7PFV
 
 | |
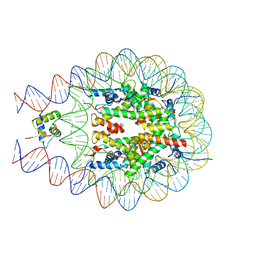
7PFW
 | |
1ECU
 
 | | SOLUTION STRUCTURE OF E2F BINDING DNA FRAGMENT GCGCGAAAC-T-GTTTCGCGC | | Descriptor: | DNA (5'-D(*GP*CP*GP*CP*GP*AP*AP*AP*CP*TP*GP*TP*TP*TP*CP*GP*CP*GP*C)-3') | | Authors: | Wu, J.H, Chang, C, Pei, J.M, Xiao, Q, Shi, Y.Y. | | Deposit date: | 2000-01-26 | | Release date: | 2000-02-02 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of E2F binding DNA fragment GCGCGAAAC-T-GTTTCGCGC studied by Molecular Dynamics Simulation and Two Dimensional NMR experiment
to be published, 2000
|
|
1QP5
 
 | | BASE-PAIRING SHIFT IN A DODECAMER CONTAINING A (CA)N TRACT | | Descriptor: | DNA (5'-D(*AP*CP*CP*GP*GP*CP*GP*CP*CP*AP*CP*A)-3'), DNA (5'-D(*TP*GP*TP*GP*GP*CP*GP*CP*CP*GP*GP*T)-3'), MAGNESIUM ION | | Authors: | Timsit, Y, Vilbois, E, Moras, D. | | Deposit date: | 1999-06-01 | | Release date: | 1999-06-14 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Base-pairing shift in the major groove of (CA)n tracts by B-DNA crystal structures.
Nature, 354, 1991
|
|
8CUC
 
 | |
3HR3
 
 | |
1TEZ
 
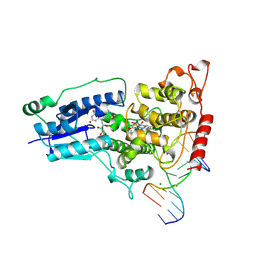 | | COMPLEX BETWEEN DNA AND THE DNA PHOTOLYASE FROM ANACYSTIS NIDULANS | | Descriptor: | 5'-D(*AP*TP*CP*GP*GP*CP*T*(TCP)P*CP*GP*C)-3', 5'-D(*TP*CP*GP*C)-3', 5'-D(P*CP*GP*AP*AP*GP*CP*CP*GP*A)-3', ... | | Authors: | Essen, L.-O, Carell, T, Mees, A, Klar, T. | | Deposit date: | 2004-05-26 | | Release date: | 2004-12-14 | | Last modified: | 2024-07-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of a photolyase bound to a CPD-like DNA lesion after in situ repair
Science, 306, 2004
|
|
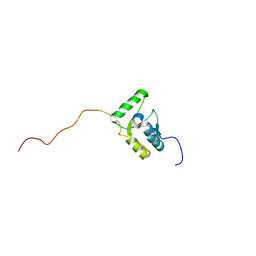
2DUN
 | |
8DXV
 
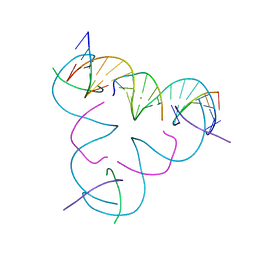 | | [T:Au+:T--(pH11-pH7; 5s)] Metal-mediated DNA base pair in tensegrity triangle grown at pH 11 and soaked in pH 7 for 5s | | Descriptor: | DNA (5'-D(*GP*AP*GP*CP*AP*GP*CP*CP*TP*GP*TP*TP*TP*GP*GP*AP*CP*AP*TP*CP*A)-3'), DNA (5'-D(P*CP*CP*AP*TP*AP*CP*A)-3'), DNA (5'-D(P*CP*TP*GP*AP*TP*GP*T)-3'), ... | | Authors: | Lu, B, Vecchioni, S, Seeman, N.C, Sha, R, Ohayon, Y.P. | | Deposit date: | 2022-08-03 | | Release date: | 2023-08-09 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (4.59 Å) | | Cite: | Crystallographic pH Titration of Silver and Mercury Mediated DNA Base Pairs
To Be Published
|
|
8DXX
 
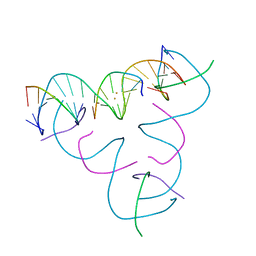 | | [T:Au+:T--(pH11-pH7; 45s)] Metal-mediated DNA base pair in tensegrity triangle grown at pH 11 and soaked in pH 7 for 45s | | Descriptor: | DNA (5'-D(*GP*AP*GP*CP*AP*GP*CP*CP*TP*GP*TP*TP*TP*GP*GP*AP*CP*AP*TP*CP*A)-3'), DNA (5'-D(P*CP*CP*AP*TP*AP*CP*A)-3'), DNA (5'-D(P*CP*TP*GP*AP*TP*GP*T)-3'), ... | | Authors: | Lu, B, Vecchioni, S, Seeman, N.C, Sha, R, Ohayon, Y.P. | | Deposit date: | 2022-08-03 | | Release date: | 2023-08-09 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (4.07 Å) | | Cite: | Crystallographic pH Titration of Silver and Mercury Mediated DNA Base Pairs
To Be Published
|
|
8DXY
 
 | | [T:Au+:T--(pH11-pH7; 180s)] Metal-mediated DNA base pair in tensegrity triangle grown at pH 11 and soaked in pH 7 for 180s | | Descriptor: | DNA (5'-D(*GP*AP*GP*CP*AP*GP*CP*CP*TP*GP*TP*TP*TP*GP*GP*AP*CP*AP*TP*CP*A)-3'), DNA (5'-D(P*CP*CP*AP*TP*AP*CP*A)-3'), DNA (5'-D(P*CP*TP*GP*AP*TP*GP*T)-3'), ... | | Authors: | Lu, B, Vecchioni, S, Seeman, N.C, Sha, R, Ohayon, Y.P. | | Deposit date: | 2022-08-03 | | Release date: | 2023-08-09 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (4.87 Å) | | Cite: | Crystallographic pH Titration of Silver and Mercury Mediated DNA Base Pairs
To Be Published
|
|
8DXW
 
 | | [T:Au+:T--(pH11-pH7; 15s)] Metal-mediated DNA base pair in tensegrity triangle grown at pH 11 and soaked in pH 7 for 15s | | Descriptor: | DNA (5'-D(*GP*AP*GP*CP*AP*GP*CP*CP*TP*GP*TP*TP*TP*GP*GP*AP*CP*AP*TP*CP*A)-3'), DNA (5'-D(P*CP*CP*AP*TP*AP*CP*A)-3'), DNA (5'-D(P*CP*TP*GP*AP*TP*GP*T)-3'), ... | | Authors: | Lu, B, Vecchioni, S, Seeman, N.C, Sha, R, Ohayon, Y.P. | | Deposit date: | 2022-08-03 | | Release date: | 2023-08-09 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (4.52 Å) | | Cite: | Crystallographic pH Titration of Silver and Mercury Mediated DNA Base Pairs
To Be Published
|
|
8DXZ
 
 | | [T:Au+:T--(pH11-pH7; 420s)] Metal-mediated DNA base pair in tensegrity triangle grown at pH 11 and soaked in pH 7 for 420s | | Descriptor: | DNA (5'-D(*GP*AP*GP*CP*AP*GP*CP*CP*TP*GP*TP*TP*TP*GP*GP*AP*CP*AP*TP*CP*A)-3'), DNA (5'-D(P*CP*CP*AP*TP*AP*CP*A)-3'), DNA (5'-D(P*CP*TP*GP*AP*TP*GP*T)-3'), ... | | Authors: | Lu, B, Vecchioni, S, Seeman, N.C, Sha, R, Ohayon, Y.P. | | Deposit date: | 2022-08-03 | | Release date: | 2023-08-09 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (4.71 Å) | | Cite: | Crystallographic pH Titration of Silver and Mercury Mediated DNA Base Pairs
To Be Published
|
|
1DCW
 
 | | STRUCTURE OF A FOUR-WAY JUNCTION IN AN INVERTED REPEAT SEQUENCE. | | Descriptor: | DNA (5'-D(*CP*CP*GP*GP*TP*AP*CP*CP*GP*G)-3'), SODIUM ION | | Authors: | Eichman, B.F, Vargason, J.M, Mooers, B.H.M, Ho, P.S. | | Deposit date: | 1999-11-05 | | Release date: | 2000-04-17 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The Holliday junction in an inverted repeat DNA sequence: sequence effects on the structure of four-way junctions.
Proc.Natl.Acad.Sci.USA, 97, 2000
|
|
1EYU
 
 | |
6FRH
 
 | | Crystal structure of Ssp DnaB Mini-Intein variant M86 | | Descriptor: | Replicative DNA helicase,Replicative DNA helicase | | Authors: | Popp, M.A, Blankenfeldt, W, Gazdag, M.E, Matern, J.J.C, Mootz, H.D. | | Deposit date: | 2018-02-15 | | Release date: | 2019-01-30 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | A functional interplay between intein and extein sequences in protein splicing compensates for the essential block B histidine.
Chem Sci, 10, 2019
|
|
6FRE
 
 | | Crystal structure of G-1F/H73A mutant of Ssp DnaB Mini-Intein variant M86 | | Descriptor: | Replicative DNA helicase,Replicative DNA helicase | | Authors: | Popp, M.A, Blankenfeldt, W, Friedel, K, Mootz, H.D. | | Deposit date: | 2018-02-15 | | Release date: | 2019-01-30 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.22 Å) | | Cite: | A functional interplay between intein and extein sequences in protein splicing compensates for the essential block B histidine.
Chem Sci, 10, 2019
|
|
7PF4
 
 | |
237D
 
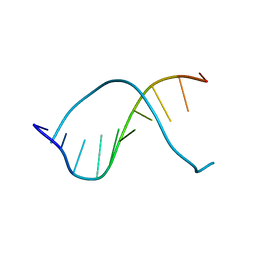 | | CRYSTAL STRUCTURE OF A DNA DECAMER SHOWING A NOVEL PSEUDO FOUR-WAY HELIX-HELIX JUNCTION | | Descriptor: | DNA (5'-D(*CP*GP*CP*AP*AP*TP*TP*GP*CP*G)-3') | | Authors: | Spink, N, Nunn, C.M, Vojtechovsky, J, Berman, H.M, Neidle, S. | | Deposit date: | 1995-09-28 | | Release date: | 1996-03-22 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of a DNA decamer showing a novel pseudo four-way helix-helix junction.
Proc.Natl.Acad.Sci.USA, 92, 1995
|
|
1C7Y
 
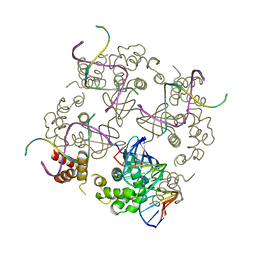 | | E.COLI RUVA-HOLLIDAY JUNCTION COMPLEX | | Descriptor: | DNA (5'-D(P*DAP*DAP*DGP*DTP*DTP*DGP*DGP*DGP*DAP*DTP*DTP*DGP*DT)-3'), DNA (5'-D(P*DCP*DAP*DAP*DTP*DCP*DCP*DCP*DAP*DAP*DCP*DTP*DT)-3'), DNA (5'-D(P*DCP*DGP*DAP*DAP*DTP*DGP*DTP*DGP*DTP*DGP*DTP*DCP*DT)-3'), ... | | Authors: | Ariyoshi, M, Nishino, T, Iwasaki, H, Shinagawa, H, Morikawa, K. | | Deposit date: | 2000-04-03 | | Release date: | 2000-07-21 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Crystal structure of the holliday junction DNA in complex with a single RuvA tetramer.
Proc.Natl.Acad.Sci.USA, 97, 2000
|
|
2N9H
 
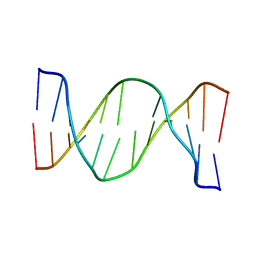 | | Glucose as a nuclease mimic in DNA | | Descriptor: | DNA (5'-D(*CP*TP*AP*GP*CP*(GL6)P*GP*TP*CP*AP*TP*C)-3'), DNA (5'-D(*GP*AP*TP*GP*AP*CP*TP*GP*CP*TP*AP*G)-3') | | Authors: | Gomez-Pinto, I, Vengut-Climent, E, Lucas, R, Avino, A, Eritja, R, Gonzalez-Ibanez, C, Morales, J, Muro, A, Penalver, P, Fonseca-Guerra, C, Bickelhaupt, M. | | Deposit date: | 2015-11-25 | | Release date: | 2016-08-31 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Glucose-Nucleobase Pseudo Base Pairs: Biomolecular Interactions within DNA.
Angew.Chem.Int.Ed.Engl., 55, 2016
|
|
2JPD
 
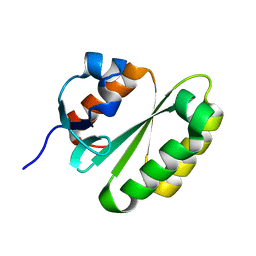 | | Solution structure of the ERCC1 central domain | | Descriptor: | DNA excision repair protein ERCC-1 | | Authors: | Tripsianes, K, Folkers, G, Zheng, C, Das, D, Grinstead, J.S, Kaptein, R, Boelens, R. | | Deposit date: | 2007-05-06 | | Release date: | 2007-09-04 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Analysis of the XPA and ssDNA-binding surfaces on the central domain of human ERCC1 reveals evidence for subfunctionalization
Nucleic Acids Res., 35, 2007
|
|
218D
 
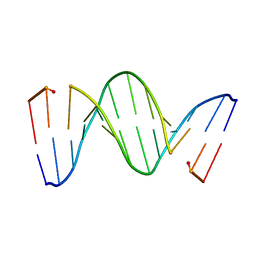 | | THE STRUCTURE OF A NEW CRYSTAL FORM OF A DNA DODECAMER CONTAINING T.(O6ME)G BASE PAIRS | | Descriptor: | DNA (5'-D(*CP*GP*TP*GP*AP*AP*TP*TP*CP*(6OG)P*CP*G)-3') | | Authors: | Vojtechovsky, J, Eaton, M.D, Gaffney, B, Jones, R, Berman, H.M. | | Deposit date: | 1995-07-18 | | Release date: | 1995-10-15 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structure of a new crystal form of a DNA dodecamer containing T.(O6Me)G base pairs.
Biochemistry, 34, 1995
|
|
6R0C
 
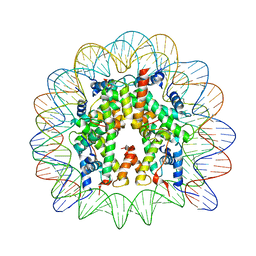 | | Human-D02 Nucleosome Core Particle with biotin-streptavidin label | | Descriptor: | DNA (142-MER), Histone H2A type 1, Histone H2B type 1-C/E/F/G/I, ... | | Authors: | Pye, V.E, Wilson, M.D, Cherepanov, P, Costa, A. | | Deposit date: | 2019-03-12 | | Release date: | 2019-09-25 | | Last modified: | 2024-05-15 | | Method: | ELECTRON MICROSCOPY (4.2 Å) | | Cite: | Retroviral integration into nucleosomes through DNA looping and sliding along the histone octamer.
Nat Commun, 10, 2019
|
|
233D
 
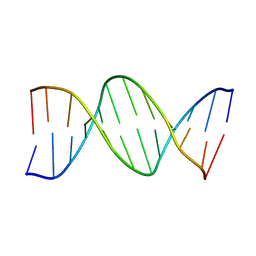 | | THE CRYSTAL STRUCTURE ANALYSIS OF D(CGCGAASSCGCG)2: A SYNTHETIC DNA DODECAMER DUPLEX CONTAINING FOUR 4'-THIO-2'-DEOXYTHYMIDINE NUCLEOTIDES | | Descriptor: | DNA (5'-D(*CP*GP*CP*GP*AP*AP*)-D(*(T49)P*(T49)P*)-D(*CP*GP*CP*G)-3') | | Authors: | Boggon, T.J, Hancox, E.L, McAuley-Hecht, K.E, Connolly, B.A, Hunter, W.N, Brown, T, Walker, R.T, Leonard, G.A. | | Deposit date: | 1995-10-16 | | Release date: | 1996-06-21 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | The crystal structure analysis of d(CGCGAASSCGCG)2, a synthetic DNA dodecamer duplex containing four 4'-thio-2'-deoxythymidine nucleotides.
Nucleic Acids Res., 24, 1996
|
|
