1CSW
 
 | |
1CEY
 
 | | ASSIGNMENTS, SECONDARY STRUCTURE, GLOBAL FOLD, AND DYNAMICS OF CHEMOTAXIS Y PROTEIN USING THREE-AND FOUR-DIMENSIONAL HETERONUCLEAR (13C,15N) NMR SPECTROSCOPY | | Descriptor: | CHEY | | Authors: | Moy, F.J, Lowry, D.F, Matsumura, P, Dahlquist, F.W, Krywko, J.E, Domaille, P.J. | | Deposit date: | 1994-11-23 | | Release date: | 1995-02-07 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Assignments, secondary structure, global fold, and dynamics of chemotaxis Y protein using three- and four-dimensional heteronuclear (13C,15N) NMR spectroscopy.
Biochemistry, 33, 1994
|
|
1CFH
 
 | | STRUCTURE OF THE METAL-FREE GAMMA-CARBOXYGLUTAMIC ACID-RICH MEMBRANE BINDING REGION OF FACTOR IX BY TWO-DIMENSIONAL NMR SPECTROSCOPY | | Descriptor: | COAGULATION FACTOR IX, FORMIC ACID | | Authors: | Freedman, S.J, Furie, B.C, Furie, B, Baleja, J.D. | | Deposit date: | 1995-02-26 | | Release date: | 1995-07-10 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Structure of the metal-free gamma-carboxyglutamic acid-rich membrane binding region of factor IX by two-dimensional NMR spectroscopy.
J.Biol.Chem., 270, 1995
|
|
1CLD
 
 | | DNA-binding protein | | Descriptor: | CADMIUM ION, CD2-LAC9 | | Authors: | Gardner, K.H, Coleman, J.E. | | Deposit date: | 1995-06-06 | | Release date: | 1995-09-15 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the Kluyveromyces lactis LAC9 Cd2 Cys6 DNA-binding domain.
Nat.Struct.Biol., 2, 1995
|
|
1CM1
 
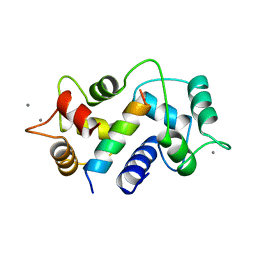 | | MOTIONS OF CALMODULIN-SINGLE-CONFORMER REFINEMENT | | Descriptor: | CALCIUM ION, CALMODULIN, CALMODULIN-DEPENDENT PROTEIN KINASE II-ALPHA | | Authors: | Wall, M.E, Phillips Jr, G.N. | | Deposit date: | 1997-09-23 | | Release date: | 1998-03-04 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Motions of calmodulin characterized using both Bragg and diffuse X-ray scattering.
Structure, 5, 1997
|
|
1COV
 
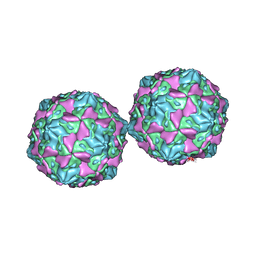 | | COXSACKIEVIRUS B3 COAT PROTEIN | | Descriptor: | COXSACKIEVIRUS COAT PROTEIN, MYRISTIC ACID, PALMITIC ACID | | Authors: | Muckelbauer, J.K, Rossmann, M.G. | | Deposit date: | 1994-10-19 | | Release date: | 1996-03-08 | | Last modified: | 2023-04-19 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Structure determination of coxsackievirus B3 to 3.5 A resolution.
Acta Crystallogr.,Sect.D, 51, 1995
|
|
1CM4
 
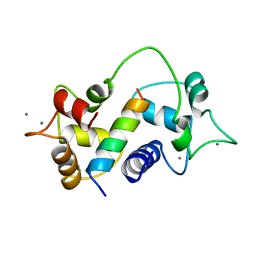 | | Motions of calmodulin-four-conformer refinement | | Descriptor: | CALCIUM ION, CALMODULIN, CALMODULIN-DEPENDENT PROTEIN KINASE II-ALPHA | | Authors: | Wall, M.E, Phillips Jr, G.N. | | Deposit date: | 1997-09-23 | | Release date: | 1998-03-04 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Motions of calmodulin characterized using both Bragg and diffuse X-ray scattering.
Structure, 5, 1997
|
|
1CL1
 
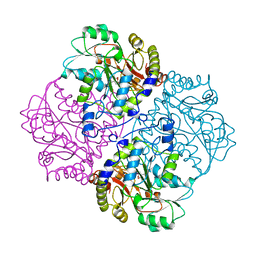 | |
1A4S
 
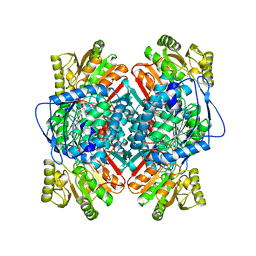 | | BETAINE ALDEHYDE DEHYDROGENASE FROM COD LIVER | | Descriptor: | BETAINE ALDEHYDE DEHYDROGENASE | | Authors: | Johansson, K, El Ahmad, M, Hjelmqvist, L, Ramaswamy, S, Jornvall, H, Eklund, H. | | Deposit date: | 1998-02-03 | | Release date: | 1998-04-08 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure of betaine aldehyde dehydrogenase at 2.1 A resolution.
Protein Sci., 7, 1998
|
|
1A74
 
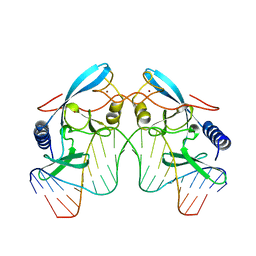 | | I-PPOL HOMING ENDONUCLEASE/DNA COMPLEX | | Descriptor: | DNA (5'-D(*TP*TP*GP*AP*CP*TP*CP*TP*CP*TP*TP*AP*AP*GP*AP*GP*A P*GP*TP*CP*A)-3'), INTRON-ENCODED ENDONUCLEASE I-PPOI, ZINC ION | | Authors: | Jurica, B.L, Flick, K.E, Monnat Junior, R.J, Stoddard, M.S. | | Deposit date: | 1998-03-19 | | Release date: | 1998-06-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | DNA binding and cleavage by the nuclear intron-encoded homing endonuclease I-PpoI.
Nature, 394, 1998
|
|
1A43
 
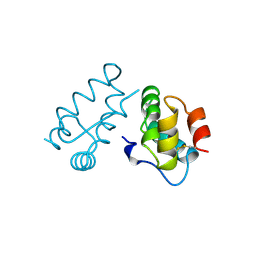 | | STRUCTURE OF THE HIV-1 CAPSID PROTEIN DIMERIZATION DOMAIN AT 2.6A RESOLUTION | | Descriptor: | HIV-1 CAPSID | | Authors: | Worthylake, D.K, Wang, H, Yoo, S, Sundquist, W.I, Hill, C.P. | | Deposit date: | 1998-02-10 | | Release date: | 1999-02-09 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structures of the HIV-1 capsid protein dimerization domain at 2.6 A resolution.
Acta Crystallogr.,Sect.D, 55, 1999
|
|
1A4Z
 
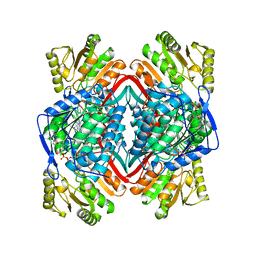 | |
1A5U
 
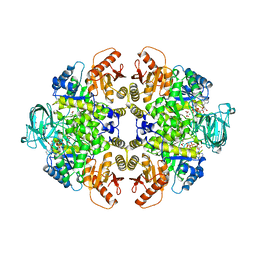 | | PYRUVATE KINASE COMPLEX WITH BIS MG-ATP-NA-OXALATE | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, OXALATE ION, ... | | Authors: | Larsen, T.M, Benning, M.M, Rayment, I, Reed, G.H. | | Deposit date: | 1998-02-18 | | Release date: | 1999-03-02 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structure of the bis(Mg2+)-ATP-oxalate complex of the rabbit muscle pyruvate kinase at 2.1 A resolution: ATP binding over a barrel.
Biochemistry, 37, 1998
|
|
1A81
 
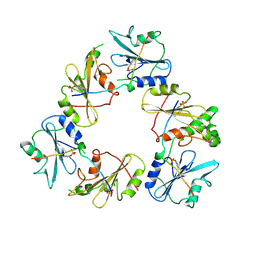 | |
1AFP
 
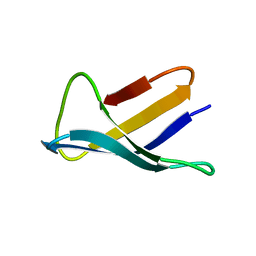 | | SOLUTION STRUCTURE OF THE ANTIFUNGAL PROTEIN FROM ASPERGILLUS GIGANTEUS. EVIDENCE FOR DISULPHIDE CONFIGURATIONAL ISOMERISM | | Descriptor: | ANTIFUNGAL PROTEIN FROM ASPERGILLUS GIGANTEUS | | Authors: | Campos-Olivas, R, Bruix, M, Santoro, J, Lacadena, J, Del Pozo, A.M, Gavilanes, J.G, Rico, M. | | Deposit date: | 1994-11-11 | | Release date: | 1995-02-07 | | Last modified: | 2024-04-10 | | Method: | SOLUTION NMR | | Cite: | NMR solution structure of the antifungal protein from Aspergillus giganteus: evidence for cysteine pairing isomerism.
Biochemistry, 34, 1995
|
|
1A73
 
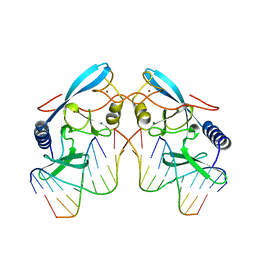 | | INTRON-ENCODED ENDONUCLEASE I-PPOI COMPLEXED WITH DNA | | Descriptor: | DNA (5'-D(*TP*TP*GP*AP*CP*TP*CP*TP*CP*TP*TP*AP*A)-3'), DNA (5'-D(P*GP*AP*GP*AP*GP*TP*CP*A)-3'), INTRON 3 (I-PPO) ENCODED ENDONUCLEASE, ... | | Authors: | Flick, K.E, Monnat Junior, R.J, Stoddard, B.L. | | Deposit date: | 1998-03-19 | | Release date: | 1998-10-14 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | DNA binding and cleavage by the nuclear intron-encoded homing endonuclease I-PpoI.
Nature, 394, 1998
|
|
1A8R
 
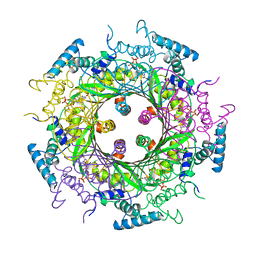 | | GTP CYCLOHYDROLASE I (H112S MUTANT) IN COMPLEX WITH GTP | | Descriptor: | GTP CYCLOHYDROLASE I, GUANOSINE-5'-TRIPHOSPHATE | | Authors: | Auerbach, G, Nar, H, Bracher, A, Bacher, A, Huber, R. | | Deposit date: | 1998-03-27 | | Release date: | 1999-05-11 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Biosynthesis of pteridines. Reaction mechanism of GTP cyclohydrolase I.
J.Mol.Biol., 326, 2003
|
|
1AMU
 
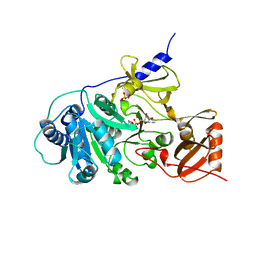 | | PHENYLALANINE ACTIVATING DOMAIN OF GRAMICIDIN SYNTHETASE 1 IN A COMPLEX WITH AMP AND PHENYLALANINE | | Descriptor: | ADENOSINE MONOPHOSPHATE, GRAMICIDIN SYNTHETASE 1, MAGNESIUM ION, ... | | Authors: | Conti, E, Stachelhaus, T, Marahiel, M.A, Brick, P. | | Deposit date: | 1997-06-18 | | Release date: | 1998-07-01 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural basis for the activation of phenylalanine in the non-ribosomal biosynthesis of gramicidin S.
EMBO J., 16, 1997
|
|
1AF2
 
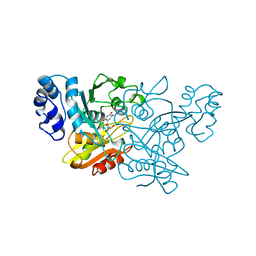 | | CRYSTAL STRUCTURE OF CYTIDINE DEAMINASE COMPLEXED WITH URIDINE | | Descriptor: | CYTIDINE DEAMINASE, URIDINE-5'-MONOPHOSPHATE, ZINC ION | | Authors: | Xiang, S, Carter, C.W. | | Deposit date: | 1997-03-20 | | Release date: | 1997-07-07 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The structure of the cytidine deaminase-product complex provides evidence for efficient proton transfer and ground-state destabilization.
Biochemistry, 36, 1997
|
|
1A4D
 
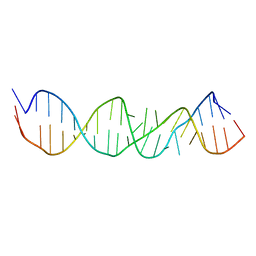 | | LOOP D/LOOP E ARM OF ESCHERICHIA COLI 5S RRNA, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | RNA (5'-R(*GP*GP*CP*CP*GP*AP*UP*GP*GP*UP*AP*GP*UP*GP*UP*GP*GP*GP*GP*UP*C)-3'), RNA (5'-R(P*UP*CP*CP*CP*CP*AP*UP*GP*CP*GP*AP*GP*AP*GP*UP*AP*GP*GP*CP*C)-3') | | Authors: | Dallas, A, Moore, P.B. | | Deposit date: | 1998-01-29 | | Release date: | 1998-04-29 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The loop E-loop D region of Escherichia coli 5S rRNA: the solution structure reveals an unusual loop that may be important for binding ribosomal proteins.
Structure, 5, 1997
|
|
1AEY
 
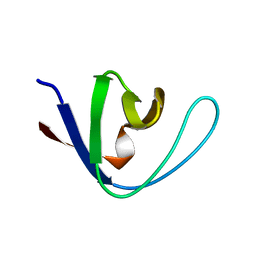 | |
1A3C
 
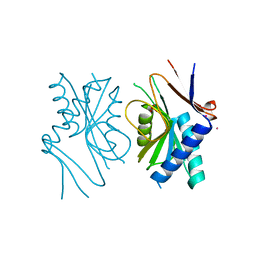 | | PYRR, THE BACILLUS SUBTILIS PYRIMIDINE BIOSYNTHETIC OPERON REPRESSOR, DIMERIC FORM | | Descriptor: | PYRIMIDINE OPERON REGULATORY PROTEIN PYRR, SAMARIUM (III) ION, SULFATE ION | | Authors: | Tomchick, D.R, Turner, R.J, Switzer, R.W, Smith, J.L. | | Deposit date: | 1998-01-20 | | Release date: | 1998-08-05 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Adaptation of an enzyme to regulatory function: structure of Bacillus subtilis PyrR, a pyr RNA-binding attenuation protein and uracil phosphoribosyltransferase.
Structure, 6, 1998
|
|
1CNY
 
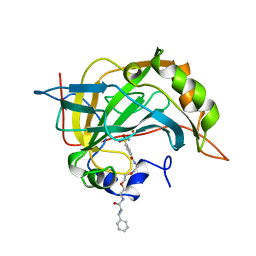 | |
1COL
 
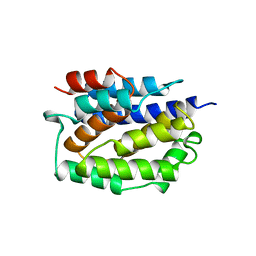 | | REFINED STRUCTURE OF THE PORE-FORMING DOMAIN OF COLICIN A AT 2.4 ANGSTROMS RESOLUTION | | Descriptor: | COLICIN A | | Authors: | Parker, M.W, Postma, J.P.M, Pattus, F, Tucker, A.D, Tsernoglou, D. | | Deposit date: | 1991-07-06 | | Release date: | 1992-07-15 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Refined structure of the pore-forming domain of colicin A at 2.4 A resolution.
J.Mol.Biol., 224, 1992
|
|
1CMK
 
 | | CRYSTAL STRUCTURES OF THE MYRISTYLATED CATALYTIC SUBUNIT OF CAMP-DEPENDENT PROTEIN KINASE REVEAL OPEN AND CLOSED CONFORMATIONS | | Descriptor: | IODIDE ION, MYRISTIC ACID, cAMP-DEPENDENT PROTEIN KINASE CATALYTIC SUBUNIT, ... | | Authors: | Zheng, J, Knighton, D.R, Xuong, N.-H, Taylor, S.S, Sowadski, J.M, Ten Eyck, L.F. | | Deposit date: | 1993-11-18 | | Release date: | 1994-05-31 | | Last modified: | 2024-06-05 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structures of the myristylated catalytic subunit of cAMP-dependent protein kinase reveal open and closed conformations.
Protein Sci., 2, 1993
|
|
