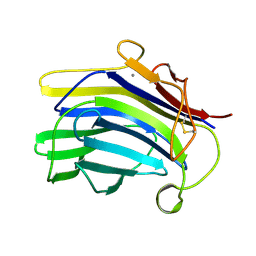
1CPN
 
 | |
1COM
 
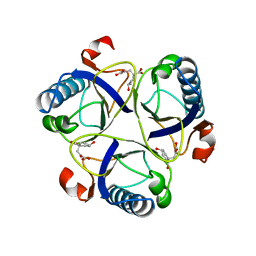 | | THE MONOFUNCTIONAL CHORISMATE MUTASE FROM BACILLUS SUBTILIS: STRUCTURE DETERMINATION OF CHORISMATE MUTASE AND ITS COMPLEXES WITH A TRANSITION STATE ANALOG AND PREPHENATE, AND IMPLICATIONS ON THE MECHANISM OF ENZYMATIC REACTION | | Descriptor: | CHORISMATE MUTASE, PREPHENIC ACID | | Authors: | Chook, Y.M, Ke, H, Lipscomb, W.N. | | Deposit date: | 1994-04-08 | | Release date: | 1994-06-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The monofunctional chorismate mutase from Bacillus subtilis. Structure determination of chorismate mutase and its complexes with a transition state analog and prephenate, and implications for the mechanism of the enzymatic reaction.
J.Mol.Biol., 240, 1994
|
|
1CPM
 
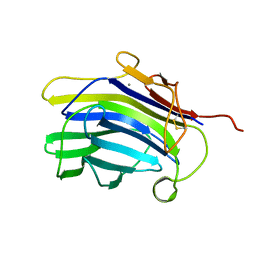 | |
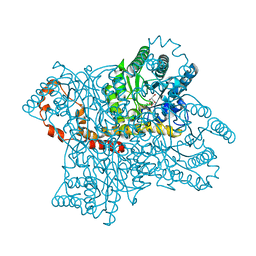
1XII
 
 | |
1XIJ
 
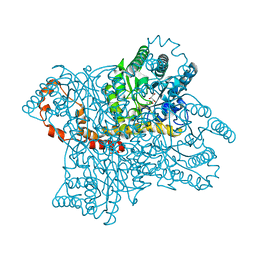 | |
1HDN
 | |
1LST
 
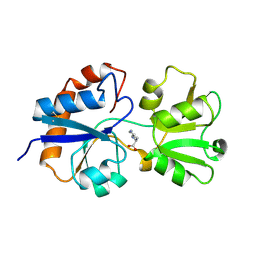 | |
1XID
 
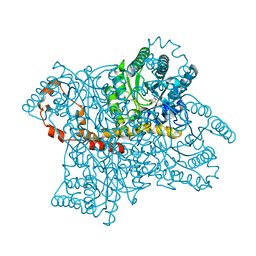 | |
1XIF
 
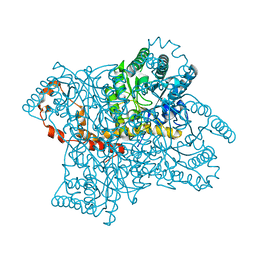 | |
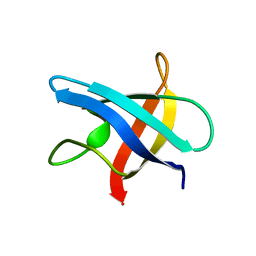
1MJC
 | |
2LAO
 
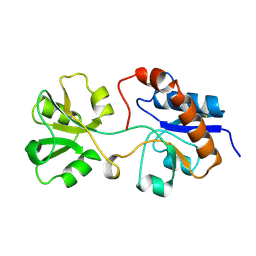 | | THREE-DIMENSIONAL STRUCTURES OF THE PERIPLASMIC LYSINE-, ARGININE-, ORNITHINE-BINDING PROTEIN WITH AND WITHOUT A LIGAND | | Descriptor: | LYSINE, ARGININE, ORNITHINE-BINDING PROTEIN | | Authors: | Kim, S.-H, Oh, B.-H, Kang, C.-H. | | Deposit date: | 1993-02-25 | | Release date: | 1994-06-22 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Three-dimensional structures of the periplasmic lysine/arginine/ornithine-binding protein with and without a ligand.
J.Biol.Chem., 268, 1993
|
|
1VTM
 
 | |
1SET
 
 | |
1SYD
 
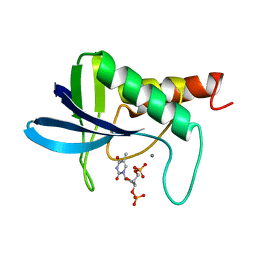 | | ENGINEERING ALTERNATIVE BETA-TURN TYPES IN STAPHYLOCOCCAL NUCLEASE | | Descriptor: | CALCIUM ION, STAPHYLOCOCCAL NUCLEASE, THYMIDINE-3',5'-DIPHOSPHATE | | Authors: | Hynes, T.R, Hodel, A, Fox, R.O. | | Deposit date: | 1994-01-07 | | Release date: | 1994-07-31 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Engineering alternative beta-turn types in staphylococcal nuclease.
Biochemistry, 33, 1994
|
|
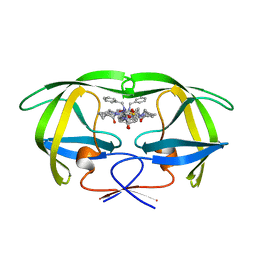
1HTG
 | |
1STB
 
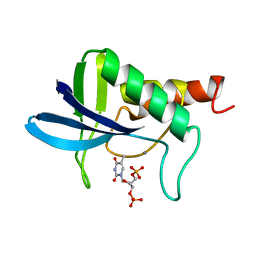 | | ACCOMMODATION OF INSERTION MUTATIONS ON THE SURFACE AND IN THE INTERIOR OF STAPHYLOCOCCAL NUCLEASE | | Descriptor: | CALCIUM ION, STAPHYLOCOCCAL NUCLEASE, THYMIDINE-3',5'-DIPHOSPHATE | | Authors: | Quirk, S, Gittis, A, Keefe, L.J, Lattman, E.E. | | Deposit date: | 1994-01-17 | | Release date: | 1994-07-31 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Accommodation of insertion mutations on the surface and in the interior of staphylococcal nuclease.
Protein Sci., 3, 1994
|
|
1SYC
 
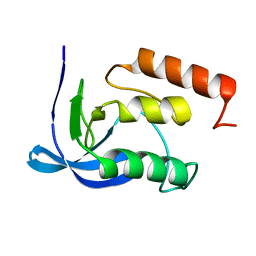 | |
137L
 
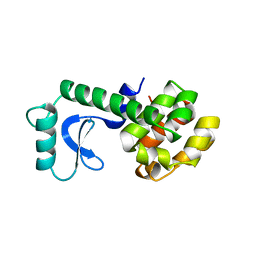 | |
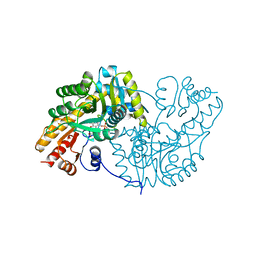
1AKB
 
 | |
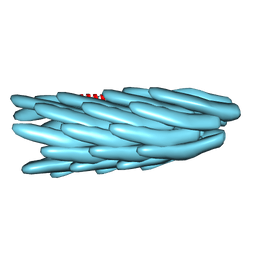
1IFM
 | |
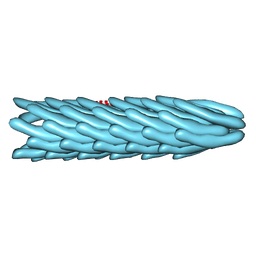
1IFK
 | |
1TND
 
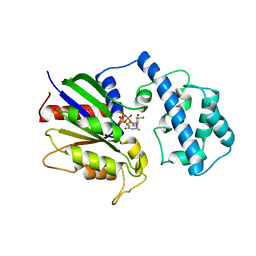 | | THE 2.2 ANGSTROMS CRYSTAL STRUCTURE OF TRANSDUCIN-ALPHA COMPLEXED WITH GTP GAMMA S | | Descriptor: | 5'-GUANOSINE-DIPHOSPHATE-MONOTHIOPHOSPHATE, CACODYLATE ION, MAGNESIUM ION, ... | | Authors: | Noel, J.P, Hamm, H.E, Sigler, P.B. | | Deposit date: | 1994-03-31 | | Release date: | 1994-07-31 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The 2.2 A crystal structure of transducin-alpha complexed with GTP gamma S.
Nature, 366, 1993
|
|
1ELS
 
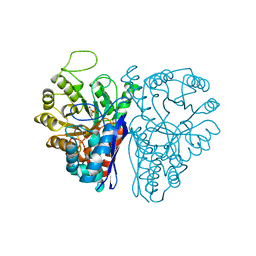 | | CATALYTIC METAL ION BINDING IN ENOLASE: THE CRYSTAL STRUCTURE OF ENOLASE-MN2+-PHOSPHONOACETOHYDROXAMATE COMPLEX AT 2.4 ANGSTROMS RESOLUTION | | Descriptor: | ENOLASE, MANGANESE (II) ION, PHOSPHONOACETOHYDROXAMIC ACID | | Authors: | Zhang, E, Hatada, M, Brewer, J.M, Lebioda, L. | | Deposit date: | 1994-04-05 | | Release date: | 1994-07-31 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Catalytic metal ion binding in enolase: the crystal structure of an enolase-Mn2+-phosphonoacetohydroxamate complex at 2.4-A resolution.
Biochemistry, 33, 1994
|
|
2DHD
 
 | |
1EPB
 
 | |
