3N2X
 
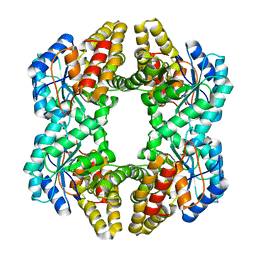 | | Crystal structure of YagE, a prophage protein belonging to the dihydrodipicolinic acid synthase family from E. coli K12 in complex with pyruvate | | Descriptor: | 1,2-ETHANEDIOL, Uncharacterized protein yagE | | Authors: | Bhaskar, V, Kumar, P.M, Manicka, S, Krishnaswamy, S. | | Deposit date: | 2010-05-19 | | Release date: | 2011-04-13 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Identification of biochemical and putative biological role of a xenolog from Escherichia coli using structural analysis.
Proteins, 79, 2011
|
|
3NEV
 
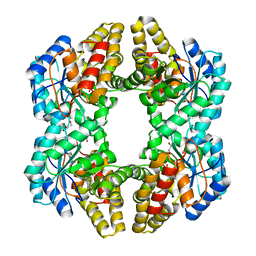 | | Crystal structure of YagE, a prophage protein from E. coli K12 in complex with KDGal | | Descriptor: | 1,2-ETHANEDIOL, 3-DEOXY-D-LYXO-HEXONIC ACID, Uncharacterized protein yagE | | Authors: | Bhaskar, V, Kumar, P.M, Manicka, S, Krishnaswamy, S. | | Deposit date: | 2010-06-09 | | Release date: | 2011-04-13 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Identification of biochemical and putative biological role of a xenolog from Escherichia coli using structural analysis.
Proteins, 79, 2011
|
|
5A70
 
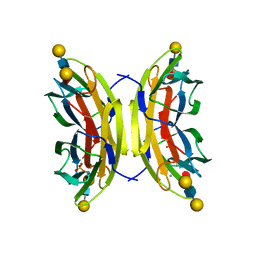 | | Structure of the LecB lectin from Pseudomonas aeruginosa strain PA14 in complex with lewis x tetrasaccharide | | Descriptor: | CALCIUM ION, LECB, SULFATE ION, ... | | Authors: | Sommer, R, Wagner, S, Varrot, A, Khaledi, A, Haussler, S, Imberty, A, Titz, A. | | Deposit date: | 2015-07-02 | | Release date: | 2016-07-27 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Induction of rare conformation of oligosaccharide by binding to calcium-dependent bacterial lectin: X-ray crystallography and modelling study.
Eur.J.Med.Chem., 177, 2019
|
|
8TXS
 
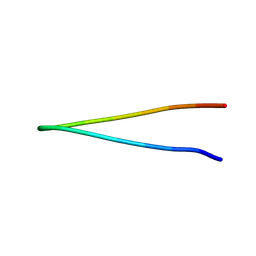 | |
8YP5
 
 | | The structure of MAP2K4 complexed with 5Z7-oxozeaenol | | Descriptor: | (3S,5Z,8S,9S,11E)-8,9,16-trihydroxy-14-methoxy-3-methyl-3,4,9,10-tetrahydro-1H-2-benzoxacyclotetradecine-1,7(8H)-dione, Dual specificity mitogen-activated protein kinase kinase 4, MAGNESIUM ION | | Authors: | Yumura, S, Kinishita, T. | | Deposit date: | 2024-03-15 | | Release date: | 2025-01-22 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Conserved gatekeeper methionine regulates the binding and access of kinase inhibitors to ATP sites of MAP2K1, 4, and 7: Clues for developing selective inhibitors.
Bioorg.Med.Chem.Lett., 112, 2024
|
|
7ZIE
 
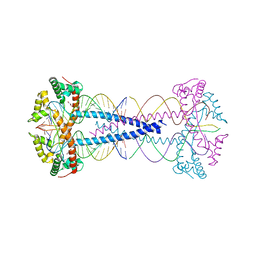 | |
7ANV
 
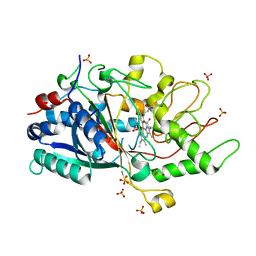 | |
8A6W
 
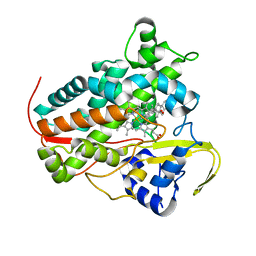 | |
8A9P
 
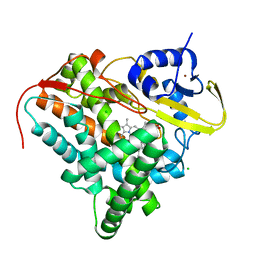 | | Crystal structure of CYP142 from Mycobacterium tuberculosis in complex with a fragment | | Descriptor: | (3-phenyl-1,2,4-oxadiazol-5-yl)methanamine, BROMIDE ION, CHLORIDE ION, ... | | Authors: | Snee, M, Katariya, M, Levy, C, Leys, D. | | Deposit date: | 2022-06-29 | | Release date: | 2023-07-12 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.63 Å) | | Cite: | Crystal structure of CYP142 from Mycobacterium tuberculosis in complex with a fragment
To Be Published
|
|
5MAL
 
 | |
6PT8
 | | Crystal Structure of CobT from Methanocaldococcus jannaschii in complex with Adenine Alpha-Ribotide and Nicotinic Acid | | Descriptor: | ALPHA-ADENOSINE MONOPHOSPHATE, NICOTINIC ACID, PHOSPHATE ION, ... | | Authors: | Schwarzwalder, A.H, Jeter, V.L, Vecellio, A.A, Erpenbach, E, Escalante, J.C, Rayment, I. | | Deposit date: | 2019-07-15 | | Release date: | 2020-07-22 | | Last modified: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structural studies of the phosphoribosyltransferase involved in cobamide biosynthesis in methanogenic archaea and cyanobacteria.
Sci Rep, 12, 2022
|
|
6TPP
 
 | |
6PTF
 
 | | Crystal Structure of CobT from Methanocaldococcus jannaschii in Apo State | | Descriptor: | UPF0284 protein MJ1598 | | Authors: | Schwarzwalder, A.H, Jeter, V.L, Vecellio, A.A, Erpenbach, E, Escalante-Semerena, J.C, Rayment, I. | | Deposit date: | 2019-07-15 | | Release date: | 2020-07-22 | | Last modified: | 2024-04-10 | | Method: | X-RAY DIFFRACTION (2.203 Å) | | Cite: | Structural studies of the phosphoribosyltransferase involved in cobamide biosynthesis in methanogenic archaea and cyanobacteria.
Sci Rep, 12, 2022
|
|
8ASO
 
 | | Nickel(II) bound to a non-canonical quadruplex | | Descriptor: | COBALT HEXAMMINE(III), DNA (5'-D(*GP*CP*AP*TP*GP*CP*T)-3'), NICKEL (II) ION | | Authors: | Lambert, M.C, Hall, J.P. | | Deposit date: | 2022-08-19 | | Release date: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.19 Å) | | Cite: | Identifying metal-DNA binding sites, what is the best method to get transition metals into a crystal system?
To Be Published
|
|
8ASM
 
 | | Cobalt(II) bound to a non-canonical quadruplex | | Descriptor: | COBALT (II) ION, COBALT HEXAMMINE(III), DNA (5'-D(*GP*CP*AP*TP*GP*CP*T)-3') | | Authors: | Lambert, M.C, Hall, J.P. | | Deposit date: | 2022-08-19 | | Release date: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Identifying metal-DNA binding sites, what is the best method to get transition metals into a crystal system?
To Be Published
|
|
5M4Z
 
 | | Crystal structure of the complex of T.spiralis thymidylate synthase with N(4)-hydroxy-2'-deoxycytidine-5'-monophosphate, crystallized in the presence of N(5,10)-methylenetetrahydrofolate | | Descriptor: | GLYCEROL, Thymidylate synthase, [(2~{R},3~{S},5~{R})-5-[(4~{E})-4-hydroxyimino-2-oxidanylidene-1,3-diazinan-1-yl]-3-oxidanyl-oxolan-2-yl]methyl dihydrogen phosphate | | Authors: | Wilk, P, Maj, P, Jarmula, A, Dowiercial, A, Rode, W. | | Deposit date: | 2016-10-19 | | Release date: | 2017-12-20 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.179 Å) | | Cite: | Molecular Mechanism of Thymidylate Synthase Inhibition by N 4 -Hydroxy-dCMP in View of Spectrophotometric and Crystallographic Studies.
Int J Mol Sci, 22, 2021
|
|
7BUP
 
 | |
5JPJ
 
 | | Crystal structure of 6aJL2-R24G | | Descriptor: | 6aJL2 protein | | Authors: | Pelaez-Aguilar, A.E, Diaz-Vilchis, A, Amero, C, Becerril, B, Rudino-Pinera, E. | | Deposit date: | 2016-05-03 | | Release date: | 2017-06-28 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of 6aJL2-R24G light chain variable domain: Does crystal packing explain amyloid fibril formation?
Biochem Biophys Rep, 20, 2019
|
|
6Q29
 
 | |
4JFA
 
 | | Crystal Structure of Plasmodium falciparum Tryptophanyl-tRNA synthetase | | Descriptor: | BETA-MERCAPTOETHANOL, POTASSIUM ION, TRYPTOPHAN, ... | | Authors: | Khan, S, Garg, A, Manickam, Y, Sharma, A. | | Deposit date: | 2013-02-28 | | Release date: | 2014-01-29 | | Last modified: | 2025-03-26 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | An appended domain results in an unusual architecture for malaria parasite tryptophanyl-tRNA synthetase
Plos One, 8, 2013
|
|
5DLP
 
 | | Acetylcholinesterase Methylene Blue no PEG | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 3,7-BIS(DIMETHYLAMINO)PHENOTHIAZIN-5-IUM, Acetylcholinesterase, ... | | Authors: | Dym, O. | | Deposit date: | 2015-09-07 | | Release date: | 2016-03-30 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The impact of crystallization conditions on structure-based drug design: A case study on the methylene blue/acetylcholinesterase complex.
Protein Sci., 25, 2016
|
|
7PU1
 
 | | High resolution X-ray structure of Thermoascus aurantiacus LPMO | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, COPPER (II) ION, ... | | Authors: | Banerjee, S, Frandsen, K.E.H, Singh, R.K, Bjerrum, M.J, Lo Leggio, L. | | Deposit date: | 2021-09-28 | | Release date: | 2022-03-16 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.06 Å) | | Cite: | Protonation State of an Important Histidine from High Resolution Structures of Lytic Polysaccharide Monooxygenases.
Biomolecules, 12, 2022
|
|
7EOX
 
 | | Protease structure from Euphorbia resinifera | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, IODIDE ION, Protease, ... | | Authors: | Siritapetawee, J, Attarataya, J, Charoenwattanasatien, R. | | Deposit date: | 2021-04-24 | | Release date: | 2022-01-12 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Sequence analysis and crystal structure of a glycosylated protease from Euphorbia resinifera latex for its proteolytic activity aspect.
Biotechnol Appl Biochem, 69, 2022
|
|
9GM4
 
 | |
9BYJ
 
 | | Crystal Structure of Hck in complex with the Src-family kinase inhibitor A-419259 | | Descriptor: | 1,2-ETHANEDIOL, 7-[trans-4-(4-methylpiperazin-1-yl)cyclohexyl]-5-(4-phenoxyphenyl)-7H-pyrrolo[2,3-d]pyrimidin-4-amine, DIMETHYL SULFOXIDE, ... | | Authors: | Selzer, A.M, Alvarado, J.J, Smithgall, T.E. | | Deposit date: | 2024-05-23 | | Release date: | 2024-10-09 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Cocrystallization of the Src-Family Kinase Hck with the ATP-Site Inhibitor A-419259 Stabilizes an Extended Activation Loop Conformation.
Biochemistry, 63, 2024
|
|
