9FEE
 
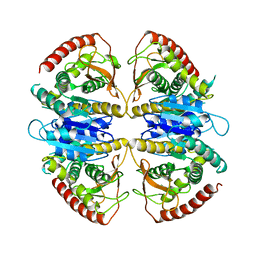 | | Cryo-EM structure of Trypanosoma cruzi glycosomal malate dehydrogenase | | Descriptor: | malate dehydrogenase | | Authors: | Lipinski, O, Sonani, R.R, Blat, A, Jemiola-Rzeminska, M, Patel, S.N, Sood, T, Dubin, G. | | Deposit date: | 2024-05-19 | | Release date: | 2024-05-29 | | Last modified: | 2025-06-11 | | Method: | ELECTRON MICROSCOPY (3.03 Å) | | Cite: | Structure of Trypanosoma peroxisomal import complex unveils conformational dynamics
Biorxiv, 2024
|
|
9FEF
 
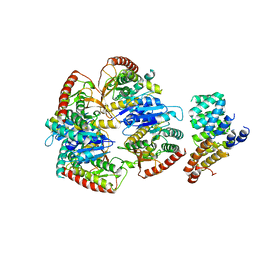 | | Cryo-EM structure of Trypanosoma cruzi (MDH)4-PEX5 complex | | Descriptor: | Peroxisome targeting signal 1 receptor, malate dehydrogenase | | Authors: | Lipinski, O, Sonani, R.R, Blat, A, Jemiola-Rzeminska, M, Patel, S.N, Sood, T, Dubin, G. | | Deposit date: | 2024-05-19 | | Release date: | 2024-05-29 | | Last modified: | 2025-06-11 | | Method: | ELECTRON MICROSCOPY (2.98 Å) | | Cite: | Structure of Trypanosoma peroxisomal import complex unveils conformational dynamics
Biorxiv, 2024
|
|
9END
 
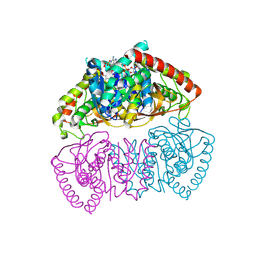 | | Crystal structure of Methanopyrus kandleri malate dehydrogenase mutant 3 | | Descriptor: | CHLORIDE ION, Malate dehydrogenase, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Coquille, S, Roche, J, Girard, E, Madern, D. | | Deposit date: | 2024-03-12 | | Release date: | 2024-07-10 | | Last modified: | 2025-01-29 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Allostery and Evolution: A Molecular Journey Through the Structural and Dynamical Landscape of an Enzyme Super Family.
Mol.Biol.Evol., 42, 2025
|
|
5NFR
 
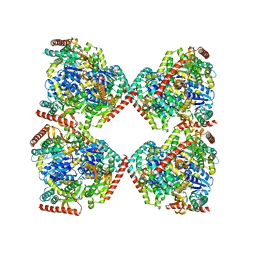 | | Crystal structure of malate dehydrogenase from Plasmodium falciparum (PfMDH) | | Descriptor: | CITRIC ACID, Malate dehydrogenase | | Authors: | Lunev, S, Romero, A.R, Batista, F.A, Wrenger, C, Groves, M.R. | | Deposit date: | 2017-03-15 | | Release date: | 2018-01-31 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Oligomeric interfaces as a tool in drug discovery: Specific interference with activity of malate dehydrogenase of Plasmodium falciparum in vitro.
PLoS ONE, 13, 2018
|
|
2A92
 
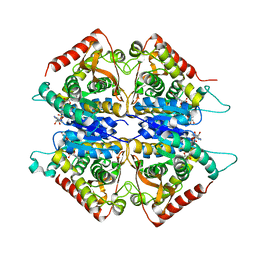 | | Crystal structure of lactate dehydrogenase from Plasmodium vivax: complex with NADH | | Descriptor: | 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, L-lactate dehydrogenase | | Authors: | Chaikuad, A, Fairweather, V, Conners, R, Joseph-Horne, T, Turgut-Balik, D, Brady, R.L. | | Deposit date: | 2005-07-11 | | Release date: | 2006-01-10 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | Structure of Lactate Dehydrogenase from Plasmodium vivax: Complexes with NADH and APADH.
Biochemistry, 44, 2005
|
|
2AA3
 | | Crystal structure of Plasmodium vivax lactate dehydrogenase complex with APADH | | Descriptor: | ACETYL PYRIDINE ADENINE DINUCLEOTIDE, REDUCED, L-lactate dehydrogenase, ... | | Authors: | Chaikuad, A, Fairweather, V, Conners, R, Joseph-Horne, T, Turgut-Balik, D, Brady, R.L. | | Deposit date: | 2005-07-13 | | Release date: | 2006-01-10 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structure of Lactate Dehydrogenase from Plasmodium vivax: Complexes with NADH and APADH.
Biochemistry, 44, 2005
|
|
8GH3
 
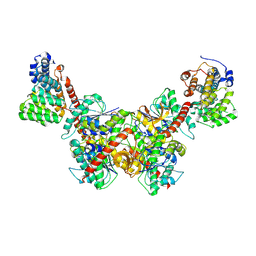 | | Structure of Trypanosoma (MDH)4-(Pex5)2, distal conformation | | Descriptor: | Peroxisome targeting signal 1 receptor, malate dehydrogenase | | Authors: | Sonani, R.R, Artur, B, Jemiola-Rzeminska, M, Lipinski, O, Patel, S.N, Sood, T, Dubin, G. | | Deposit date: | 2023-03-09 | | Release date: | 2024-03-27 | | Last modified: | 2024-10-09 | | Method: | ELECTRON MICROSCOPY (3.53 Å) | | Cite: | Noncanonical interactions and conformational dynamics in cargo-Pex5-Pex14 ternary complex for peroxisomal import
Biorxiv, 2023
|
|
8GH2
 | | Structure of Trypanosoma (MDH)4-(Pex5)2, close conformation | | Descriptor: | Peroxisome targeting signal 1 receptor, malate dehydrogenase | | Authors: | Sonani, R.R, Artur, B, Jemiola-Rzeminska, M, Lipinski, O, Patel, S.N, Sood, T, Dubin, G. | | Deposit date: | 2023-03-09 | | Release date: | 2024-03-27 | | Last modified: | 2024-10-09 | | Method: | ELECTRON MICROSCOPY (3.66 Å) | | Cite: | Noncanonical interactions and conformational dynamics in cargo-Pex5-Pex14 ternary complex for peroxisomal import
Biorxiv, 2023
|
|
8GI0
 
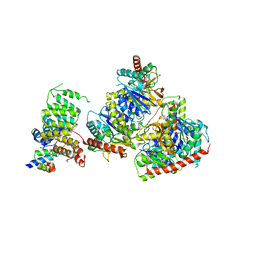 | | Structure of Trypanosoma docking complex | | Descriptor: | Peroxisomal membrane protein PEX14, Peroxisome targeting signal 1 receptor, malate dehydrogenase | | Authors: | Sonani, R.R, Artur, B, Jemiola-Rzeminska, M, Lipinski, O, Patel, S.N, Sood, T, Dubin, G. | | Deposit date: | 2023-03-13 | | Release date: | 2024-03-27 | | Last modified: | 2025-04-02 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Noncanonical interactions and conformational dynamics in cargo-Pex5-Pex14 ternary complex for peroxisomal import
Biorxiv, 2023
|
|
8GGH
 | | Structure of Trypanosoma (MDH)4-PEX5, distal conformation | | Descriptor: | Peroxisome targeting signal 1 receptor, malate dehydrogenase | | Authors: | Sonani, R.R, Artur, B, Jemiola-Rzeminska, M, Lipinski, O, Patel, S.N, Sood, T, Dubin, G. | | Deposit date: | 2023-03-08 | | Release date: | 2024-03-27 | | Last modified: | 2024-10-09 | | Method: | ELECTRON MICROSCOPY (3.29 Å) | | Cite: | Noncanonical interactions and conformational dynamics in cargo-Pex5-Pex14 ternary complex for peroxisomal import
Biorxiv, 2023
|
|
8GGD
 | | Structure of Trypanosoma (MDH)4-Pex5, close conformation | | Descriptor: | Peroxisome targeting signal 1 receptor, malate dehydrogenase | | Authors: | Sonani, R.R, Artur, B, Jemiola-Rzeminska, M, Lipinski, O, Patel, S.N, Sood, T, Dubin, G. | | Deposit date: | 2023-03-08 | | Release date: | 2024-03-27 | | Last modified: | 2024-10-09 | | Method: | ELECTRON MICROSCOPY (3.33 Å) | | Cite: | Noncanonical interactions and conformational dynamics in cargo-Pex5-Pex14 ternary complex for peroxisomal import
Biorxiv, 2023
|
|
2X8L
 
 | |
7BY9
 
 | | Malate Dehydrogenase from Geobacillus stearothermophilus (gs-MDH) complexed with Oxaloacetic Acid (OAA) and Nicotinamide Adenine Dinucleotide (NAD) | | Descriptor: | Malate dehydrogenase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, OXALOACETATE ION | | Authors: | Shimozawa, Y, Nakamura, T, Himiyama, T, Nishiya, Y. | | Deposit date: | 2020-04-22 | | Release date: | 2021-03-24 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural analysis and reaction mechanism of malate dehydrogenase from Geobacillus stearothermophilus.
J.Biochem., 170, 2021
|
|
7BY8
 | |
7BYA
 
 | | Malate Dehydrogenase from Geobacillus stearothermophilus (gs-MDH) complexed with Oxaloacetic Acid (OAA) and Adenosine 5'-Diphosphoribose (APR) | | Descriptor: | ADENOSINE-5-DIPHOSPHORIBOSE, Malate dehydrogenase, OXALOACETATE ION | | Authors: | Shimozawa, Y, Nakamura, T, Himiyama, T, Nishiya, Y. | | Deposit date: | 2020-04-22 | | Release date: | 2021-03-24 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural analysis and reaction mechanism of malate dehydrogenase from Geobacillus stearothermophilus.
J.Biochem., 170, 2021
|
|
2FN7
 
 | |
2HJR
 
 | | Crystal Structure of Cryptosporidium parvum malate dehydrogenase | | Descriptor: | ADENOSINE-5-DIPHOSPHORIBOSE, CITRIC ACID, Malate dehydrogenase | | Authors: | Wernimont, A.K, Dong, A, Lew, J, Hassani, A, Ren, H, Qiu, W, Kozieradzki, I, Weigelt, J, Sundstrom, M, Edwards, A.M, Arrowsmith, C.H, Bochkarev, A, Hui, R, Amani, M, Structural Genomics Consortium (SGC) | | Deposit date: | 2006-06-30 | | Release date: | 2006-08-01 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Genome-scale protein expression and structural biology of Plasmodium falciparum and related Apicomplexan organisms.
Mol.Biochem.Parasitol., 151, 2007
|
|
9QCG
 
 | |
5K0Z
 | | Cryo-EM structure of lactate dehydrogenase (LDH) in inhibitor-bound state | | Descriptor: | L-lactate dehydrogenase B chain | | Authors: | Merk, A, Bartesaghi, A, Banerjee, S, Falconieri, V, Rao, P, Earl, L, Milne, J, Subramaniam, S. | | Deposit date: | 2016-05-17 | | Release date: | 2016-06-08 | | Last modified: | 2024-03-06 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | Breaking Cryo-EM Resolution Barriers to Facilitate Drug Discovery.
Cell, 165, 2016
|
|
6LDH
 
 | |
1LLD
 
 | |
4CL3
 
 | | 1.70 A resolution structure of the malate dehydrogenase from Chloroflexus aurantiacus | | Descriptor: | ACETATE ION, CADMIUM ION, CHLORIDE ION, ... | | Authors: | Talon, R, Madern, D, Girard, E. | | Deposit date: | 2014-01-11 | | Release date: | 2014-02-05 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.699 Å) | | Cite: | An Experimental Point of View on Hydration/Solvation in Halophilic Proteins.
Front.Microbiol., 5, 2014
|
|
5ES3
 | | Co-crystal structure of LDH liganded with oxamate | | Descriptor: | L-lactate dehydrogenase A chain, OXAMIC ACID | | Authors: | Nowicki, M.W, Wear, M.A, McNae, I.W, Blackburn, E.A. | | Deposit date: | 2015-11-16 | | Release date: | 2016-01-13 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | A Streamlined, Automated Protocol for the Production of Milligram Quantities of Untagged Recombinant Rat Lactate Dehydrogenase A Using AKTAxpressTM.
Plos One, 10, 2015
|
|
4E0B
 
 | | 2.17 Angstrom resolution crystal structure of malate dehydrogenase from Vibrio vulnificus CMCP6 | | Descriptor: | ACETATE ION, Malate dehydrogenase | | Authors: | Halavaty, A.S, Wawrzak, Z, Onopriyenko, O, Kwon, K, Anderson, W.F, Savchenko, A, Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2012-03-02 | | Release date: | 2012-03-21 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.17 Å) | | Cite: | 2.17 Angstrom resolution crystal structure of malate dehydrogenase from Vibrio vulnificus CMCP6
To be Published
|
|
8S88
 
 | | Crystal structure of human L-lactate Dehydrogenase B protein in complex with NADH, oxamate and fluoxetine | | Descriptor: | (3S)-N-methyl-3-phenyl-3-[4-(trifluoromethyl)phenoxy]propan-1-amine, 1,4-DIHYDRONICOTINAMIDE ADENINE DINUCLEOTIDE, 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, ... | | Authors: | Van Gysel, M, Wouters, J. | | Deposit date: | 2024-03-06 | | Release date: | 2025-03-19 | | Method: | X-RAY DIFFRACTION (2.07 Å) | | Cite: | Crystal structure of human L-lactate Dehydrogenase B protein in complex with NADH, oxamate and fluoxetine
To Be Published
|
|
