | Release date: | 2006-04-11 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural basis of docking interactions between ERK2 and MAP kinase phosphatase 3
Proc.Natl.Acad.Sci.Usa, 103, 2006
|
|
2FYT
 
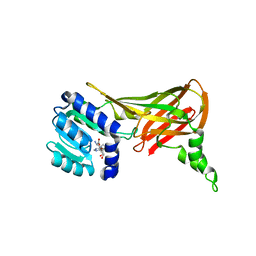 | | Human HMT1 hnRNP methyltransferase-like 3 (S. cerevisiae) protein | | Descriptor: | Protein arginine N-methyltransferase 3, S-ADENOSYL-L-HOMOCYSTEINE | | Authors: | Min, J.R, Wu, H, Zeng, H, Loppnau, P, Sundstrom, M, Arrowsmith, C.H, Edwards, A.M, Bochkarev, A, Plotnikov, A.N, Structural Genomics Consortium (SGC) | | Deposit date: | 2006-02-08 | | Release date: | 2006-03-14 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Crystal Structure of Human HMT1 hnRNP
methyltransferase-like 3 in complex with SAH.
To be Published
|
|
2FYU
 
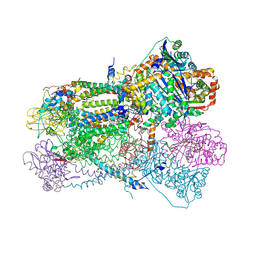 | | Crystal structure of bovine heart mitochondrial bc1 with jg144 inhibitor | | Descriptor: | (5S)-3-ANILINO-5-(2,4-DIFLUOROPHENYL)-5-METHYL-1,3-OXAZOLIDINE-2,4-DIONE, Cytochrome b, Cytochrome c1, ... | | Authors: | Xia, D, Esser, L. | | Deposit date: | 2006-02-08 | | Release date: | 2006-08-29 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Surface-modulated motion switch: Capture and release of iron-sulfur protein in the cytochrome bc1 complex.
Proc.Natl.Acad.Sci.Usa, 103, 2006
|
|
2FYV
 
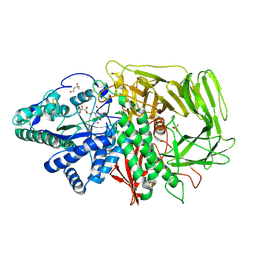 | | Golgi alpha-mannosidase II complex with an amino-salacinol carboxylate analog | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, 6-DEOXY-6-[(2R,3R,4R)-3,4-DIHYDROXY-2-(HYDROXYMETHYL)PYRROLIDIN-1-YL]-L-GULONIC ACID, ... | | Authors: | Kuntz, D.A, Hamlet, T, Rose, D.R. | | Deposit date: | 2006-02-08 | | Release date: | 2006-12-26 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Synthesis, enzymatic activity, and X-ray crystallography of an unusual class of amino acids.
Bioorg.Med.Chem., 14, 2006
|
|
