8XK8
 
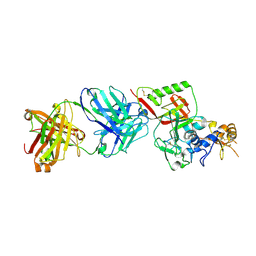 | | N1D10 Fab bound to SFTSV glycoprotein-Gn | | Descriptor: | Envelopment polyprotein, mAb N1D10 Fab heavy chain, mAb N1D10 Fab light chain | | Authors: | Zhao, H, Deng, Z. | | Deposit date: | 2023-12-22 | | Release date: | 2024-07-10 | | Last modified: | 2024-07-24 | | Method: | X-RAY DIFFRACTION (3.53 Å) | | Cite: | A broadly protective antibody targeting the glycoprotein Gn inhibits severe fever with thrombocytopenia syndrome virus infection.
To Be Published
|
|
8XJ0
 
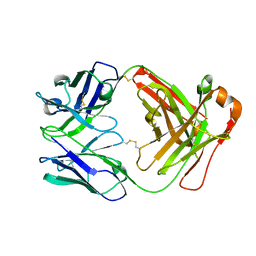 | | Crystal structure of AmFab mutant - P40C/E165C (Light chain), G10C/P210C(Heavy chain) | | Descriptor: | Adalimumab Fab Heavy chain, Adalimumab Fab Light chain | | Authors: | Senda, M, Yoshikawa, M, Nakamura, H, Ohkuri, T, Senda, T. | | Deposit date: | 2023-12-20 | | Release date: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Stabilization of adalimumab Fab through the introduction of disulfide bonds between the variable and constant domains.
Biochem.Biophys.Res.Commun., 700, 2024
|
|
8XIO
 
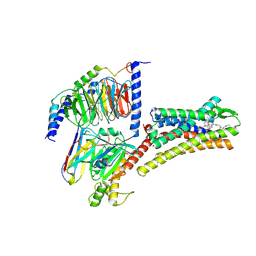 | | Structure of L797591-SSTR1 G protein complex | | Descriptor: | (2~{S})-~{N}-[(4~{S})-6-azanyl-2,2,4-trimethyl-hexyl]-3-naphthalen-1-yl-2-[[2-phenylethyl(2-pyridin-2-ylethyl)carbamoyl]amino]propanamide, G-alpha-i, Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, ... | | Authors: | Wang, Y, Xu, Y, Xu, H.E, Zhuang, Y. | | Deposit date: | 2023-12-19 | | Release date: | 2024-07-03 | | Method: | ELECTRON MICROSCOPY (2.65 Å) | | Cite: | Selective ligand recognition and activation of somatostatin receptors SSTR1 and SSTR3
To Be Published
|
|
8XI9
 
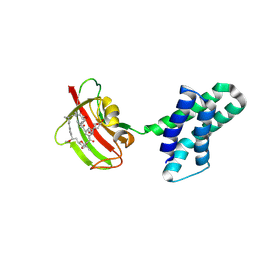 | | Crystal structure of FRB-FKBP fusion protein in complex with rapamycin | | Descriptor: | FRB-FKBP fusion protein, RAPAMYCIN IMMUNOSUPPRESSANT DRUG | | Authors: | Inobe, T, Sakaguchi, R, Obita, T, Mukaiyama, A, Yokoyama, T, Mizuguchi, M, Akiyama, S. | | Deposit date: | 2023-12-19 | | Release date: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural insights into rapamycin-induced oligomerization of a FRB-FKBP fusion protein.
Febs Lett., 2024
|
|
8XI6
 
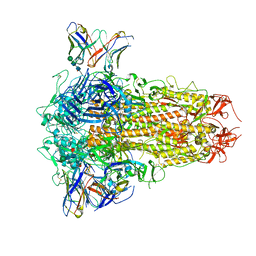 | | SARS-CoV-2 Omicron BQ.1.1 Variant Spike Protein Complexed with MO11 Fab | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-[alpha-L-fucopyranose-(1-6)]2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Ishimaru, H, Nishimura, M, Shigematsu, H, Marini, M.I, Hasegawa, N, Takamiya, R, Iwata, S, Mori, Y. | | Deposit date: | 2023-12-19 | | Release date: | 2024-04-24 | | Last modified: | 2024-05-29 | | Method: | ELECTRON MICROSCOPY (2.3 Å) | | Cite: | Epitopes of an antibody that neutralizes a wide range of SARS-CoV-2 variants in a conserved subdomain 1 of the spike protein.
J.Virol., 98, 2024
|
|
8XHU
 
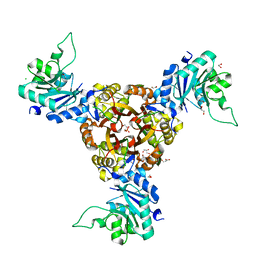 | | Crystal structure of Helicobacter pylori IspDF | | Descriptor: | 1,2-ETHANEDIOL, Bifunctional enzyme IspD/IspF, CHLORIDE ION, ... | | Authors: | Chen, X, Wu, D. | | Deposit date: | 2023-12-18 | | Release date: | 2024-04-10 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Two natural compounds as potential inhibitors against the Helicobacter pylori and Acinetobacter baumannii IspD enzymes.
Int J Antimicrob Agents, 63, 2024
|
|
8XHR
 
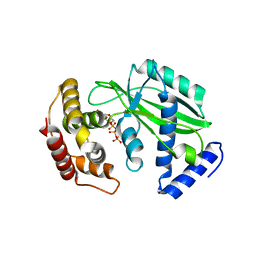 | |
8XGW
 
 | |
8XGV
 
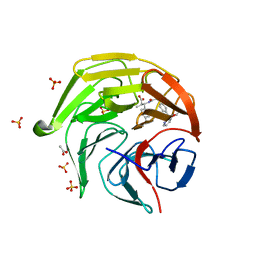 | | Optimization Efforts for Identification of Novel Highly Potent Keap1-Nrf2 Protein-Protein Interaction (PPI) Inhibitors | | Descriptor: | (2~{R},3~{S})-3-[[(2~{S})-2-[4-[(3-ethoxypyridin-2-yl)methyl]phenyl]-2-fluoranyl-ethanoyl]amino]-2-methyl-3-(4-methylphenyl)propanoic acid, ACETATE ION, Kelch-like ECH-associated protein 1, ... | | Authors: | Otake, K, Hara, Y, Ubukata, M, Inoue, M, Nagahashi, N, Motoda, D, Ogawa, N, Hantani, Y, Hantani, R, Adachi, T, Nomura, A, Yamaguchi, K, Maekawa, M, Mamada, H, Motomura, T, Sato, M, Harada, K. | | Deposit date: | 2023-12-15 | | Release date: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.42 Å) | | Cite: | Optimization Efforts for Identification of Novel Highly Potent Keap1-Nrf2 Protein-Protein Interaction Inhibitors.
J.Med.Chem., 67, 2024
|
|
8XGR
 
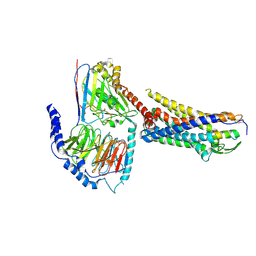 | | ETB-eGt complex bound to endothelin-1 | | Descriptor: | Camelid antibody VHH fragment, Endothelin receptor type B, Endothelin-1, ... | | Authors: | Oshima, H.S, Sano, F.K, Akasaka, H, Iwama, A, Shihoya, W, Nureki, O. | | Deposit date: | 2023-12-15 | | Release date: | 2024-04-03 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Optimizing cryo-EM structural analysis of G i -coupling receptors via engineered G t and Nb35 application.
Biochem.Biophys.Res.Commun., 693, 2024
|
|
8XGK
 
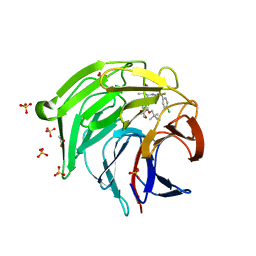 | | Optimization Efforts for Identification of Novel Highly Potent Keap1-Nrf2 Protein-Protein Interaction Ihhibitors | | Descriptor: | (2~{R},3~{S})-3-[[(2~{S})-2-(4-chlorophenyl)-2-fluoranyl-ethanoyl]amino]-3-[3-(2-cyano-2-methyl-propoxy)-4-methyl-phenyl]-2-methyl-propanoic acid, ACETATE ION, Kelch-like ECH-associated protein 1, ... | | Authors: | Otake, K, Hara, Y, Ubukata, M, Inoue, M, Nagahashi, N, Motoda, D, Ogawa, N, Hantani, Y, Hantani, R, Adachi, T, Nomura, A, Yamaguchi, K, Maekawa, M, Mamada, H, Motomura, T, Sato, M, Harada, K. | | Deposit date: | 2023-12-15 | | Release date: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.32 Å) | | Cite: | Optimization Efforts for Identification of Novel Highly Potent Keap1-Nrf2 Protein-Protein Interaction Inhibitors.
J.Med.Chem., 67, 2024
|
|
8XFY
 
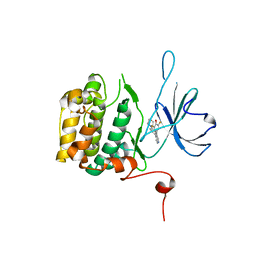 | | The Crystal Structure of RSK2 from Biortus. | | Descriptor: | 2,6-bis(fluoranyl)-4-[4-(4-morpholin-4-ylphenyl)pyridin-3-yl]phenol, Ribosomal protein S6 kinase alpha-3 | | Authors: | Wang, F, Cheng, W, Lv, Z, Lin, D, Pan, W. | | Deposit date: | 2023-12-14 | | Release date: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | The Crystal Structure of RSK2 from Biortus.
To Be Published
|
|
8XFR
 
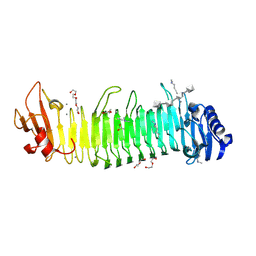 | |
8XFQ
 
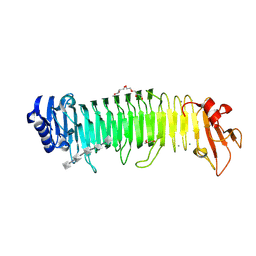 | |
8XF3
 
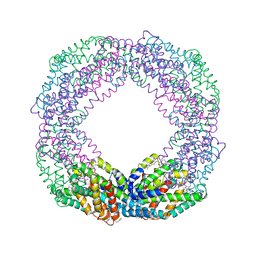 | | Crystal structure of the reassembled C-phycocyanin octamer from Thermoleptolyngbya sp. O-77 | | Descriptor: | C-phycocyanin alpha chain, C-phycocyanin beta chain, PHOSPHATE ION, ... | | Authors: | Nguyen, H.K, Teramoto, T, Kakuta, Y, Yoon, K.S. | | Deposit date: | 2023-12-13 | | Release date: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Disassembly and reassembly of the non-conventional thermophilic C-phycocyanin.
J.Biosci.Bioeng., 137, 2024
|
|
8XF2
 
 | | Crystal structure of the reassembled C-phycocyanin hexamer from Thermoleptolyngbya sp. O-77 | | Descriptor: | C-phycocyanin alpha chain, C-phycocyanin beta chain, PHOSPHATE ION, ... | | Authors: | Nguyen, H.K, Teramoto, T, Kakuta, Y, Yoon, K.S. | | Deposit date: | 2023-12-13 | | Release date: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Disassembly and reassembly of the non-conventional thermophilic C-phycocyanin.
J.Biosci.Bioeng., 137, 2024
|
|
8XF1
 
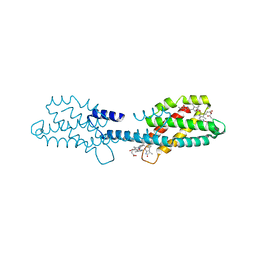 | | Crystal structure of the dissociated C-phycocyanin beta-chain from Thermoleptolyngbya sp. O-77 | | Descriptor: | C-phycocyanin beta chain, PHYCOCYANOBILIN | | Authors: | Nguyen, H.K, Teramoto, T, Kakuta, Y, Yoon, K.S. | | Deposit date: | 2023-12-13 | | Release date: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Disassembly and reassembly of the non-conventional thermophilic C-phycocyanin.
J.Biosci.Bioeng., 137, 2024
|
|
8XF0
 
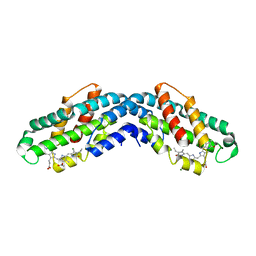 | | Crystal structure of the dissociated C-phycocyanin alpha-chain from Thermoleptolyngbya sp. O-77 | | Descriptor: | C-phycocyanin alpha chain, MAGNESIUM ION, PHYCOCYANOBILIN | | Authors: | Nguyen, H.K, Teramoto, T, Kakuta, Y, Yoon, K.S. | | Deposit date: | 2023-12-13 | | Release date: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Disassembly and reassembly of the non-conventional thermophilic C-phycocyanin.
J.Biosci.Bioeng., 137, 2024
|
|
8XEQ
 
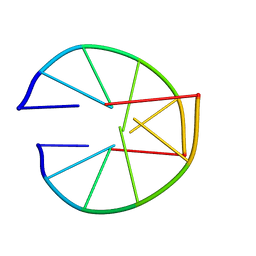 | |
8XEJ
 
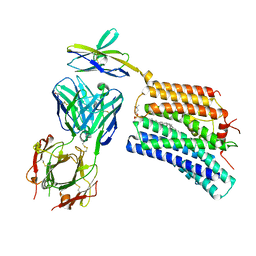 | | Cryo-EM structure of human XKR8-basigin complex in lipid nanodisc | | Descriptor: | 1,2-DILINOLEOYL-SN-GLYCERO-3-PHOSPHOCHOLINE, Fab heavy chain, Fab light chain, ... | | Authors: | Sakuragi, T.S, Kanai, R.K, Kikkawa, M.K, Toyoshima, C.T, Nagata, S.N. | | Deposit date: | 2023-12-12 | | Release date: | 2024-02-28 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (3.66 Å) | | Cite: | The role of the C-terminal tail region as a plug to regulate XKR8 lipid scramblase.
J.Biol.Chem., 300, 2024
|
|
8XE9
 
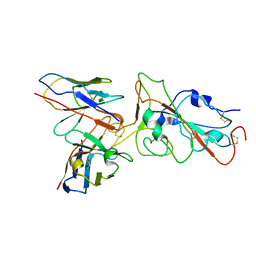 | | XBB.1.5 RBD in complex with BD55-1205 | | Descriptor: | BD55-1205 heavy chain, BD55-1205 light chain, Spike protein S2' | | Authors: | Feng, L.L. | | Deposit date: | 2023-12-11 | | Release date: | 2024-03-06 | | Last modified: | 2024-04-24 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | XBB.1.5 RBD in complex with BD55-1205
To Be Published
|
|
8XE7
 
 | |
8XE5
 
 | | norbelladine 4'-O-methyltransferase complexed with Mg, SAH, and 3,4-dihydroxybenzaldehyde | | Descriptor: | GLYCEROL, MAGNESIUM ION, Norbelladine 4'-O-methyltransferase, ... | | Authors: | Saw, Y.Y.H, Nakashima, Y, Morita, H. | | Deposit date: | 2023-12-11 | | Release date: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (2.07 Å) | | Cite: | Structure-Based Catalytic Mechanism of Amaryllidaceae O-Methyltransferases
Acs Catalysis, 2024
|
|
8XE4
 
 | | norbelladine 4'-O-methyltransferase complexed with Mg, SAH, and norbelladine | | Descriptor: | GLYCEROL, MAGNESIUM ION, Norbelladine, ... | | Authors: | Saw, Y.Y.H, Nakashima, Y, Morita, H. | | Deposit date: | 2023-12-11 | | Release date: | 2024-08-07 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Structure-Based Catalytic Mechanism of Amaryllidaceae O-Methyltransferases
Acs Catalysis, 2024
|
|
8XE3
 
 | |
