1TXS
 
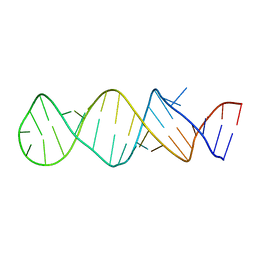 | | STEM-LOOP D OF THE CLOVERLEAF DOMAIN OF ENTEROVIRAL 5'UTR RNA | | Descriptor: | Enteroviral 5'-UTR | | Authors: | Du, Z, Yu, J, Ulyanov, N.B, Andino, R, James, T.L. | | Deposit date: | 2004-07-06 | | Release date: | 2004-10-05 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of a Consensus Stem-Loop D RNA Domain that Plays Important Roles in Regulating Translation and Replication in Enteroviruses and Rhinoviruses
Biochemistry, 43, 2004
|
|
1U0P
 
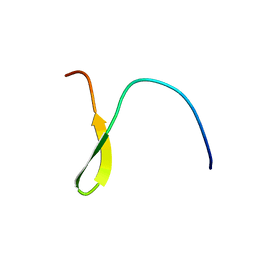 | | Stable A-state hairpin of T4 fibritin foldon | | Descriptor: | fibritin | | Authors: | Meier, S, Guthe, S, Kiefhaber, T, Grzesiek, S. | | Deposit date: | 2004-07-14 | | Release date: | 2005-02-22 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Foldon, the natural trimerization domain of T4 fibritin, dissociates into a monomeric A-state form containing a stable beta-hairpin: atomic details of trimer dissociation and
local beta-hairpin stability from residual dipolar couplings
J.Mol.Biol., 344, 2004
|
|
1T5M
 
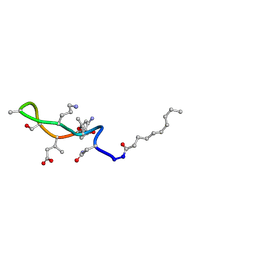 | | Structural transitions as determinants of the action of the calcium-dependent antibiotic daptomycin | | Descriptor: | DAPTOMYCIN, DECANOIC ACID | | Authors: | Jung, D, Rozek, A, Okon, M, Hancock, R.E. | | Deposit date: | 2004-05-04 | | Release date: | 2004-08-31 | | Last modified: | 2012-12-12 | | Method: | SOLUTION NMR | | Cite: | Structural Transitions as Determinants of the Action of the Calcium-Dependent Antibiotic Daptomycin.
Chem.Biol., 11, 2004
|
|
1VD2
 
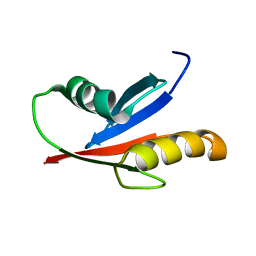 | | Solution Structure of the PB1 domain of PKCiota | | Descriptor: | Protein kinase C, iota type | | Authors: | Hirano, Y, Yoshinaga, S, Yokochi, M, Ogura, K, Noda, Y, Sumimoto, H, Inagaki, F. | | Deposit date: | 2004-03-18 | | Release date: | 2004-09-14 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution structure of atypical protein kinase C PB1 domain and its mode of interaction with ZIP/p62 and MEK5
J.Biol.Chem., 279, 2004
|
|
1HS7
 
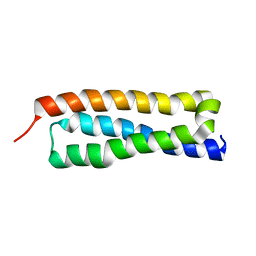 | | VAM3P N-TERMINAL DOMAIN SOLUTION STRUCTURE | | Descriptor: | SYNTAXIN VAM3 | | Authors: | Dulubova, I, Yamaguchi, T, Wang, Y, Sudhof, T.C, Rizo, J. | | Deposit date: | 2000-12-24 | | Release date: | 2001-03-07 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Vam3p structure reveals conserved and divergent properties of syntaxins.
Nat.Struct.Biol., 8, 2001
|
|
1J2M
 
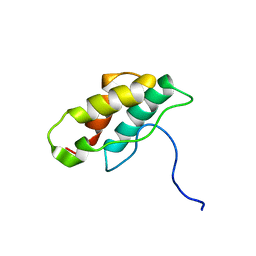 | | Solution structure of CPI-17(22-120) | | Descriptor: | 17-kDa PKC-potentiated inhibitory protein of PP1 | | Authors: | Ohki, S, Eto, M, Takada, R, Shimizu, M, Brautigan, D.L, Kainosho, M. | | Deposit date: | 2003-01-07 | | Release date: | 2003-06-17 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Distinctive Solution Conformation of Phosphatase Inhibitor CPI-17 Substituted with Aspartate at the Phosphorylation-site Threonine Residue
J.Mol.Biol., 326, 2003
|
|
1IRZ
 
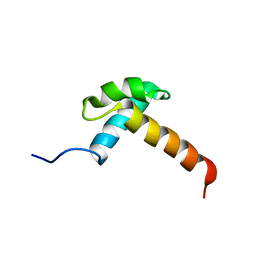 | |
1K5W
 | | THREE-DIMENSIONAL STRUCTURE OF THE SYNAPTOTAGMIN 1 C2B-DOMAIN: SYNAPTOTAGMIN 1 AS A PHOSPHOLIPID BINDING MACHINE | | Descriptor: | CALCIUM ION, Synaptotagmin I | | Authors: | Fernandez, I, Arac, D, Ubach, J, Gerber, S.H, Shin, O, Gao, Y, Anderson, R.G.W, Sudhof, T.C, Rizo, J. | | Deposit date: | 2001-10-12 | | Release date: | 2002-01-23 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Three-dimensional structure of the synaptotagmin 1 C2B-domain: synaptotagmin 1 as a phospholipid binding machine.
Neuron, 32, 2001
|
|
1UB1
 
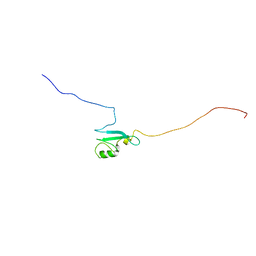 | | Solution structure of the matrix attachment region-binding domain of chicken MeCP2 | | Descriptor: | attachment region binding protein | | Authors: | Heitmann, B, Maurer, T, Weitzel, J.M, Stratling, W.H, Kalbitzer, H.R, Brunner, E, Structural Proteomics in Europe (SPINE) | | Deposit date: | 2003-03-27 | | Release date: | 2003-08-05 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the matrix attachment region-binding domain of chicken MeCP2
EUR.J.BIOCHEM., 270, 2003
|
|
1I46
 
 | |
1I4C
 | |
1I6D
 
 | | SOLUTION STRUCTURE OF THE FUNCTIONAL DOMAIN OF PARACOCCUS DENITRIFICANS CYTOCHROME C552 IN THE REDUCED STATE | | Descriptor: | CYTOCHROME C552, HEME C | | Authors: | Reincke, B, Perez, C, Pristovsek, P, Luecke, C, Ludwig, C, Loehr, F, Rogov, V.V, Ludwig, B, Rueterjans, H. | | Deposit date: | 2001-03-02 | | Release date: | 2001-10-17 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Solution structure and dynamics of the functional domain of Paracoccus denitrificans cytochrome c(552) in both redox states.
Biochemistry, 40, 2001
|
|
1I4B
 
 | |
1RVH
 
 | | SOLUTION STRUCTURE OF THE DNA DODECAMER GCAAAATTTTGC | | Descriptor: | 5'-D(*GP*CP*AP*AP*AP*AP*TP*TP*TP*TP*GP*C)-3' | | Authors: | Stefl, R, Wu, H, Ravindranathan, S, Sklenar, V, Feigon, J. | | Deposit date: | 2003-12-13 | | Release date: | 2004-02-10 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | DNA A-tract bending in three dimensions: Solving the dA4T4 vs. dT4A4 conundrum.
Proc.Natl.Acad.Sci.USA, 101, 2004
|
|
1RVI
 | | SOLUTION STRUCTURE OF THE DNA DODECAMER CGTTTTAAAACG | | Descriptor: | 5'-D(*CP*GP*TP*TP*TP*TP*AP*AP*AP*AP*CP*G)-3' | | Authors: | Stefl, R, Wu, H, Ravindranathan, S, Sklenar, V, Feigon, J. | | Deposit date: | 2003-12-13 | | Release date: | 2004-02-10 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | DNA A-tract bending in three dimensions: Solving the dA4T4 vs. dT4A4 conundrum.
Proc.Natl.Acad.Sci.USA, 101, 2004
|
|
1L1I
 
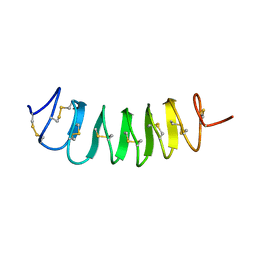 | | Solution Structure of the Tenebrio molitor Antifreeze Protein | | Descriptor: | Thermal hysteresis protein isoform YL-1 (2-14) | | Authors: | Daley, M.E, Spyracopoulos, L, Jia, Z, Davies, P.L, Sykes, B.D. | | Deposit date: | 2002-02-16 | | Release date: | 2002-05-22 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Structure and dynamics of a beta-helical antifreeze protein.
Biochemistry, 41, 2002
|
|
1KIO
 
 | | SOLUTION STRUCTURE OF THE SMALL SERINE PROTEASE INHIBITOR SGCI[L30R, K31M] | | Descriptor: | SERINE PROTEASE INHIBITOR I | | Authors: | Gaspari, Z, Patthy, A, Graf, L, Perczel, A. | | Deposit date: | 2001-12-03 | | Release date: | 2001-12-12 | | Last modified: | 2021-10-27 | | Method: | SOLUTION NMR | | Cite: | Comparative structure analysis of proteinase inhibitors from the desert locust, Schistocerca gregaria.
Eur.J.Biochem., 269, 2002
|
|
1I6E
 
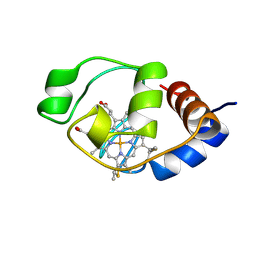 | | SOLUTION STRUCTURE OF THE FUNCTIONAL DOMAIN OF PARACOCCUS DENITRIFICANS CYTOCHROME C552 IN THE OXIDIZED STATE | | Descriptor: | CYTOCHROME C552, HEME C | | Authors: | Reincke, B, Perez, C, Pristovsek, P, Luecke, C, Ludwig, C, Loehr, F, Rogov, V.V, Ludwig, B, Rueterjans, H. | | Deposit date: | 2001-03-02 | | Release date: | 2001-10-17 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Solution structure and dynamics of the functional domain of Paracoccus denitrificans cytochrome c(552) in both redox states.
Biochemistry, 40, 2001
|
|
1LA4
 
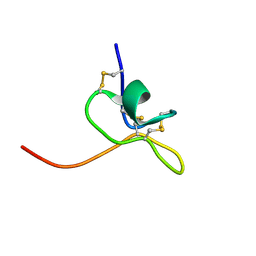 | | Solution Structure of SGTx1 | | Descriptor: | SGTx1 | | Authors: | Lee, C.W, Roh, S.H, Kim, S, Endoh, H, Kodera, Y, Maeda, T, Swartz, K.J, Kim, J.I. | | Deposit date: | 2002-03-28 | | Release date: | 2003-11-11 | | Last modified: | 2022-02-23 | | Method: | SOLUTION NMR | | Cite: | Solution Structure and Functional Characterization of SGTx1, a Modifier of Kv2.1 Channel Gating
Biochemistry, 43, 2004
|
|
1LL8
 
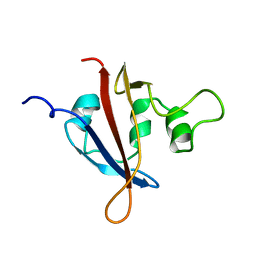 | |
1J2N
 
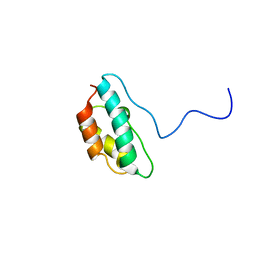 | | Solution structure of CPI-17(22-120) T38D | | Descriptor: | 17-kDa PKC-potentiated inhibitory protein of PP1 | | Authors: | Ohki, S, Eto, M, Shimizu, M, Takada, R, Brautigan, D.L, Kainosho, M. | | Deposit date: | 2003-01-07 | | Release date: | 2003-06-17 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Distinctive Solution Conformation of Phosphatase Inhibitor CPI-17 Substituted with Aspartate at the Phosphorylation-site Threonine Residue
J.Mol.Biol., 326, 2003
|
|
1T6R
 
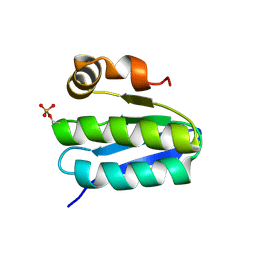 | | Solution structure of TM1442, a putative anti sigma factor antagonist in phosphorylated state | | Descriptor: | Putative anti-sigma factor antagonist TM1442 | | Authors: | Etezady-Esfarjani, T, Placzek, W, Herrmann, T, Lesley, S.A, Wuthrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2004-05-07 | | Release date: | 2005-05-24 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Solution structures of the putative anti-sigma-factor antagonist TM1442 from Thermotoga maritima in the free and phosphorylated states.
Magn.Reson.Chem., 44 Spec No, 2006
|
|
1LWR
 
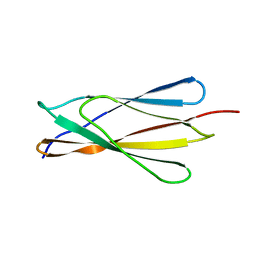 | | Solution structure of the NCAM fibronectin type III module 2 | | Descriptor: | Neural Cell Adhesion Molecule 1, 140 kDa isoform | | Authors: | Kiselyov, V.V, Skladchikova, G, Hinsby, A.M, Jensen, P.H, Kulahin, N, Pedersen, N, Tsetlin, V, Poulsen, F.M, Berezin, V, Bock, E. | | Deposit date: | 2002-06-03 | | Release date: | 2003-06-03 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural basis for a direct interaction between FGFR1 and NCAM and evidence for a regulatory role of ATP
Structure, 11, 2003
|
|
1LVJ
 
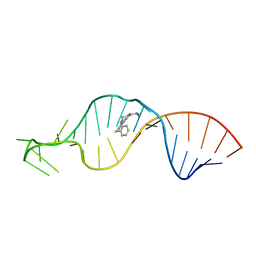 | |
1SS6
 
 | | Solution structure of SEP domain from human p47 | | Descriptor: | NSFL1 cofactor p47 | | Authors: | Soukenik, M, Leidert, M, Sievert, V, Buessow, K, Leitner, D, Labudde, D, Ball, L.J, Oschkinat, H. | | Deposit date: | 2004-03-23 | | Release date: | 2004-11-09 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The SEP domain of p47 acts as a reversible competitive inhibitor of cathepsin L
FEBS Lett., 576, 2004
|
|
