6HO4
 
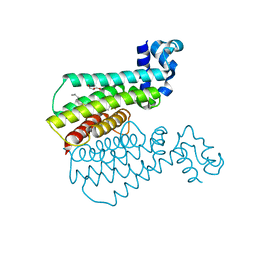 | | TRANSCRIPTIONAL REPRESSOR ETHR FROM MYCOBACTERIUM TUBERCULOSIS IN COMPLEX WITH BDM44693 | | Descriptor: | 4-[2-(4-methylpentyl)-1,3-thiazol-4-yl]-~{N}-[3,3,3-tris(fluoranyl)propyl]benzamide, GLYCEROL, HTH-type transcriptional regulator EthR | | Authors: | Wintjens, R, Wohlkonig, A. | | Deposit date: | 2018-09-17 | | Release date: | 2018-12-26 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | A comprehensive analysis of the protein-ligand interactions in crystal structures of Mycobacterium tuberculosis EthR.
Biochim Biophys Acta Proteins Proteom, 1867, 2018
|
|
8H58
 
 | | Crystal structure of YhaJ effector binding domain | | Descriptor: | HTH-type transcriptional regulator YhaJ, SODIUM ION | | Authors: | Kim, M, Ryu, S.E. | | Deposit date: | 2022-10-12 | | Release date: | 2023-10-25 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.639 Å) | | Cite: | Structural basis of transcription factor YhaJ for DNT detection.
Iscience, 26, 2023
|
|
2OBA
 
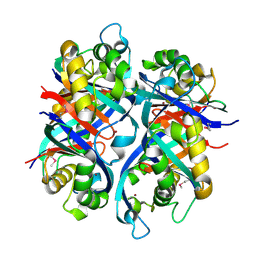 | | Pseudomonas aeruginosa 6-pyruvoyl tetrahydrobiopterin synthase | | Descriptor: | Probable 6-pyruvoyl tetrahydrobiopterin synthase, ZINC ION | | Authors: | McGrath, T.E, Kisselman, G, Battaile, K, Romanov, V, Wu-Brown, J, Guthrie, J, Virag, C, Mansoury, K, Edwards, A.M, Pai, E.F, Chirgadze, N.Y. | | Deposit date: | 2006-12-18 | | Release date: | 2007-01-30 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.33 Å) | | Cite: | Pseudomonas aeruginosa 6-pyruvoyl tetrahydrobiopterin synthase
TO BE PUBLISHED
|
|
3PBE
 
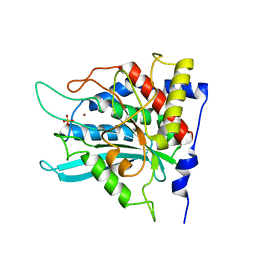 | | Crystal structure of the mutant W207F of human secretory glutaminyl cyclase | | Descriptor: | Glutaminyl-peptide cyclotransferase, SULFATE ION, ZINC ION | | Authors: | Huang, K.F, Liaw, S.S, Huang, W.L, Chia, C.Y, Lo, Y.C, Chen, Y.L, Wang, A.H.J. | | Deposit date: | 2010-10-20 | | Release date: | 2011-02-02 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Structures of human Golgi-resident glutaminyl cyclase and its complexes with inhibitors reveal a large loop movement upon inhibitor binding
J.Biol.Chem., 286, 2011
|
|
3PDC
 
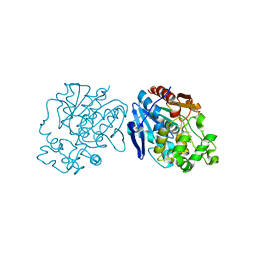 | |
8HR2
 
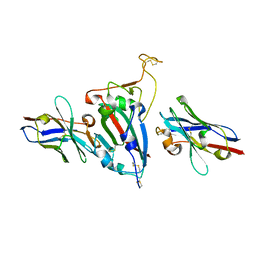 | | Ternary Crystal Complex Structure of RBD with NB1B5 and NB1C6 | | Descriptor: | NB1B5, NB1C6, Spike protein S1 | | Authors: | Sun, Z. | | Deposit date: | 2022-12-14 | | Release date: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Structure basis of two nanobodies neutralizing SARS-CoV-2 Omicron variant by targeting ultra-conservative epitopes.
J.Struct.Biol., 215, 2023
|
|
6IAM
 
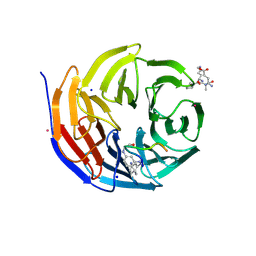 | |
3MVF
 
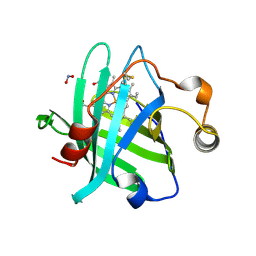 | |
6BZN
 
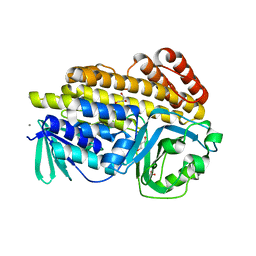 | |
5XJY
 
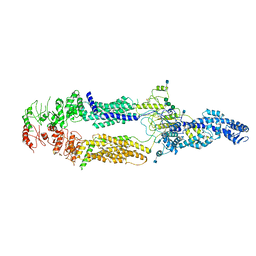 | | Cryo-EM structure of human ABCA1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ATP-binding cassette sub-family A member 1, ... | | Authors: | Qian, H.W, Yan, N, Gong, X. | | Deposit date: | 2017-05-04 | | Release date: | 2017-06-07 | | Last modified: | 2020-07-29 | | Method: | ELECTRON MICROSCOPY (4.1 Å) | | Cite: | Structure of the Human Lipid Exporter ABCA1.
Cell, 169, 2017
|
|
2KL2
 
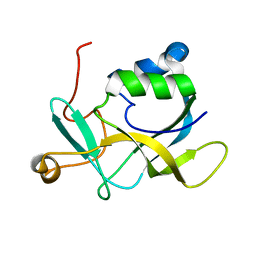 | | NMR solution structure of A2LD1 (gi:13879369) | | Descriptor: | AIG2-like domain-containing protein 1 | | Authors: | Pedrini, B, Serrano, P, Mohanty, B, Geralt, M, Herrmann, T, Wuthrich, K, Wilson, I, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2009-06-30 | | Release date: | 2009-07-14 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Comparison of NMR and crystal structures highlights conformational isomerism in protein active sites.
Acta Crystallogr.,Sect.F, 66, 2010
|
|
1PS2
 
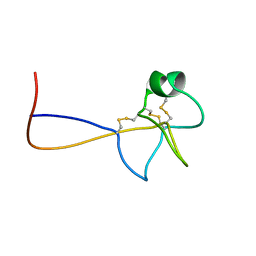 | | HIGH RESOLUTION NMR SOLUTION STRUCTURE OF HUMAN PS2, 19 STRUCTURES | | Descriptor: | PS2 | | Authors: | Williams, M.A, Polshakov, V.I, Gargaro, A.R, Feeney, J. | | Deposit date: | 1997-01-07 | | Release date: | 1997-07-07 | | Last modified: | 2021-11-03 | | Method: | SOLUTION NMR | | Cite: | High-resolution solution structure of human pNR-2/pS2: a single trefoil motif protein.
J.Mol.Biol., 267, 1997
|
|
6BGH
 
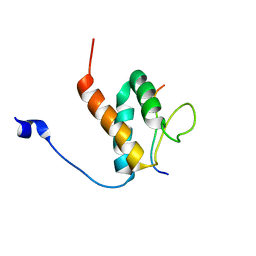 | |
7GTF
 
 | | PanDDA Analysis group deposition -- Crystal structure of PTP1B in complex with XST00000754b | | Descriptor: | 1,2-benzoxazol-3-ol, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Tyrosine-protein phosphatase non-receptor type 1 | | Authors: | Mehlman, T, Ginn, H.M, Keedy, D.A. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | An expanded view of ligandability in the allosteric enzyme PTP1B from computational reanalysis of large-scale crystallographic data.
Biorxiv, 2024
|
|
7GSQ
 
 | |
7GSK
 
 | |
7GTO
 
 | | PanDDA Analysis group deposition -- Crystal structure of PTP1B in complex with FMOOA000602a | | Descriptor: | (6aR,8R,12R,12aS)-5-methyl-5,6a,7,10,11,12a-hexahydro-6H,9H-pyrazolo[1',2':1,2]pyrazolo[4,3-c]quinolin-9-one, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Tyrosine-protein phosphatase non-receptor type 1 | | Authors: | Mehlman, T, Ginn, H.M, Keedy, D.A. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | An expanded view of ligandability in the allosteric enzyme PTP1B from computational reanalysis of large-scale crystallographic data.
Biorxiv, 2024
|
|
7GTD
 
 | |
7GSU
 
 | |
7GTR
 
 | |
7GSY
 
 | |
7GTS
 
 | |
7GSR
 
 | |
7GT5
 
 | | PanDDA Analysis group deposition -- Crystal structure of PTP1B in complex with FMOOA000529a | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Tyrosine-protein phosphatase non-receptor type 1, methyl [(3R,4S)-3-ethyl-4-hydroxy-1,1-dioxo-3,4-dihydro-1lambda~6~,2-benzothiazin-2(1H)-yl]acetate | | Authors: | Mehlman, T, Ginn, H.M, Keedy, D.A. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.61 Å) | | Cite: | An expanded view of ligandability in the allosteric enzyme PTP1B from computational reanalysis of large-scale crystallographic data.
Biorxiv, 2024
|
|
7GS7
 
 | | PanDDA Analysis group deposition -- Crystal structure of PTP1B in complex with FMOPL000621a | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 4-(1,2,3-thiadiazol-4-yl)phenyl ethylcarbamate, Tyrosine-protein phosphatase non-receptor type 1 | | Authors: | Mehlman, T, Ginn, H.M, Keedy, D.A. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | An expanded view of ligandability in the allosteric enzyme PTP1B from computational reanalysis of large-scale crystallographic data.
Biorxiv, 2024
|
|
