6C6D
 
 | |
4HCS
 
 | | Structure of Novel subfamily CX chemokine solved by sulfur SAD | | Descriptor: | Uncharacterized protein | | Authors: | Rajasekaran, D, Fan, C, Meng, W, Pflugrath, J.W, Lolis, E.J. | | Deposit date: | 2012-10-01 | | Release date: | 2013-10-16 | | Last modified: | 2014-04-23 | | Method: | X-RAY DIFFRACTION (1.28 Å) | | Cite: | Structural insight into the evolution of a new chemokine family from zebrafish.
Proteins, 82, 2014
|
|
2BDN
 
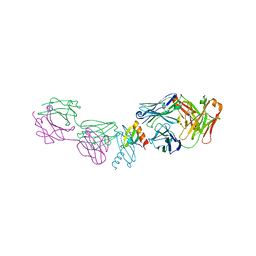 | | Crystal structure of human MCP-1 bound to a blocking antibody, 11K2 | | Descriptor: | Antibody heavy chain 11K2, Antibody light chain 11K2, Small inducible cytokine A2 | | Authors: | Boriack-Sjodin, P.A, Rushe, M, Reid, C, Jarpe, M, van Vlijmen, H, Bailly, V. | | Deposit date: | 2005-10-20 | | Release date: | 2006-06-13 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.53 Å) | | Cite: | Structure activity relationships of monocyte chemoattractant proteins in complex with a blocking antibody.
Protein Eng.Des.Sel., 19, 2006
|
|
4HED
 
 | | Zebrafish chemokine CXL1 | | Descriptor: | Uncharacterized protein | | Authors: | Rajasekaran, D, Fan, C, Meng, W, Pflugrath, J.W, Lolis, E.J. | | Deposit date: | 2012-10-03 | | Release date: | 2013-08-21 | | Last modified: | 2014-04-23 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Structural insight into the evolution of a new chemokine family from zebrafish.
Proteins, 82, 2014
|
|
6EHZ
 
 | |
6LOG
 
 | |
4HSV
 
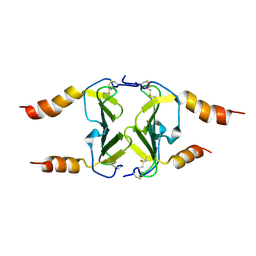 | | Crystal Structure of CXCL4L1 | | Descriptor: | Platelet factor 4 variant | | Authors: | Kuo, J.H, Liu, J.S, Wu, W.G, Sue, S.C. | | Deposit date: | 2012-10-30 | | Release date: | 2013-04-10 | | Last modified: | 2013-07-31 | | Method: | X-RAY DIFFRACTION (2.083 Å) | | Cite: | Alternative C-terminal helix orientation alters chemokine function: structure of the anti-angiogenic chemokine, CXCL4L1
J.Biol.Chem., 288, 2013
|
|
2EOT
 
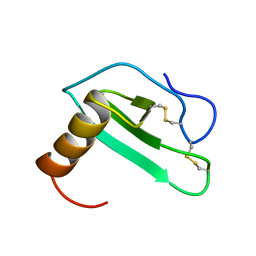 | | SOLUTION STRUCTURE OF EOTAXIN, AN ENSEMBLE OF 32 NMR SOLUTION STRUCTURES | | Descriptor: | EOTAXIN | | Authors: | Crump, M.P, Rajarathnam, K, Kim, K.-S, Clark-Lewis, I, Sykes, B.D. | | Deposit date: | 1998-06-29 | | Release date: | 1998-11-11 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Solution structure of eotaxin, a chemokine that selectively recruits eosinophils in allergic inflammation.
J.Biol.Chem., 273, 1998
|
|
2FJ2
 
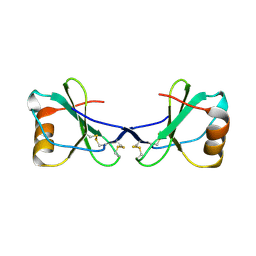 | | Crystal Structure of Viral Macrophage Inflammatory Protein-II | | Descriptor: | Viral macrophage inflammatory protein-II | | Authors: | Li, Y, Liu, D, Cao, R, Kumar, S, Dong, C.Z, wilson, S.R, Gao, Y.G, Huang, Z. | | Deposit date: | 2005-12-30 | | Release date: | 2006-12-26 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of chemically synthesized vMIP-II.
Proteins, 67, 2007
|
|
5IZB
 | |
2KEC
 
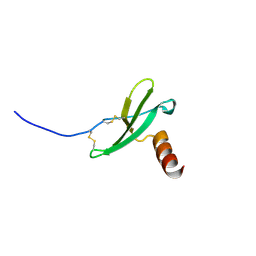 | |
2KED
 
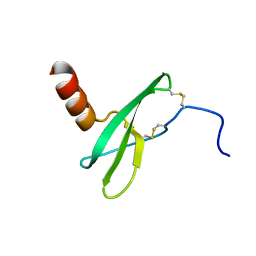 | |
7T1E
 
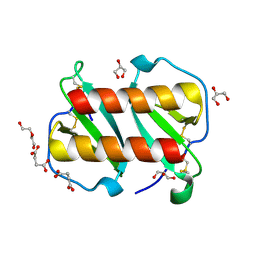 | | Structure of monomeric and dimeric human CCL20 | | Descriptor: | C-C motif chemokine 20, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Peterson, F.C, Riutta, S.J, Volkman, B.F. | | Deposit date: | 2021-12-01 | | Release date: | 2022-12-14 | | Last modified: | 2024-06-26 | | Method: | X-RAY DIFFRACTION (1.459 Å) | | Cite: | The Chemokine, CCL20, and Its Receptor, CCR6, in the Pathogenesis and Treatment of Psoriasis and Psoriatic Arthritis
J Psoriasis Psoriatic Arthritis, 8, 2023
|
|
2KEE
 
 | |
2KUM
 
 | | Solution structure of the human chemokine CCL27 | | Descriptor: | C-C motif chemokine 27 | | Authors: | Kirkpatrick, J.P, Jansma, A, Hsu, A, Handel, T.M, Nietlispach, D. | | Deposit date: | 2010-02-22 | | Release date: | 2010-03-02 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | NMR analysis of the structure, dynamics, and unique oligomerization properties of the chemokine CCL27.
J.Biol.Chem., 285, 2010
|
|
5L2U
 
 | |
5L7M
 
 | | Murin CXCL13 solution structure | | Descriptor: | C-X-C motif chemokine 13 | | Authors: | Monneau, Y.R, Lortat-Jacob, H. | | Deposit date: | 2016-06-03 | | Release date: | 2017-06-21 | | Last modified: | 2023-06-14 | | Method: | SOLUTION NMR | | Cite: | Solution structure of CXCL13 and heparan sulfate binding show that GAG binding site and cellular signalling rely on distinct domains.
Open Biol, 7, 2017
|
|
5LTL
 
 | |
7JNY
 
 | | Crystal structure of CXCL13 | | Descriptor: | C-X-C motif chemokine 13 | | Authors: | Rosenberg Jr, E.M, Rajasekaran, D, Murphy, J.W, Pantouris, G, Lolis, E.J. | | Deposit date: | 2020-08-05 | | Release date: | 2020-10-07 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | The N-terminal length and side-chain composition of CXCL13 affect crystallization, structure and functional activity.
Acta Crystallogr D Struct Biol, 76, 2020
|
|
1A15
 
 | | SDF-1ALPHA | | Descriptor: | STROMAL DERIVED FACTOR-1ALPHA, SULFATE ION | | Authors: | Dealwis, C.G, Fernandez, E.J, Lolis, E. | | Deposit date: | 1997-12-22 | | Release date: | 1998-08-12 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of chemically synthesized [N33A] stromal cell-derived factor 1alpha, a potent ligand for the HIV-1 "fusin" coreceptor.
Proc.Natl.Acad.Sci.USA, 95, 1998
|
|
1B2T
 
 | |
1B3A
 
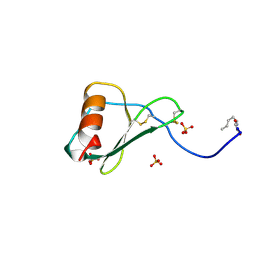 | | TOTAL CHEMICAL SYNTHESIS AND HIGH-RESOLUTION CRYSTAL STRUCTURE OF THE POTENT ANTI-HIV PROTEIN AOP-RANTES | | Descriptor: | PENTYLOXYAMINO-ACETALDEHYDE, PROTEIN (RANTES), SULFATE ION | | Authors: | Wilken, J, Hoover, D, Thompson, D.A, Barlow, P.N, Mcsparron, H, Picard, L, Wlodawer, A, Lubkowski, J, Kent, S.B.H. | | Deposit date: | 1998-12-07 | | Release date: | 1999-04-23 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Total chemical synthesis and high-resolution crystal structure of the potent anti-HIV protein AOP-RANTES.
Chem.Biol., 6, 1999
|
|
5WK3
 
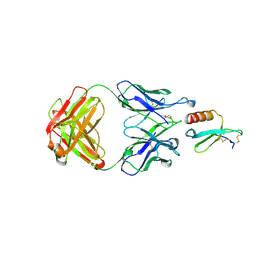 | | CRYSTAL STRUCTURE OF THE COMPLEX BETWEEN CCL17 AND M116 FAB | | Descriptor: | C-C motif chemokine 17, GLYCEROL, M116 HEAVY CHAIN, ... | | Authors: | Teplyakov, A, Obmolova, G, Gilliland, G.L. | | Deposit date: | 2017-07-24 | | Release date: | 2017-12-20 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural insights into chemokine CCL17 recognition by antibody M116.
Biochem Biophys Rep, 13, 2018
|
|
1B53
 
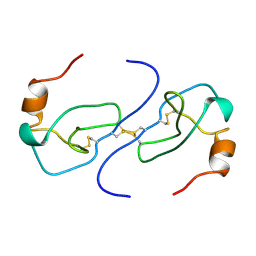 | | NMR STRUCTURE OF HUMAN MIP-1A D26A, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | MIP-1A | | Authors: | Waltho, J.P, Higgins, L.D, Craven, C.J, Tan, P, Dudgeon, T. | | Deposit date: | 1999-01-11 | | Release date: | 1999-07-22 | | Last modified: | 2021-11-03 | | Method: | SOLUTION NMR | | Cite: | Identification of amino acid residues critical for aggregation of human CC chemokines macrophage inflammatory protein (MIP)-1alpha, MIP-1beta, and RANTES. Characterization of active disaggregated chemokine variants.
J.Biol.Chem., 274, 1999
|
|
1B50
 
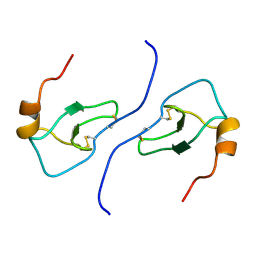 | | NMR STRUCTURE OF HUMAN MIP-1A D26A, 10 STRUCTURES | | Descriptor: | MIP-1A | | Authors: | Waltho, J.P, Higgins, L.D, Craven, C.J, Tan, P, Dudgeon, T. | | Deposit date: | 1999-01-11 | | Release date: | 1999-07-22 | | Last modified: | 2021-11-03 | | Method: | SOLUTION NMR | | Cite: | Identification of amino acid residues critical for aggregation of human CC chemokines macrophage inflammatory protein (MIP)-1alpha, MIP-1beta, and RANTES. Characterization of active disaggregated chemokine variants.
J.Biol.Chem., 274, 1999
|
|
