4MYP
 
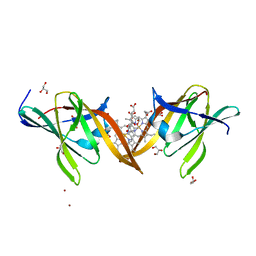 | | Structure of the central NEAT domain, N2, of the listerial Hbp2 protein complexed with heme | | Descriptor: | GLYCEROL, Iron-regulated surface determinant protein A, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Malmirchegini, G.R, Sawaya, M.R, Clubb, R.T. | | Deposit date: | 2013-09-27 | | Release date: | 2014-10-22 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Novel Mechanism of Hemin Capture by Hbp2, the Hemoglobin-binding Hemophore from Listeria monocytogenes.
J.Biol.Chem., 289, 2014
|
|
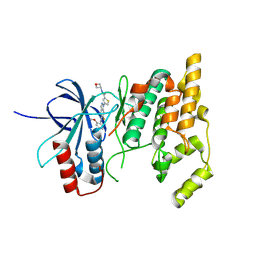
8VTF
 
 | |
4N76
 
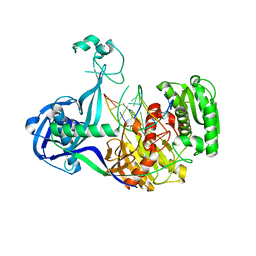 | | Structure of Thermus thermophilus Argonaute bound to guide DNA and cleaved target DNA with Mn2+ | | Descriptor: | 5'-D(P*TP*AP*CP*TP*AP*CP*CP*TP*CP*G)-3', 5'-D(P*TP*GP*AP*GP*GP*TP*AP*GP*TP*AP*GP*GP*TP*TP*GP*TP*AP*TP*AP*GP*T)-3', Argonaute, ... | | Authors: | Sheng, G, Zhao, H, Wang, J, Rao, Y, Wang, Y. | | Deposit date: | 2013-10-15 | | Release date: | 2014-01-15 | | Last modified: | 2025-02-12 | | Method: | X-RAY DIFFRACTION (2.89 Å) | | Cite: | Structure-based cleavage mechanism of Thermus thermophilus Argonaute DNA guide strand-mediated DNA target cleavage.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
8VKX
 
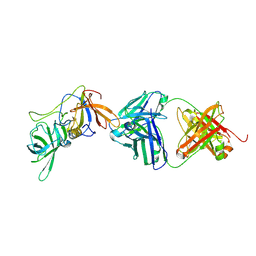 | | VX22 bound to GII.4 P domain | | Descriptor: | VP1, VX22 heavy chain, VX22 light chain | | Authors: | Olia, A.S, Park, J, Lindesmith, L.C, Kwong, P.D. | | Deposit date: | 2024-01-10 | | Release date: | 2025-02-19 | | Last modified: | 2025-09-03 | | Method: | X-RAY DIFFRACTION (3.35 Å) | | Cite: | Broadly neutralizing antibodies targeting pandemic GII.4 variants or seven GII genotypes of human norovirus.
Sci Transl Med, 17, 2025
|
|
4N8C
 
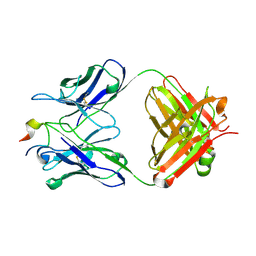 | | Three-dimensional structure of the extracellular domain of Matrix protein 2 of influenza A virus | | Descriptor: | Extracellular domain of influenza Matrix protein 2, Heavy chain of monoclonal antibody, Light chain of monoclonal antibody | | Authors: | Cho, K.J, Seok, J.H, Kim, S, Roose, K, Schepens, B, Fiers, W, Saelens, X, Kim, K.H. | | Deposit date: | 2013-10-17 | | Release date: | 2014-10-22 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure of the extracellular domain of matrix protein 2 of influenza A virus in complex with a protective monoclonal antibody
J.Virol., 89, 2015
|
|
8VXX
 
 | | Mango II bound to 365A-061 | | Descriptor: | 3-[2-({[(6P)-2-fluoro-6-(1H-pyrazol-1-yl)phenyl]methyl}amino)-2-oxoethyl]-2-[(E)-(1-methylquinolin-4(1H)-ylidene)methyl]-1,3-benzothiazol-3-ium, Chains: A,B,C, POTASSIUM ION | | Authors: | Passalacqua, L.F.M, Ferre-D'Amare, A.R, Schneekloth, J.S. | | Deposit date: | 2024-02-06 | | Release date: | 2025-02-19 | | Last modified: | 2025-10-08 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structure-informed design of an ultrabright RNA-activated fluorophore.
Nat.Chem., 17, 2025
|
|
4N8N
 
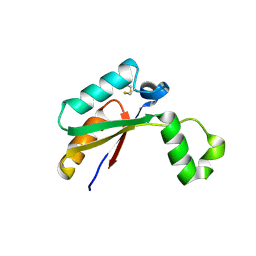 | | Crystal structure of Mycobacterial FtsX extracellular domain | | Descriptor: | Cell division protein FtsX, POTASSIUM ION | | Authors: | Mavrici, D. | | Deposit date: | 2013-10-17 | | Release date: | 2014-05-28 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.874 Å) | | Cite: | Mycobacterium tuberculosis FtsX extracellular domain activates the peptidoglycan hydrolase, RipC.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
8VK0
 
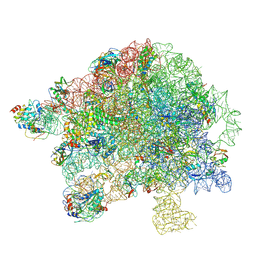 | | Structure of Mycobacterium smegmatis 50S ribosomal subunit bound to HflX:50S-HflX-A | | Descriptor: | 23S ribosomal RNA, 50S Ribosomal Protein L13, 50S Ribosomal Protein L18, ... | | Authors: | Majumdar, S, Koripella, R.K, Sharma, M.R, Manjari, S.R, Banavali, N.K, Agrawal, R.K. | | Deposit date: | 2024-01-08 | | Release date: | 2025-02-19 | | Last modified: | 2025-04-30 | | Method: | ELECTRON MICROSCOPY (3.14 Å) | | Cite: | HflX-mediated drug resistance through ribosome splitting and rRNA disordering in mycobacteria.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
5WIW
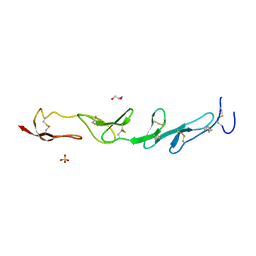 
 | |
8VR8
 
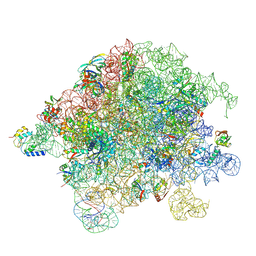 | | Structure of Mycobacterium smegmatis 50S ribosomal subunit bound to HflX and chloramphenicol:50S-HflX-B-Clm | | Descriptor: | 23S ribosomal RNA, 50S Ribosomal Protein L13, 50S Ribosomal Protein L20, ... | | Authors: | Majumdar, S, Koripella, R.K, Sharma, M.R, Manjari, S.R, Banavali, N.K, Agrawal, R.K. | | Deposit date: | 2024-01-20 | | Release date: | 2025-02-19 | | Last modified: | 2025-04-30 | | Method: | ELECTRON MICROSCOPY (3.25 Å) | | Cite: | HflX-mediated drug resistance through ribosome splitting and rRNA disordering in mycobacteria.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
6SSA
 
 | | Human Leukocyte Antigen Class I A02 Carrying LLWNGPMQV | | Descriptor: | 1,2-ETHANEDIOL, Beta-2-microglobulin, CALCIUM ION, ... | | Authors: | Rizkallah, P.J, Bovay, A. | | Deposit date: | 2019-09-06 | | Release date: | 2020-07-15 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | Identification of a superagonist variant of the immunodominant Yellow fever virus epitope NS4b214-222by combinatorial peptide library screening.
Mol.Immunol., 125, 2020
|
|
4N96
 
 | | E. coli sliding clamp in complex with 6-nitroindazole | | Descriptor: | 6-NITROINDAZOLE, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Yin, Z, Oakley, A.J. | | Deposit date: | 2013-10-19 | | Release date: | 2013-11-06 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Discovery of lead compounds targeting the bacterial sliding clamp using a fragment-based approach.
J.Med.Chem., 57, 2014
|
|
8VIO
 
 | | Structure of Mycobacterium smegmatis HflX bound to a 70S ribosome | | Descriptor: | 16S ribosomal RNA, 23S ribosomal RNA, 30S Ribosomal Protein S14, ... | | Authors: | Majumdar, S, Koripella, R.K, Sharma, M.R, Manjari, S.R, Banavali, N.K, Agrawal, R.K. | | Deposit date: | 2024-01-05 | | Release date: | 2025-02-19 | | Last modified: | 2025-04-30 | | Method: | ELECTRON MICROSCOPY (3.26 Å) | | Cite: | HflX-mediated drug resistance through ribosome splitting and rRNA disordering in mycobacteria.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
8VK7
 
 | | Structure of Mycobacterium smegmatis 50S ribosomal subunit bound to HflX:50S-HflX-B | | Descriptor: | 23S ribosomal RNA, 50S Ribosomal Protein L13, 50S Ribosomal Protein L18, ... | | Authors: | Majumdar, S, Koripella, R.K, Sharma, M.R, Manjari, S.R, Banavali, N.K, Agrawal, R.K. | | Deposit date: | 2024-01-08 | | Release date: | 2025-02-19 | | Last modified: | 2025-04-30 | | Method: | ELECTRON MICROSCOPY (3.09 Å) | | Cite: | HflX-mediated drug resistance through ribosome splitting and rRNA disordering in mycobacteria.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
8VKW
 
 | | Structure of Mycobacterium smegmatis 50S ribosomal subunit bound to delNTE-HflX | | Descriptor: | 23S ribosomal RNA, 50S Ribosomal Protein L13, 50S Ribosomal Protein L18, ... | | Authors: | Majumdar, S, Koripella, R.K, Sharma, M.R, Manjari, S.R, Banavali, N.K, Agrawal, R.K. | | Deposit date: | 2024-01-10 | | Release date: | 2025-02-19 | | Last modified: | 2025-04-30 | | Method: | ELECTRON MICROSCOPY (3.44 Å) | | Cite: | HflX-mediated drug resistance through ribosome splitting and rRNA disordering in mycobacteria.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
4N9O
 
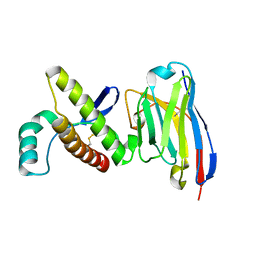 | | Probing the N-terminal beta-sheet conversion in the crystal structure of the human prion protein bound to a Nanobody | | Descriptor: | Major prion protein, Nanobody Nb484 | | Authors: | Abskharon, R.N.N, Giachin, G, Wohlkonig, A, Soror, S.H, Pardon, E, Legname, G, Steyaert, J. | | Deposit date: | 2013-10-21 | | Release date: | 2014-01-22 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Probing the N-Terminal beta-Sheet Conversion in the Crystal Structure of the Human Prion Protein Bound to a Nanobody.
J.Am.Chem.Soc., 136, 2014
|
|
8VKI
 
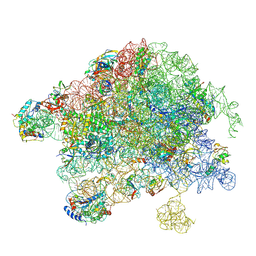 | | Structure of Mycobacterium smegmatis 50S ribosomal subunit bound to HflX:50S-HflX-C | | Descriptor: | 23S ribosomal RNA, 50S Ribosomal Protein L13, 50S Ribosomal Protein L18, ... | | Authors: | Majumdar, S, Koripella, R.K, Sharma, M.R, Manjari, S.R, Banavali, N.K, Agrawal, R.K. | | Deposit date: | 2024-01-09 | | Release date: | 2025-02-19 | | Last modified: | 2025-04-30 | | Method: | ELECTRON MICROSCOPY (2.96 Å) | | Cite: | HflX-mediated drug resistance through ribosome splitting and rRNA disordering in mycobacteria.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
4N9X
 
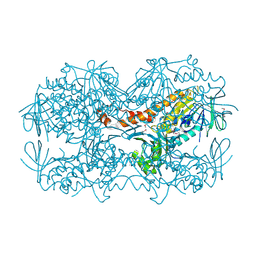 | | Crystal Structure of the OCTAPRENYL-METHYL-METHOXY-BENZQ MOLECULE from Erwina carotovora subsp. atroseptica strain SCRI 1043 / ATCC BAA-672, Northeast Structural Genomics Consortium (NESG) Target EwR161 | | Descriptor: | Putative monooxygenase | | Authors: | Kuzin, A, Chen, Y, Lew, S, Seetharaman, J, Mao, L, Xiao, R, Owens, L.A, Wang, D, Everett, J.K, Acton, T.B, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2013-10-21 | | Release date: | 2013-11-20 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.503 Å) | | Cite: | Crystal Structure of the OCTAPRENYL-METHYL-METHOXY-BENZQ MOLECULE from Erwina carotovora subsp. atroseptica strain SCRI 1043 / ATCC BAA-672, Northeast Structural Genomics Consortium (NESG) Target EwR161
To be Published
|
|
8VR4
 
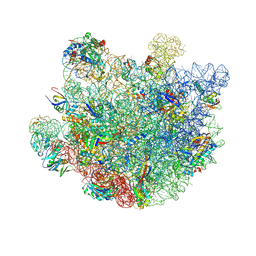 | | Structure of Mycobacterium smegmatis 50S ribosomal subunit bound to HflX and erythromycin:50S-HflX-A-Ery | | Descriptor: | 23S ribosomal RNA, 50S Ribosomal Protein L13, 50S Ribosomal Protein L18, ... | | Authors: | Majumdar, S, Koripella, R.K, Sharma, M.R, Manjari, S.R, Banavali, N.K, Agrawal, R.K. | | Deposit date: | 2024-01-20 | | Release date: | 2025-02-19 | | Last modified: | 2025-04-30 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | HflX-mediated drug resistance through ribosome splitting and rRNA disordering in mycobacteria.
Proc.Natl.Acad.Sci.USA, 122, 2025
|
|
4NA8
 
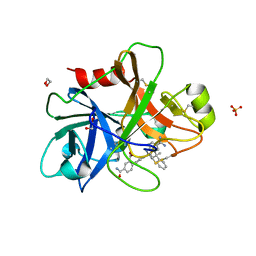 | | Factor XIa in complex with the inhibitor 5-aminocarbonyl-2-[3-[(2s,4r)-6-carbamimidoyl-4-methyl-4-phenyl-2,3-dihydro-1h-quinolin-2-yl]phenyl]benzoic acid | | Descriptor: | 1,2-ETHANEDIOL, 5-aminocarbonyl-2-[3-[(2S,4R)-6-carbamimidoyl-4-methyl-4-phenyl-2,3-dihydro-1H-quinolin-2-yl]phenyl]benzoic acid, Coagulation factor XI, ... | | Authors: | Wei, A. | | Deposit date: | 2013-10-21 | | Release date: | 2014-02-12 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Tetrahydroquinoline Derivatives as Potent and Selective Factor XIa Inhibitors.
J.Med.Chem., 57, 2014
|
|
8VY0
 
 | | Mango II bound to 365A-087 | | Descriptor: | 2-[(E)-(1-methylquinolin-4(1H)-ylidene)methyl]-3-[2-oxo-2-({[(2P)-2-(1H-pyrazol-1-yl)phenyl]methyl}amino)ethyl]-1,3-benzothiazol-3-ium, Chains: A,B,C, POTASSIUM ION | | Authors: | Passalacqua, L.F.M, Ferre-D'Amare, A.R, Schneekloth, J.S. | | Deposit date: | 2024-02-06 | | Release date: | 2025-02-19 | | Last modified: | 2025-10-08 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structure-informed design of an ultrabright RNA-activated fluorophore.
Nat.Chem., 17, 2025
|
|
6SW2
 
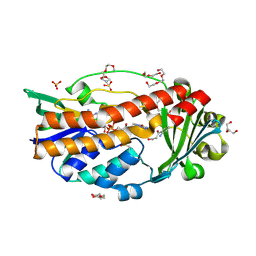 | |
8W3Y
 
 | |
1DGL
 
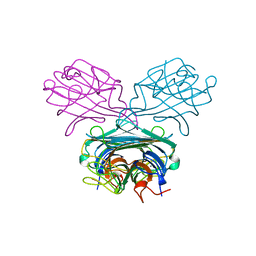 | | LECTIN FROM DIOCLEA GRANDIFLORA COMPLEXED TO TRIMANNOSIDE | | Descriptor: | CALCIUM ION, LECTIN, MANGANESE (II) ION, ... | | Authors: | Rozwarski, D.A, Swami, B.M, Brewer, C.F, Sacchettini, J.C. | | Deposit date: | 1998-02-10 | | Release date: | 1999-06-01 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of the lectin from Dioclea grandiflora complexed with core trimannoside of asparagine-linked carbohydrates.
J.Biol.Chem., 273, 1998
|
|
8VMD
 
 | |
