1K9H
 
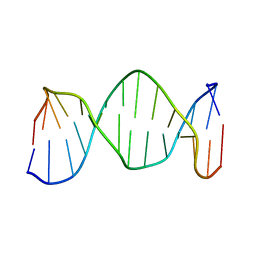 | | NMR structure of DNA TGTGAGCGCTCACA | | Descriptor: | 5'-D(*TP*GP*TP*GP*AP*GP*CP*GP*CP*TP*CP*AP*CP*A)-3' | | Authors: | Kaluarachchi, K, Gorenstein, D.G, Luxon, B.A. | | Deposit date: | 2001-10-29 | | Release date: | 2001-11-07 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | How Do Proteins Recognize DNA? Solution Structure and Local Conformational Dynamics of Lac Operators by 2D NMR
J.Biomol.Struct.Dyn., Conversation 11, 2000
|
|
2F2D
 
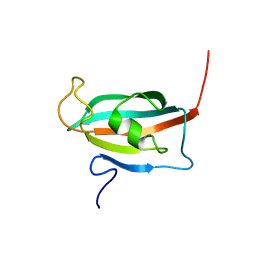 | | Solution structure of the FK506-binding domain of human FKBP38 | | Descriptor: | 38 kDa FK-506 binding protein homolog, FKBP38 | | Authors: | Maestre-Martinez, M, Edlich, F, Jarczowski, F, Weiwad, M, Fischer, G, Luecke, C. | | Deposit date: | 2005-11-16 | | Release date: | 2006-05-02 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the FK506-Binding Domain of Human FKBP38
J.BIOMOL.NMR, 34, 2006
|
|
2AKL
 
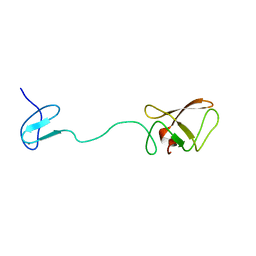 | | Solution structure for phn-A like protein PA0128 from Pseudomonas aeruginosa | | Descriptor: | ZINC ION, phnA-like protein pa0128 | | Authors: | Srisailam, S, Yee, A, Lemak, A, Lukin, J.A, Bansal, S, Prestegard, J.H, Arrowsmith, C.H, Northeast Structural Genomics Consortium (NESG), Ontario Centre for Structural Proteomics (OCSP) | | Deposit date: | 2005-08-03 | | Release date: | 2006-07-18 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Sequence Specific Resonance Assignment of a Hypothetical Protein PA0128 from Pseudomonas Aeruginosa
J.Biomol.Nmr, 36, 2006
|
|
2B86
 
 | |
2I1P
 
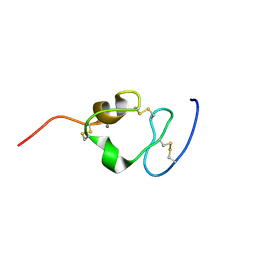 | | Solution structure of the twelfth cysteine-rich ligand-binding repeat in rat megalin | | Descriptor: | CALCIUM ION, Low-density lipoprotein receptor-related protein 2 | | Authors: | Wolf, C.A, Dancea, F, Shi, M, Bade-Noskova, V, Rueterjans, H, Kerjaschki, D, Luecke, C. | | Deposit date: | 2006-08-14 | | Release date: | 2007-02-13 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the twelfth cysteine-rich ligand-binding repeat in rat megalin.
J.Biomol.Nmr, 37, 2007
|
|
2KRR
 
 | | Solution structure of the RBD1,2 domains from human nucleolin | | Descriptor: | Nucleolin | | Authors: | Arumugam, N, Miller, C, Maliekal, J, Bates, P.J, Trent, J.O, Lane, A.N. | | Deposit date: | 2009-12-22 | | Release date: | 2010-05-05 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the RBD1,2 domains from human nucleolin.
J.Biomol.Nmr, 47, 2010
|
|
2KCU
 
 | | NMR solution structure of an uncharacterized protein from Chlorobium tepidum. Northeast Structural Genomics target CtR107 | | Descriptor: | protein CtR107 | | Authors: | Mills, J.L, Zhang, Q, Sukumaran, D.K, Wang, D, Jiang, M, Foote, E.L, Xiao, R, Nair, R, Everett, J.K, Swapna, G.V.T, Acton, T.B, Rost, B, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-12-29 | | Release date: | 2009-01-20 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of an uncharacterized protein from Chlorobium tepidum. Northeast Structural Genomics target CtR107
To be Published
|
|
2KCK
 
 | | NMR solution structure of the Northeast Structural Genomics Consortium (NESG) target MrR121A | | Descriptor: | TPR repeat | | Authors: | Barb, A.W, Lee, H.-W, Wang, X, Lee, D, Jiang, M, Ciccosanti, C, Xiao, R, Nair, R, Everett, J.K, Swapna, G.V.T, Acton, T.B, Rost, B, Montelione, G.T, Prestegard, J.H, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-12-22 | | Release date: | 2009-01-27 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of the Northeast Structural Genomics Target MrR121A
To be Published
|
|
2JWM
 
 | |
2MCJ
 
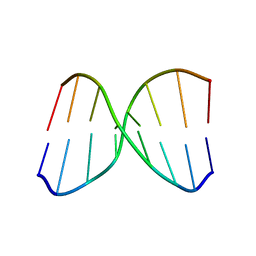 | | NMR structure of spermine modified DNA duplex | | Descriptor: | DNA_(5'-D(*CP*AP*GP*(N4S)P*CP*GP*AP*C)-3'), DNA_(5'-D(*GP*TP*CP*GP*GP*CP*TP*G)-3') | | Authors: | Brzezinska, J, Gdaniec, Z, Popenda, L, Markiewicz, W.T. | | Deposit date: | 2013-08-20 | | Release date: | 2014-01-22 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Polyaminooligonucleotide: NMR structure of duplex DNA containing a nucleoside with spermine residue, N-[4,9,13-triazatridecan-1-yl]-2'-deoxycytidine.
Biochim.Biophys.Acta, 1840, 2014
|
|
2MCI
 
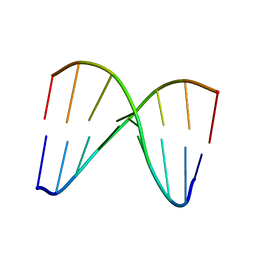 | | NMR structure of DNA duplex | | Descriptor: | DNA_(5'-D(*CP*AP*GP*CP*CP*GP*AP*C)-3'), DNA_(5'-D(*GP*TP*CP*GP*GP*CP*TP*G)-3') | | Authors: | Brzezinska, J, Gdaniec, Z, Popenda, L, Markiewicz, W.T. | | Deposit date: | 2013-08-20 | | Release date: | 2014-01-22 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Polyaminooligonucleotide: NMR structure of duplex DNA containing a nucleoside with spermine residue, N-[4,9,13-triazatridecan-1-yl]-2'-deoxycytidine.
Biochim.Biophys.Acta, 1840, 2014
|
|
2JZ5
 
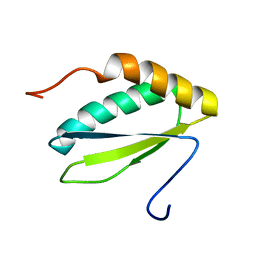 | | NMR solution structure of protein VPA0419 from Vibrio parahaemolyticus. Northeast Structural Genomics target VpR68 | | Descriptor: | Uncharacterized protein VPA0419 | | Authors: | Singarapu, K.K, Sukumaran, D.K, Eletski, A, Parish, D, Zhao, L, Jiang, M, Maglaqui, M, Xiao, R, Liu, J, Baran, M.C, Swapna, G, Acton, T, Rost, B, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2007-12-28 | | Release date: | 2008-02-19 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structures of proteins VPA0419 from Vibrio parahaemolyticus and yiiS from Shigella flexneri provide structural coverage for protein domain family PFAM 04175.
Proteins, 78, 2010
|
|
2OED
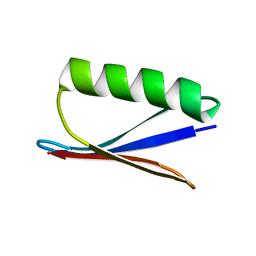 
 | | GB3 solution structure obtained by refinement of X-ray structure with dipolar couplings | | Descriptor: | Immunoglobulin G-binding protein G | | Authors: | Ulmer, T.S, Ramirez, B.E, Delaglio, F, Bax, A, Grishaev, A. | | Deposit date: | 2006-12-29 | | Release date: | 2007-01-30 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Evaluation of backbone proton positions and dynamics in a small protein by liquid crystal NMR spectroscopy
J.Am.Chem.Soc., 125, 2003
|
|
2PVE
 
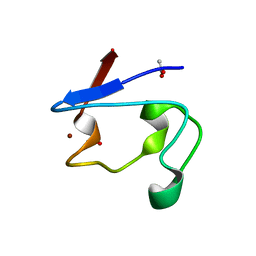 | | NMR and X-ray Analysis of Structural Additivity in Metal Binding Site-Swapped Hybrids of Rubredoxin | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, Rubredoxin, ... | | Authors: | LeMaster, D.M, Anderson, J.S, Wang, L, Guo, Y, Li, H, Hernandez, G. | | Deposit date: | 2007-05-09 | | Release date: | 2007-12-18 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (0.79 Å) | | Cite: | NMR and X-ray analysis of structural additivity in metal binding site-swapped hybrids of rubredoxin.
Bmc Struct.Biol., 7, 2007
|
|
2CVR
 
 | | NMR solution structure of sso7d mutant, K12L, 12 conformers | | Descriptor: | DNA-binding protein 7a | | Authors: | Arosio, I, Recca, T, Consonni, R, Alberti, E, Fusi, P, Zetta, L. | | Deposit date: | 2005-06-13 | | Release date: | 2006-08-29 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structural Determinants Responsible for the Thermostability of Sso7d
and its Single Point Mutants
To be Published
|
|
2LGB
 
 | | Modified A22Gly-B31Arg Human Insulin | | Descriptor: | Insulin A chain, Insulin B chain | | Authors: | Bocian, W, Kozerski, L. | | Deposit date: | 2011-07-25 | | Release date: | 2012-04-11 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Recombinant A22(G)-B31 (R)-human insulin. A22 addition introduces conformational mobility in B chain C-terminus.
J.Biomol.Nmr, 52, 2012
|
|
2PVX
 
 | | NMR and X-ray Analysis of Structural Additivity in Metal Binding Site-Swapped Hybrids of Rubredoxin | | Descriptor: | Rubredoxin, ZINC ION | | Authors: | Wang, L, LeMaster, D.M, Hernandez, G, Li, H. | | Deposit date: | 2007-05-10 | | Release date: | 2007-12-18 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.04 Å) | | Cite: | NMR and X-ray analysis of structural additivity in metal binding site-swapped hybrids of rubredoxin
Bmc Struct.Biol., 7, 2007
|
|
2KSS
 
 | | NMR structure of Myxococcus xanthus antirepressor CarS1 | | Descriptor: | Carotenogenesis protein carS | | Authors: | Jimenez, M, Gonzalez, C, Padmanabhan, S, Leon, E, Navarro-Aviles, G, Elias-Arnanz, M. | | Deposit date: | 2010-01-13 | | Release date: | 2010-05-12 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | A bacterial antirepressor with SH3 domain topology mimics operator DNA in sequestering the repressor DNA recognition helix.
Nucleic Acids Res., 38, 2010
|
|
2HSX
 
 | | NMR Structure of the nonstructural protein 1 (nsp1) from the SARS coronavirus | | Descriptor: | Leader protein; p65 homolog; NSP1 (EC 3.4.22.-) | | Authors: | Almeida, M.S, Herrmann, T, Geralt, M, Johnson, M.A, Saikatendu, K, Joseph, J, Subramanian, R.C, Neuman, B.W, Buchmeier, M.J, Stevens, R.C, Kuhn, P, Wilson, I.A, Wuthrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2006-07-24 | | Release date: | 2007-02-06 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Novel beta-barrel fold in the nuclear magnetic resonance structure of the replicase nonstructural protein 1 from the severe acute respiratory syndrome coronavirus.
J.Virol., 81, 2007
|
|
2LJ5
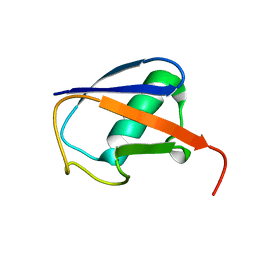 
 | |
2JOH
 
 | |
2KFM
 
 | |
2KG2
 
 | | Solution structure of a PDZ protein | | Descriptor: | Tax1-binding protein 3 | | Authors: | Durney, M.A, Birrane, G, Anklin, C, Soni, A, Ladias, J.A.A. | | Deposit date: | 2009-03-02 | | Release date: | 2010-01-19 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the human Tax-interacting protein-1.
J.Biomol.Nmr, 45, 2009
|
|
2K7O
 
 | | Ca2+-S100B, refined with RDCs | | Descriptor: | CALCIUM ION, Protein S100-B | | Authors: | Wright, N.T, Inman, K.G, Levine, J.A, Weber, D.J. | | Deposit date: | 2008-08-17 | | Release date: | 2008-11-18 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Refinement of the solution structure and dynamic properties of Ca(2+)-bound rat S100B.
J.Biomol.Nmr, 42, 2008
|
|
2L49
 
 | | The solution structure of the P2 C,the immunity repressor of the P2 bacteriophage | | Descriptor: | C protein | | Authors: | Massad, T, Papadopolos, E, Stenmark, P, Damberg, P. | | Deposit date: | 2010-10-01 | | Release date: | 2010-10-13 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | The C repressor of the P2 bacteriophage.
J.Biomol.Nmr, 64, 2016
|
|
