3DNC
 
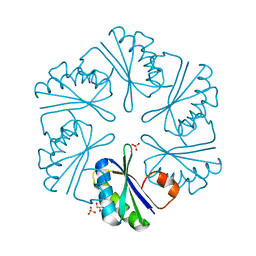 | | Carboxysome shell protein, CcmK2 C-terminal deletion mutant, with a closer spacing between hexamers | | Descriptor: | Carbon dioxide-concentrating mechanism protein ccmK homolog 2, GLYCEROL, SULFATE ION | | Authors: | Tanaka, S, Sawaya, M.R, Yeates, T.O. | | Deposit date: | 2008-07-01 | | Release date: | 2009-01-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Insights from multiple structures of the shell proteins from the beta-carboxysome.
Protein Sci., 18, 2009
|
|
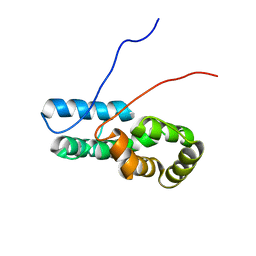
7TZ8
 
 | |
4JDX
 
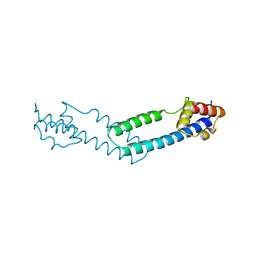 | |
3PRU
 
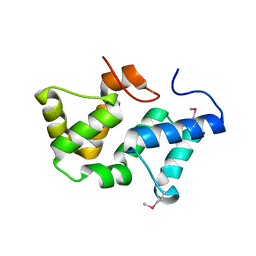 | | Crystal Structure of Phycobilisome 32.1 kDa linker polypeptide, phycocyanin-associated, rod 1 (fragment 14-158) from Synechocystis sp. PCC 6803, Northeast Structural Genomics Consortium Target SgR182A | | Descriptor: | CHLORIDE ION, Phycobilisome 32.1 kDa linker polypeptide, phycocyanin-associated, ... | | Authors: | Kuzin, A, Su, M, Patel, P, Xiao, R, Ciccosanti, C, Lee, D, Everett, J.K, Nair, R, Acton, T.B, Rost, B, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2010-11-30 | | Release date: | 2010-12-15 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.677 Å) | | Cite: | Northeast Structural Genomics Consortium Target SgR182A
To be Published
|
|
4JDQ
 
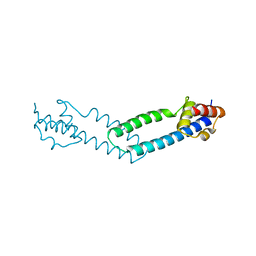 | |
2OV3
 
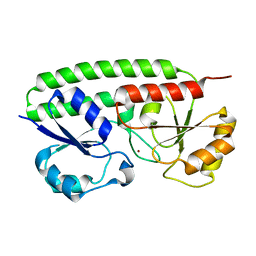 | |
5Y2W
 
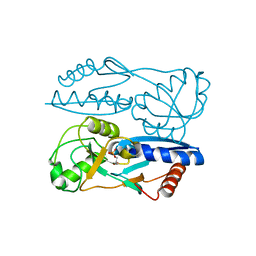 | | Structure of Synechocystis PCC6803 CcmR regulatory domain in complex with 2-PG | | Descriptor: | 2-PHOSPHOGLYCOLIC ACID, Rubisco operon transcriptional regulator | | Authors: | Jiang, Y.L, Wang, X.P, Sun, H, Cheng, W, Cao, D.D, Han, S.J, Li, W.F, Chen, Y, Zhou, C.Z. | | Deposit date: | 2017-07-27 | | Release date: | 2017-12-27 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Coordinating carbon and nitrogen metabolic signaling through the cyanobacterial global repressor NdhR.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
2VQA
 
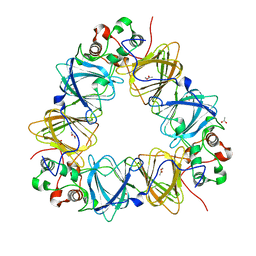 | | Protein-folding location can regulate Mn versus Cu- or Zn-binding. Crystal Structure of MncA. | | Descriptor: | ACETATE ION, MANGANESE (II) ION, SLL1358 PROTEIN | | Authors: | Tottey, S, Waldron, K.J, Firbank, S.J, Reale, B, Bessant, C, Sato, K, Gray, J, Banfield, M.J, Dennison, C, Robinson, N.J. | | Deposit date: | 2008-03-12 | | Release date: | 2008-10-28 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Protein-Folding Location Can Regulate Manganese-Binding Versus Copper- or Zinc-Binding.
Nature, 455, 2008
|
|
1IIT
 
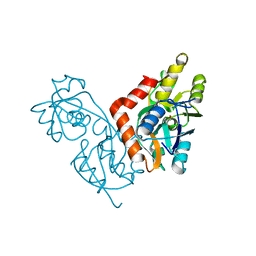 | | GLUR0 LIGAND BINDING CORE COMPLEX WITH L-SERINE | | Descriptor: | SERINE, Slr1257 protein | | Authors: | Mayer, M.L, Olson, R, Gouaux, E. | | Deposit date: | 2001-04-24 | | Release date: | 2001-09-19 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Mechanisms for ligand binding to GluR0 ion channels: crystal structures of the glutamate and serine complexes and a closed apo state.
J.Mol.Biol., 311, 2001
|
|
1IIW
 
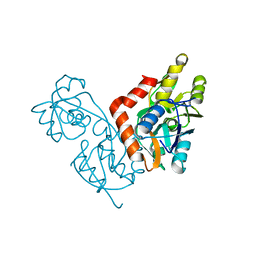 | |
6JO0
 
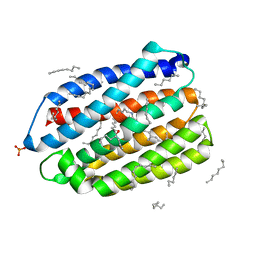 | | Crystal structure of the DTS-motif rhodopsin from Phaeocystis globosa virus 12T | | Descriptor: | (2S)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, DECANE, DODECANE, ... | | Authors: | Hosaka, T, Kimura-Someya, T, Shirouzu, M. | | Deposit date: | 2019-03-19 | | Release date: | 2019-10-02 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.651 Å) | | Cite: | A distinct lineage of giant viruses brings a rhodopsin photosystem to unicellular marine predators.
Proc.Natl.Acad.Sci.USA, 116, 2019
|
|
1II5
 
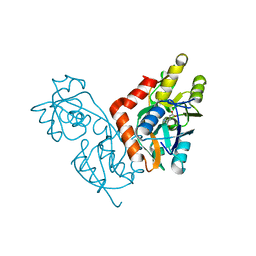 | |
2OV1
 
 | |
8QFV
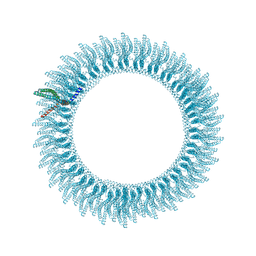 
 | | 305A Vipp1 helical tubes in the presence of EPL | | Descriptor: | Protein sll0617 | | Authors: | Junglas, B, Sachse, C. | | Deposit date: | 2023-09-05 | | Release date: | 2024-09-11 | | Last modified: | 2025-03-26 | | Method: | ELECTRON MICROSCOPY (5.1 Å) | | Cite: | Structural basis for Vipp1 membrane binding: from loose coats and carpets to ring and rod assemblies.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8QHX
 
 | | 280A Vipp1 H1-6 helical tubes in the presence of EPL | | Descriptor: | Membrane-associated protein Vipp1 | | Authors: | Junglas, B, Sachse, C. | | Deposit date: | 2023-09-11 | | Release date: | 2024-09-18 | | Last modified: | 2025-07-09 | | Method: | ELECTRON MICROSCOPY (6.9 Å) | | Cite: | Structural basis for Vipp1 membrane binding: from loose coats and carpets to ring and rod assemblies.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8QI0
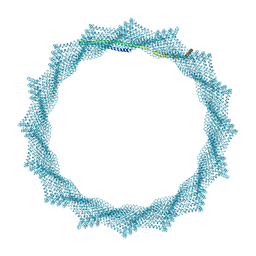 
 | | 340A Vipp1 H1-6 helical tubes | | Descriptor: | Membrane-associated protein Vipp1 | | Authors: | Junglas, B, Sachse, C. | | Deposit date: | 2023-09-11 | | Release date: | 2024-09-18 | | Last modified: | 2025-03-26 | | Method: | ELECTRON MICROSCOPY (6.7 Å) | | Cite: | Structural basis for Vipp1 membrane binding: from loose coats and carpets to ring and rod assemblies.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8QI4
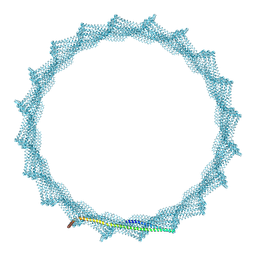 
 | | 400A Vipp1 H1-6 helical tubes | | Descriptor: | Membrane-associated protein Vipp1 | | Authors: | Junglas, B, Sachse, C. | | Deposit date: | 2023-09-11 | | Release date: | 2024-09-18 | | Last modified: | 2025-03-26 | | Method: | ELECTRON MICROSCOPY (7 Å) | | Cite: | Structural basis for Vipp1 membrane binding: from loose coats and carpets to ring and rod assemblies.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8QHY
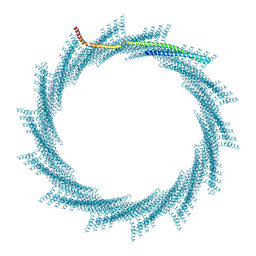 
 | | 300A Vipp1 H1-6 helical tubes in the presence of EPL | | Descriptor: | Membrane-associated protein Vipp1 | | Authors: | Junglas, B, Sachse, C. | | Deposit date: | 2023-09-11 | | Release date: | 2024-09-18 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (7 Å) | | Cite: | Structural basis for Vipp1 membrane binding: from loose coats and carpets to ring and rod assemblies.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8QI5
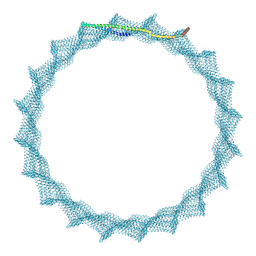 
 | | 405A Vipp1 H1-6 helical tubes | | Descriptor: | Membrane-associated protein Vipp1 | | Authors: | Junglas, B, Sachse, C. | | Deposit date: | 2023-09-11 | | Release date: | 2024-09-18 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (6.7 Å) | | Cite: | Structural basis for Vipp1 membrane binding: from loose coats and carpets to ring and rod assemblies.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8QHV
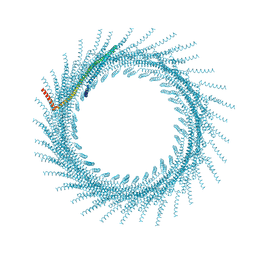 
 | | 275A Vipp1 helical tubes in the presence of EPL | | Descriptor: | Membrane-associated protein Vipp1 | | Authors: | Junglas, B, Sachse, C. | | Deposit date: | 2023-09-11 | | Release date: | 2024-09-18 | | Last modified: | 2025-07-09 | | Method: | ELECTRON MICROSCOPY (7 Å) | | Cite: | Structural basis for Vipp1 membrane binding: from loose coats and carpets to ring and rod assemblies.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8QI2
 | | 370A Vipp1 H1-6 helical tubes | | Descriptor: | Membrane-associated protein Vipp1 | | Authors: | Junglas, B, Sachse, C. | | Deposit date: | 2023-09-11 | | Release date: | 2024-09-18 | | Last modified: | 2025-07-09 | | Method: | ELECTRON MICROSCOPY (6.7 Å) | | Cite: | Structural basis for Vipp1 membrane binding: from loose coats and carpets to ring and rod assemblies.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8QI1
 | | 360A Vipp1 H1-6 helical tubes | | Descriptor: | Membrane-associated protein Vipp1 | | Authors: | Junglas, B, Sachse, C. | | Deposit date: | 2023-09-11 | | Release date: | 2024-09-18 | | Last modified: | 2025-07-09 | | Method: | ELECTRON MICROSCOPY (6.6 Å) | | Cite: | Structural basis for Vipp1 membrane binding: from loose coats and carpets to ring and rod assemblies.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8QHZ
 | | 320A Vipp1 H1-6 helical tubes | | Descriptor: | Membrane-associated protein Vipp1 | | Authors: | Junglas, B, Sachse, C. | | Deposit date: | 2023-09-11 | | Release date: | 2024-09-18 | | Last modified: | 2025-03-26 | | Method: | ELECTRON MICROSCOPY (6.3 Å) | | Cite: | Structural basis for Vipp1 membrane binding: from loose coats and carpets to ring and rod assemblies.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8QI6
 | | 420A Vipp1 H1-6 helical tubes | | Descriptor: | Membrane-associated protein Vipp1 | | Authors: | Junglas, B, Sachse, C. | | Deposit date: | 2023-09-11 | | Release date: | 2024-09-18 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (7.8 Å) | | Cite: | Structural basis for Vipp1 membrane binding: from loose coats and carpets to ring and rod assemblies.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8QHW
 
 | | 290A Vipp1 helical tubes in the presence of EPL | | Descriptor: | Membrane-associated protein Vipp1 | | Authors: | Junglas, B, Sachse, C. | | Deposit date: | 2023-09-11 | | Release date: | 2024-09-18 | | Last modified: | 2025-03-26 | | Method: | ELECTRON MICROSCOPY (7.1 Å) | | Cite: | Structural basis for Vipp1 membrane binding: from loose coats and carpets to ring and rod assemblies.
Nat.Struct.Mol.Biol., 32, 2025
|
|
