5WBX
 
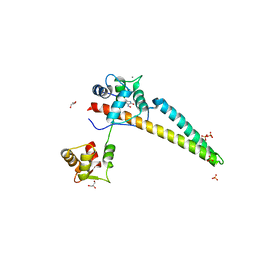 | |
5WC5
 
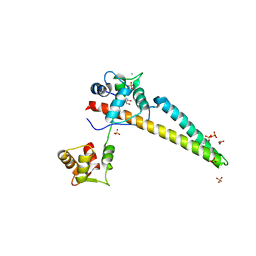 | |
2Q69
 
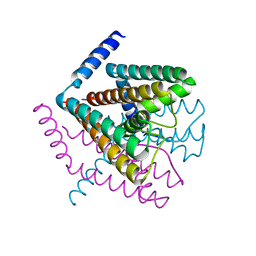 | | Crystal Structure of Nak channel D66N mutant | | Descriptor: | CALCIUM ION, Potassium channel protein, SODIUM ION | | Authors: | Alam, A, Shi, N, Jiang, Y. | | Deposit date: | 2007-06-04 | | Release date: | 2007-10-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural insight into Ca2+ specificity in tetrameric cation channels.
Proc.Natl.Acad.Sci.Usa, 104, 2007
|
|
1C2U
 
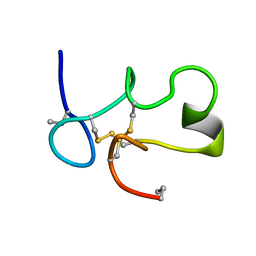 | | SOLUTION STRUCTURE OF [ABU3,35]SHK12-28,17-32 | | Descriptor: | SYNTHETIC PEPTIDE ANALOGUE OF SHK TOXIN | | Authors: | Pennington, M.W, Lanigan, M.D, Kalman, K, Manhir, V.M, Rauer, H, McVaugh, C.T, Behm, D, Donaldson, D, Chandy, K.G, Kem, W.R, Norton, R.S. | | Deposit date: | 1999-07-27 | | Release date: | 1999-11-10 | | Last modified: | 2021-11-03 | | Method: | SOLUTION NMR | | Cite: | Role of disulfide bonds in the structure and potassium channel blocking activity of ShK toxin.
Biochemistry, 38, 1999
|
|
2Q6A
 
 | | Crystal Structure of Nak channel D66E mutant | | Descriptor: | CALCIUM ION, Potassium channel protein, SODIUM ION | | Authors: | Alam, A, Shi, N, Jiang, Y. | | Deposit date: | 2007-06-04 | | Release date: | 2007-10-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural insight into Ca2+ specificity in tetrameric cation channels.
Proc.Natl.Acad.Sci.Usa, 104, 2007
|
|
2Q68
 
 | | Crystal Structure of Nak channel D66A, S70E double mutants | | Descriptor: | CALCIUM ION, Potassium channel protein, SODIUM ION | | Authors: | Alam, A, Shi, N, Jiang, Y. | | Deposit date: | 2007-06-04 | | Release date: | 2007-10-02 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural insight into Ca2+ specificity in tetrameric cation channels.
Proc.Natl.Acad.Sci.Usa, 104, 2007
|
|
5KUK
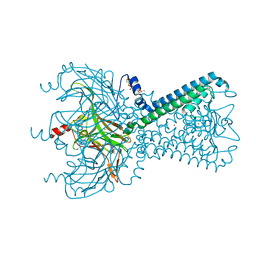 
 | | Crystal Structure of Inward Rectifier Kir2.2 K62W Mutant | | Descriptor: | ATP-sensitive inward rectifier potassium channel 12, DECYL-BETA-D-MALTOPYRANOSIDE, POTASSIUM ION | | Authors: | Lee, S.-J, Ren, F, Heyman, S, Yuan, P, Nichols, C.G. | | Deposit date: | 2016-07-13 | | Release date: | 2016-08-10 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural basis of control of inward rectifier Kir2 channel gating by bulk anionic phospholipids.
J.Gen.Physiol., 148, 2016
|
|
5KUM
 
 | | Crystal Structure of Inward Rectifier Kir2.2 K62W Mutant In Complex with PIP2 | | Descriptor: | ATP-sensitive inward rectifier potassium channel 12, DECYL-BETA-D-MALTOPYRANOSIDE, POTASSIUM ION, ... | | Authors: | Lee, S.-J, Ren, F, Heyman, S, Yuan, P, Nichols, C.G. | | Deposit date: | 2016-07-13 | | Release date: | 2016-08-10 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural basis of control of inward rectifier Kir2 channel gating by bulk anionic phospholipids.
J.Gen.Physiol., 148, 2016
|
|
2BMT
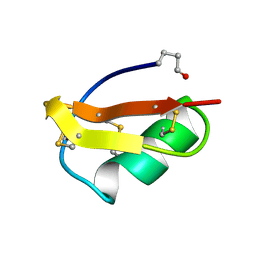 
 | | SCORPION TOXIN BMTX2 FROM BUTHUS MARTENSII KARSCH, NMR, 25 STRUCTURES | | Descriptor: | TOXIN BMTX2 | | Authors: | Blanc, E, Romi-Lebrun, R, Bornet, O, Nakajima, T, Darbon, H. | | Deposit date: | 1998-06-16 | | Release date: | 1999-01-13 | | Last modified: | 2024-10-23 | | Method: | SOLUTION NMR | | Cite: | Solution structure of two new toxins from the venom of the Chinese scorpion Buthus martensi Karsch blockers of potassium channels.
Biochemistry, 37, 1998
|
|
5NWI
 
 | | 14-3-3c in complex with CPP | | Descriptor: | 14-3-3 c-1 protein, ACETATE ION, Potassium channel KAT1 | | Authors: | Saponaro, A, Porro, A, Chaves-Sanjuan, A, Nardini, M, Thiel, G, Moroni, A. | | Deposit date: | 2017-05-06 | | Release date: | 2017-11-22 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Fusicoccin Activates KAT1 Channels by Stabilizing Their Interaction with 14-3-3 Proteins.
Plant Cell, 29, 2017
|
|
5NWJ
 
 | | 14-3-3c in complex with CPP7 | | Descriptor: | 14-3-3-like protein C, ACETATE ION, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Saponaro, A, Porro, A, Chaves-Sanjuan, A, Nardini, M, Thiel, G, Moroni, A. | | Deposit date: | 2017-05-06 | | Release date: | 2017-11-22 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.07 Å) | | Cite: | Fusicoccin Activates KAT1 Channels by Stabilizing Their Interaction with 14-3-3 Proteins.
Plant Cell, 29, 2017
|
|
4J9U
 
 | | Crystal Structure of the TrkH/TrkA potassium transport complex | | Descriptor: | HEXATANTALUM DODECABROMIDE, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, POTASSIUM ION, ... | | Authors: | Cao, Y, Jin, X, Huang, H, Levin, E.J, Zhou, M, New York Consortium on Membrane Protein Structure (NYCOMPS) | | Deposit date: | 2013-02-17 | | Release date: | 2013-04-03 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.8 Å) | | Cite: | Gating of the TrkH ion channel by its associated RCK protein TrkA.
Nature, 496, 2013
|
|
1BIG
 
 | | SCORPION TOXIN BMTX1 FROM BUTHUS MARTENSII KARSCH, NMR, 25 STRUCTURES | | Descriptor: | TOXIN BMTX1 | | Authors: | Blanc, E, Romi-Lebrun, R, Bornet, O, Nakajima, T, Darbon, H. | | Deposit date: | 1998-06-16 | | Release date: | 1999-01-13 | | Last modified: | 2019-12-25 | | Method: | SOLUTION NMR | | Cite: | Solution structure of two new toxins from the venom of the Chinese scorpion Buthus martensi Karsch blockers of potassium channels.
Biochemistry, 37, 1998
|
|
5EBL
 
 | | KcsA T75G in the Conductive State | | Descriptor: | Antibody Fab Fragment Light Chain, DIACYL GLYCEROL, NONAN-1-OL, ... | | Authors: | Matulef, K, Valiyaveetil, F.I. | | Deposit date: | 2015-10-19 | | Release date: | 2016-04-20 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Individual Ion Binding Sites in the K(+) Channel Play Distinct Roles in C-type Inactivation and in Recovery from Inactivation.
Structure, 24, 2016
|
|
5EC1
 
 | | KcsA with V76ester mutation | | Descriptor: | Antibody Fab Fragment Light Chain, DIACYL GLYCEROL, NONAN-1-OL, ... | | Authors: | Matulef, K, Valiyaveetil, F.I. | | Deposit date: | 2015-10-20 | | Release date: | 2016-04-20 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Individual Ion Binding Sites in the K(+) Channel Play Distinct Roles in C-type Inactivation and in Recovery from Inactivation.
Structure, 24, 2016
|
|
5EBM
 
 | | KcsA T75G mutant in the nonconductive state | | Descriptor: | Antibody Fab Fragment Light Chain, DIACYL GLYCEROL, NONAN-1-OL, ... | | Authors: | Matulef, K, Valiyaveetil, F.I. | | Deposit date: | 2015-10-19 | | Release date: | 2016-04-20 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Individual Ion Binding Sites in the K(+) Channel Play Distinct Roles in C-type Inactivation and in Recovery from Inactivation.
Structure, 24, 2016
|
|
5EBW
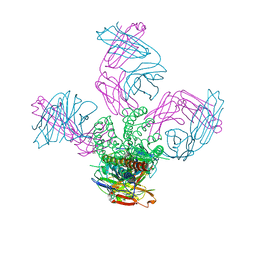 
 | | KcsA with G77ester mutation | | Descriptor: | Antibody Fab Fragment Light Chain, DIACYL GLYCEROL, NONAN-1-OL, ... | | Authors: | Matulef, K, Valiyaveetil, F.I. | | Deposit date: | 2015-10-19 | | Release date: | 2016-04-20 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Individual Ion Binding Sites in the K(+) Channel Play Distinct Roles in C-type Inactivation and in Recovery from Inactivation.
Structure, 24, 2016
|
|
5EC2
 
 | | KcsA with V76ester+G77dA mutations | | Descriptor: | Antibody Fab Fragment Light Chain, DIACYL GLYCEROL, NONAN-1-OL, ... | | Authors: | Matulef, K, Valiyaveetil, F.I. | | Deposit date: | 2015-10-20 | | Release date: | 2016-04-20 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Individual Ion Binding Sites in the K(+) Channel Play Distinct Roles in C-type Inactivation and in Recovery from Inactivation.
Structure, 24, 2016
|
|
3OUF
 
 | | Structure of a K+ selective NaK mutant | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, POTASSIUM ION, Potassium channel protein | | Authors: | Derebe, M.G, Sauer, D.B, Zeng, W, Alam, A, Shi, N, Jiang, Y. | | Deposit date: | 2010-09-14 | | Release date: | 2011-01-12 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Tuning the ion selectivity of tetrameric cation channels by changing the number of ion binding sites.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
2A0L
 
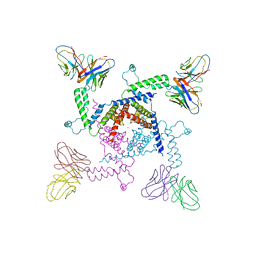 | | Crystal structure of KvAP-33H1 Fv complex | | Descriptor: | 33H1 Fv fragment, POTASSIUM ION, Voltage-gated potassium channel | | Authors: | Lee, S.Y, Lee, A, Chen, J, Mackinnon, R. | | Deposit date: | 2005-06-16 | | Release date: | 2005-11-01 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.9 Å) | | Cite: | Structure of the KvAP voltage-dependent K+ channel and its dependence on the lipid membrane.
Proc.Natl.Acad.Sci.Usa, 102, 2005
|
|
3BEH
 
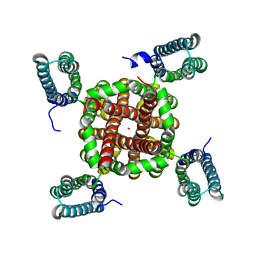 | |
1V7F
 
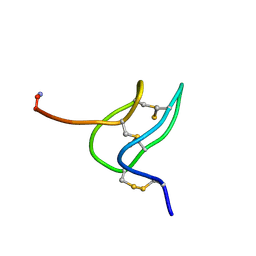 | | Solution structure of phrixotoxin 1 | | Descriptor: | Phrixotoxin 1 | | Authors: | Chagot, B, Escoubas, P, Villegas, E, Bernard, C, Ferrat, G, Corzo, G, Lazdunski, M, Darbon, H. | | Deposit date: | 2003-12-16 | | Release date: | 2004-11-23 | | Last modified: | 2024-10-30 | | Method: | SOLUTION NMR | | Cite: | Solution structure of Phrixotoxin 1, a specific peptide inhibitor of Kv4 potassium channels from the venom of the theraphosid spider Phrixotrichus auratus
Protein Sci., 13, 2004
|
|
1S6C
 
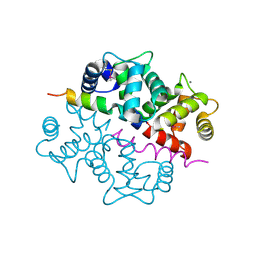 | | Crystal structure of the complex between KChIP1 and Kv4.2 N1-30 | | Descriptor: | CALCIUM ION, Kv4 potassium channel-interacting protein KChIP1b, Potassium voltage-gated channel subfamily D member 2 | | Authors: | Zhou, W, Qian, Y, Kunjilwar, K, Pfaffinger, P.J, Choe, S. | | Deposit date: | 2004-01-23 | | Release date: | 2004-02-24 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural insights into the functional interaction of KChIP1 with Shal-type K(+) channels.
Neuron, 41, 2004
|
|
3LDC
 
 | | High resolution open MthK pore structure crystallized in 100 mM K+ | | Descriptor: | Calcium-gated potassium channel mthK, POTASSIUM ION | | Authors: | Ye, S. | | Deposit date: | 2010-01-12 | | Release date: | 2010-08-11 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Novel insights into K(+) selectivity from high-resolution structures of an open K(+) channel pore.
Nat.Struct.Mol.Biol., 17, 2010
|
|
3LDD
 
 | |
