1O1E
 
 | | MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE | | Descriptor: | SKELETAL MUSCLE ACTIN, SKELETAL MUSCLE MYOSIN II, SKELETAL MUSCLE MYOSIN II ESSENTIAL LIGHT CHAIN, ... | | Authors: | Chen, L.F, Winkler, H, Reedy, M.K, Reedy, M.C, Taylor, K.A. | | Deposit date: | 2002-11-19 | | Release date: | 2002-12-04 | | Last modified: | 2023-12-27 | | Method: | ELECTRON MICROSCOPY (70 Å) | | Cite: | Molecular Modeling of Averaged Rigor Crossbridges from Tomograms of Insect Flight Muscle
J.Struct.Biol., 138, 2002
|
|
1O1F
 | | MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE | | Descriptor: | SKELETAL MUSCLE ACTIN, SKELETAL MUSCLE MYOSIN II, SKELETAL MUSCLE MYOSIN II ESSENTIAL LIGHT CHAIN, ... | | Authors: | Chen, L.F, Winkler, H, Reedy, M.K, Reedy, M.C, Taylor, K.A. | | Deposit date: | 2002-11-15 | | Release date: | 2002-12-04 | | Last modified: | 2023-12-27 | | Method: | ELECTRON MICROSCOPY (70 Å) | | Cite: | Molecular Modeling of Averaged Rigor Crossbridges from Tomograms of Insect Flight Muscle
J.Struct.Biol., 138, 2002
|
|
1O1G
 | | MOLECULAR MODELS OF AVERAGED RIGOR CROSSBRIDGES FROM TOMOGRAMS OF INSECT FLIGHT MUSCLE | | Descriptor: | SKELETAL MUSCLE ACTIN, SKELETAL MUSCLE MYOSIN II, SKELETAL MUSCLE MYOSIN II ESSENTIAL LIGHT CHAIN, ... | | Authors: | Chen, L.F, Winkler, H, Reedy, M.K, Reedy, M.C, Taylor, K.A. | | Deposit date: | 2002-11-19 | | Release date: | 2002-12-11 | | Last modified: | 2023-12-27 | | Method: | ELECTRON MICROSCOPY (70 Å) | | Cite: | Molecular modeling of averaged rigor crossbridges from tomograms of insect flight muscle.
J.Struct.Biol., 138, 2002
|
|
1O1H
 
 | | STRUCTURE OF GLUCOSE ISOMERASE DERIVATIZED WITH KR. | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CALCIUM ION, ... | | Authors: | Nowak, E, Panjikar, S, Tucker, P.A. | | Deposit date: | 2002-11-07 | | Release date: | 2002-11-27 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Structure of Glucose Isomerase Derivatized with Kr.
To be Published
|
|
1O1I
 
 | |
1O1J
 
 | |
1O1K
 
 | |
1O1L
 
 | |
1O1M
 | |
1O1N
 
 | |
1O1O
 
 | |
1O1P
 | |
1O1R
 
 | | Structure of FPT bound to GGPP | | Descriptor: | GERANYLGERANYL DIPHOSPHATE, Protein farnesyltransferase alpha subunit, Protein farnesyltransferase beta subunit, ... | | Authors: | Turek-Etienne, T.C, Strickland, C.L, Distefano, M.D. | | Deposit date: | 2003-01-14 | | Release date: | 2003-05-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Biochemical and Structural Studies with Prenyl Diphosphate Analogues Provide Insights
into Isoprenoid Recognition by Protein Farnesyl Transferase
Biochemistry, 42, 2003
|
|
1O1S
 
 | | Structure of FPT bound to isoprenoid analog 3b | | Descriptor: | (2E,6E)-8-[(3-BENZOYLBENZYL)OXY]-3,7-DIMETHYLOCTA-2,6-DIENYL TRIHYDROGEN DIPHOSPHATE, Protein farnesyltransferase alpha subunit, Protein farnesyltransferase beta subunit, ... | | Authors: | Turek-Etienne, T.C, Strickland, C.L, Distefano, M.D. | | Deposit date: | 2003-01-14 | | Release date: | 2003-05-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Biochemical and Structural Studies with Prenyl Diphosphate Analogues Provide Insights into
Isoprenoid Recognition by Protein Farnesyl Transferase
Biochemistry, 42, 2003
|
|
1O1T
 
 | | Structure of FPT bound to the CVIM-FPP product | | Descriptor: | MAGNESIUM ION, N-ACETYL-S-[(2E,6E)-3,7,11-TRIMETHYLDODECA-2,6,10-TRIENYL]-L-CYSTEINYL-D-VALYL-L-ISOLEUCYL-L-METHIONINE, Protein farnesyltransferase alpha subunit, ... | | Authors: | Turek-Etienne, T.C, Strickland, C.L, Distefano, M.D. | | Deposit date: | 2003-01-14 | | Release date: | 2003-05-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Biochemical and Structural Studies with Prenyl Diphosphate Analogues Provide Insights into
Isoprenoid Recognition by Protein Farnesyl Transferase
Biochemistry, 42, 2003
|
|
1O1U
 
 | | human ileal lipid-binding protein (ILBP) in free form | | Descriptor: | Gastrotropin | | Authors: | Kurz, M, Brachvogel, V, Matter, H, Stengelin, S, Thuering, H, Kramer, W. | | Deposit date: | 2003-02-10 | | Release date: | 2003-02-25 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Insights into the bile acid transportation system: the human ileal lipid-binding protein-cholyltaurine complex and its comparison with homologous structures.
Proteins, 50, 2003
|
|
1O1V
 
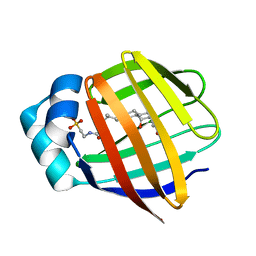 | | Human Ileal Lipid-Binding Protein (ILBP) in Complex with Cholyltaurine | | Descriptor: | Gastrotropin, TAUROCHOLIC ACID | | Authors: | Kurz, M, Brachvogel, V, Matter, H, Stengelin, S, Thuering, H, Kramer, W. | | Deposit date: | 2003-02-10 | | Release date: | 2003-02-18 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Insights into the bile acid transportation system: the human ileal lipid-binding protein-cholyltaurine complex and
its comparison with homologous structures.
Proteins, 50, 2003
|
|
1O1W
 
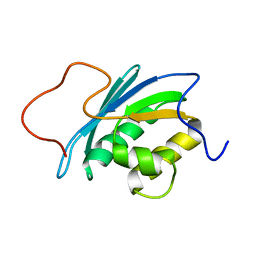 | | SOLUTION STRUCTURE OF THE RNASE H DOMAIN OF THE HIV-1 REVERSE TRANSCRIPTASE IN THE PRESENCE OF MAGNESIUM | | Descriptor: | RIBONUCLEASE H | | Authors: | Pari, K, Mueller, G.A, Derose, E.F, Kirby, T.W, London, R.E. | | Deposit date: | 2003-02-12 | | Release date: | 2003-02-18 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the Rnase H Domain of the HIV-1 Reverse Transcriptase in the Presence of Magnesium
Biochemistry, 42, 2003
|
|
1O1X
 
 | |
1O1Y
 
 | |
1O1Z
 
 | |
1O20
 
 | |
1O22
 
 | |
1O23
 
 | | CRYSTAL STRUCTURE OF LACTOSE SYNTHASE IN THE PRESENCE OF UDP-GLUCOSE | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, ALPHA-LACTALBUMIN, BETA-1,4-GALACTOSYLTRANSFERASE, ... | | Authors: | Ramakrishnan, B, Shah, P.S, Qasba, P.K. | | Deposit date: | 2003-01-29 | | Release date: | 2003-02-25 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.32 Å) | | Cite: | Alpha-Lactalbumin (La) Stimulates Milk Beta-1,4-Galactosyltransferase I (Beta 4Gal-T1) to Transfer Glucose from Udp-Glucose to N-Acetylglucosamine. Crystal Structure of Beta 4Gal-T1 X La Complex with Udp-Glc.
J.Biol.Chem., 276, 2001
|
|
1O24
 
 | | Crystal structure of Thymidylate Synthase Complementing Protein (TM0449) from Thermotoga maritima at 2.0 A resolution | | Descriptor: | Thymidylate synthase thyX | | Authors: | Mathews, I.I, Deacon, A.M, Canaves, J.M, McMullan, D, Lesley, S.A, Agarwalla, S, Kuhn, P, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2003-02-18 | | Release date: | 2003-06-24 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Functional Analysis of Substrate and Cofactor Complex Structures of a Thymidylate Synthase-Complementing Protein
Structure, 11, 2003
|
|
