7A2N
 
 | |
7A2M
 
 | |
7A2R
 
 | |
2HDA
 
 | | Yes SH3 domain | | Descriptor: | Proto-oncogene tyrosine-protein kinase Yes, SULFATE ION | | Authors: | Camara-Artigas, A, Luque, I, Ruiz-Sanz, J, Mateo, P.L, Martin-Garcia, J.M. | | Deposit date: | 2006-06-20 | | Release date: | 2007-04-17 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystallographic structure of the SH3 domain of the human c-Yes tyrosine kinase: Loop flexibility and amyloid aggregation.
Febs Lett., 581, 2007
|
|
8P8K
 
 | | Acyl-ACP thioesterase from Lemna paucicostata in complex with a thiazolopyridine | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 5-[2,6-bis(fluoranyl)phenyl]-6-chloranyl-[1,3]thiazolo[4,5-b]pyridine, Acyl-ACP thioesterase | | Authors: | Freigang, J. | | Deposit date: | 2023-06-01 | | Release date: | 2023-09-20 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | A Study in Scaffold Hopping: Discovery and Optimization of Thiazolopyridines as Potent Herbicides That Inhibit Acyl-ACP Thioesterase.
J.Agric.Food Chem., 71, 2023
|
|
7ZLX
 
 | |
8QRT
 
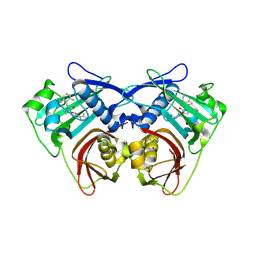 | | Acyl-ACP thioesterase from Lemna paucicostata in complex with a spirolactam | | Descriptor: | (5~{S})-7-[[2,6-bis(fluoranyl)phenyl]methyl]-3-(3-methylthiophen-2-yl)-1-oxa-2,7-diazaspiro[4.4]non-2-en-6-one, Acyl-acp thioesterase | | Authors: | Freigang, J. | | Deposit date: | 2023-10-09 | | Release date: | 2024-02-21 | | Last modified: | 2025-04-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Discovery and optimization of spirocyclic lactams that inhibit acyl-ACP thioesterase.
Pest Manag Sci, 81, 2025
|
|
8QS0
 
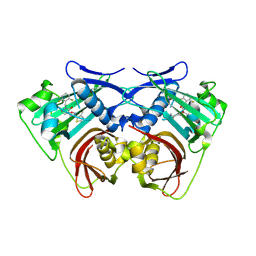 | | Acyl-ACP thioesterase from Lemna paucicostata in complex with a spirolactam | | Descriptor: | (5~{S})-9-[[2,6-bis(fluoranyl)phenyl]methyl]-3-(3-methylthiophen-2-yl)-1-oxa-2,9-diazaspiro[4.5]dec-2-en-10-one, Acyl-acp thioesterase | | Authors: | Freigang, J. | | Deposit date: | 2023-10-10 | | Release date: | 2024-02-21 | | Last modified: | 2025-04-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Discovery and optimization of spirocyclic lactams that inhibit acyl-ACP thioesterase.
Pest Manag Sci, 81, 2025
|
|
2FQX
 
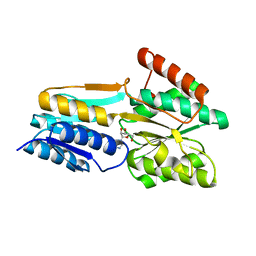 | | PnrA from Treponema pallidum complexed with guanosine | | Descriptor: | GUANOSINE, Membrane lipoprotein tmpC | | Authors: | Brautigam, C.A, Deka, R.K, Tomchick, D.R, Machius, M, Norgard, M.V. | | Deposit date: | 2006-01-18 | | Release date: | 2006-02-14 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The PnrA (Tp0319; TmpC) lipoprotein represents a new family of bacterial purine nucleoside receptor encoded within an ATP-binding cassette (ABC)-like operon in Treponema pallidum
J.Biol.Chem., 281, 2006
|
|
2FQW
 
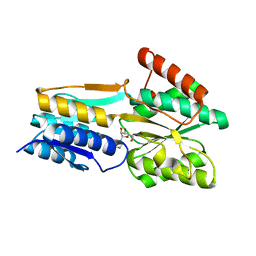 | | PnrA from Treponema pallidum as purified from E. coli (bound to inosine) | | Descriptor: | INOSINE, Membrane lipoprotein tmpC | | Authors: | Brautigam, C.A, Deka, R.K, Tomchick, D.R, Machius, M, Norgard, M.V. | | Deposit date: | 2006-01-18 | | Release date: | 2006-02-14 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.71 Å) | | Cite: | The PnrA (Tp0319; TmpC) lipoprotein represents a new family of bacterial purine nucleoside receptor encoded within an ATP-binding cassette (ABC)-like operon in Treponema pallidum
J.Biol.Chem., 281, 2006
|
|
5OAZ
 
 | |
5OAV
 
 | |
8OJW
 
 | |
8OJX
 
 | | Streptavidin S112YK121E artificial metalloenzyme for carboamination | | Descriptor: | IRIDIUM ION, Streptavidin, trichloro{(1,2,3,4,5-eta)-1,2,3,4-tetramethyl-5-[2-({5-[(3aS,4S,6aR)-2-oxohexahydro-1H-thieno[3,4-d]imidazol-4-yl]pentanoyl}amino)ethyl]cyclopentadienyl}rhodium(1+) | | Authors: | Igareta, N.V, Ward, T.R. | | Deposit date: | 2023-03-25 | | Release date: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Streptavidin WT artificial metalloenzyme for carboamination
To Be Published
|
|
5OB0
 
 | |
5OB2
 
 | |
2G93
 
 | |
5OB1
 | |
7PVR
 | |
7PVQ
 | |
7PVX
 
 | |
7PW2
 
 | |
7PVS
 
 | |
7PVV
 
 | |
7PW0
 | |
