5V6M
 
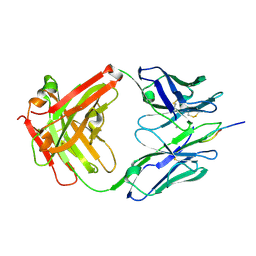 | | Crystal Structure of Rabbit Anti-HIV-1 gp120 V3 Fab 10A3 in complex with V3 peptide ConB | | Descriptor: | CALCIUM ION, Envelope glycoprotein gp120 V3 peptide of Con B sequence, Heavy chain of Fab fragment of rabbit anti-HIV1 gp120 V3 mAb 10A3, ... | | Authors: | Pan, R, Kong, X.-P. | | Deposit date: | 2017-03-17 | | Release date: | 2018-01-17 | | Last modified: | 2019-12-11 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Increased epitope complexity correlated with antibody affinity maturation and a novel binding mode revealed by structures of rabbit antibodies against the third variable loop (V3) of HIV-1 gp120.
J. Virol., 2018
|
|
5V6L
 
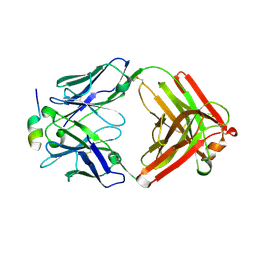 | | Crystal Structure of Rabbit Anti-HIV-1 gp120 V3 Fab 10A37 in complex with V3 peptide JR-FL | | Descriptor: | Envelope glycoprotein, v3 region, Heavy chain of Fab fragment of rabbit anti-HIV1 gp120 V3 mAb 10A37, ... | | Authors: | Pan, R, Kong, X.-P. | | Deposit date: | 2017-03-17 | | Release date: | 2018-01-17 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.549 Å) | | Cite: | Increased epitope complexity correlated with antibody affinity maturation and a novel binding mode revealed by structures of rabbit antibodies against the third variable loop (V3) of HIV-1 gp120.
J. Virol., 2018
|
|
4R2G
 
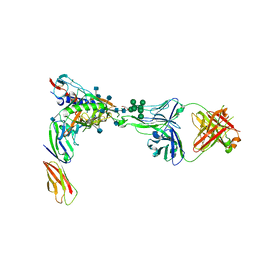 | | Crystal Structure of PGT124 Fab bound to HIV-1 JRCSF gp120 core and to CD4 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, ... | | Authors: | Garces, F, Wilson, I.A. | | Deposit date: | 2014-08-11 | | Release date: | 2014-10-08 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (3.283 Å) | | Cite: | Structural Evolution of Glycan Recognition by a Family of Potent HIV Antibodies.
Cell(Cambridge,Mass.), 159, 2014
|
|
2ESX
 
 | | The structure of the V3 region within gp120 of JR-FL HIV-1 strain (minimized average structure) | | Descriptor: | Envelope polyprotein GP160 | | Authors: | Rosen, O, Sharon, M, Samson, A.O, Quadt, S.R, Anglister, J. | | Deposit date: | 2005-10-27 | | Release date: | 2006-09-19 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Molecular switch for alternative conformations of the HIV-1 V3 region: Implications for phenotype conversion.
Proc.Natl.Acad.Sci.Usa, 13, 2006
|
|
2ESZ
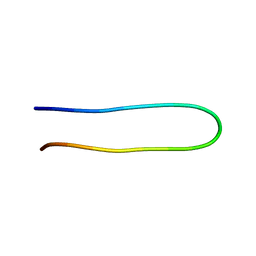 
 | | The structure of the V3 region within gp120 of JR-FL HIV-1 strain (ensemble) | | Descriptor: | Envelope polyprotein GP160 | | Authors: | Rosen, O, Sharon, M, Samson, A.O, Quadt, S.R, Anglister, J. | | Deposit date: | 2005-10-27 | | Release date: | 2006-09-19 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Molecular switch for alternative conformations of the HIV-1 V3 region: Implications for phenotype conversion.
Proc.Natl.Acad.Sci.Usa, 13, 2006
|
|
1CE4
 
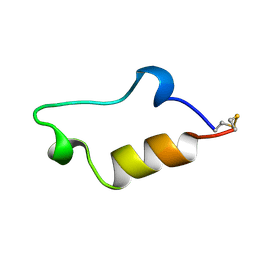 | |
