2JZ4
 
 | | Putative 32 kDa myrosinase binding protein At3g16450.1 from Arabidopsis thaliana | | Descriptor: | Jasmonate inducible protein isolog | | Authors: | Takeda, N, Sugimori, N, Torizawa, T, Terauchi, T, Ono, A.M, Yagi, H, Yamaguchi, Y, Kato, K, Ikeya, T, Guntert, P, Aceti, D.J, Markley, J.L, Kainosho, M, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2007-12-28 | | Release date: | 2008-02-19 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Structure of the putative 32 kDa myrosinase-binding protein from Arabidopsis (At3g16450.1) determined by SAIL-NMR.
Febs J., 275, 2008
|
|
2JZ5
 
 | | NMR solution structure of protein VPA0419 from Vibrio parahaemolyticus. Northeast Structural Genomics target VpR68 | | Descriptor: | Uncharacterized protein VPA0419 | | Authors: | Singarapu, K.K, Sukumaran, D.K, Eletski, A, Parish, D, Zhao, L, Jiang, M, Maglaqui, M, Xiao, R, Liu, J, Baran, M.C, Swapna, G, Acton, T, Rost, B, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2007-12-28 | | Release date: | 2008-02-19 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structures of proteins VPA0419 from Vibrio parahaemolyticus and yiiS from Shigella flexneri provide structural coverage for protein domain family PFAM 04175.
Proteins, 78, 2010
|
|
2JZ6
 
 | | Solution structure of 50S ribosomal protein L28 from Thermotoga maritima. Northeast Structural Genomics Consortium target VR97 | | Descriptor: | 50S ribosomal protein L28 | | Authors: | Wu, Y, Singarapu, K.K, Guido, V, Yee, A, Sukumaran, D, Arrowsmith, C.H, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2007-12-28 | | Release date: | 2008-01-29 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution structure of 50S ribosomal protein L28 from Thermotoga maritima. Northeast Structural Genomics Consortium target VR97.
To be Published
|
|
2JZ7
 
 | |
2JZ8
 
 | | Solution NMR structure of BH09830 from Bartonella henselae modeled with one Zn+2 bound. Northeast Structural Genomics Consortium target BnR55 | | Descriptor: | Uncharacterized protein BH09830, ZINC ION | | Authors: | Ding, K, Cort, J.R, Wang, D, Janjua, H, Owens, L, Xiao, R, Liu, J, Baran, M.C, Swapna, G.V.T, Acton, T.B, Rost, B, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2007-12-30 | | Release date: | 2008-01-15 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of BH09830 from Bartonella henselae modeled with one Zn+2 bound.
To be Published
|
|
2JZA
 
 | | Solution NMR structure of nitrite reductase [NAD(P)H] small subunit from Erwinia carotovora. Northeast Structural Genomics Consortium target EwR120 | | Descriptor: | Nitrite reductase [NAD(P)H] small subunit | | Authors: | Sathyamoorthy, B, Eletsky, A, Wang, D, Stokes, K, Owens, L, Xiao, R, Liu, J, Baran, M.C, Swapna, G.V.T, Acton, T.B, Rost, B, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2007-12-31 | | Release date: | 2008-01-22 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of nitrite reductase [NAD(P)H] small subunit from Erwinia carotovora.
To be Published
|
|
2JZB
 
 | |
2JZC
 
 | | NMR solution structure of ALG13: The sugar donor subunit of a yeast N-acetylglucosamine transferase. Northeast Structural Genomics Consortium target YG1 | | Descriptor: | UDP-N-acetylglucosamine transferase subunit ALG13 | | Authors: | Wang, X, Weldeghorghis, T, Zhang, G, Imepriali, B, Montelione, G.T, Prestegard, J.H, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-01-04 | | Release date: | 2008-02-19 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution structure of Alg13: the sugar donor subunit of a yeast N-acetylglucosamine transferase.
Structure, 16, 2008
|
|
2JZD
 
 | | NMR structure of the domain 527-651 of the SARS-CoV nonstructural protein nsp3 | | Descriptor: | Replicase polyprotein 1ab | | Authors: | Chatterjee, A, Johnson, M.A, Serrano, P, Pedrini, B, Joseph, J, Saikatendu, K, Neuman, B, Stevens, R.C, Wilson, I.A, Buchmeier, M.J, Kuhn, P, Wuthrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2008-01-04 | | Release date: | 2008-02-05 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Nuclear magnetic resonance structure shows that the severe acute respiratory syndrome coronavirus-unique domain contains a macrodomain fold.
J.Virol., 83, 2009
|
|
2JZE
 
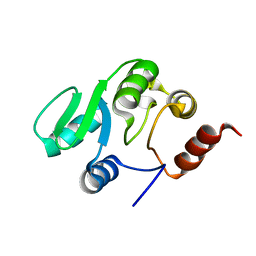 | | NMR structure of the domain 527-651 of the SARS-CoV nonstructural protein nsp3, single conformer closest to the mean coordinates of an ensemble of twenty energy minimized conformers | | Descriptor: | Replicase polyprotein 1ab | | Authors: | Chatterjee, A, Johnson, M.A, Serrano, P, Pedrini, B, Joseph, J, Saikatendu, K, Neuman, B, Stevens, R.C, Wilson, I.A, Buchmeier, M.J, Kuhn, P, Wuthrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2008-01-04 | | Release date: | 2008-02-05 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Nuclear magnetic resonance structure shows that the severe acute respiratory syndrome coronavirus-unique domain contains a macrodomain fold.
J.Virol., 83, 2009
|
|
2JZF
 
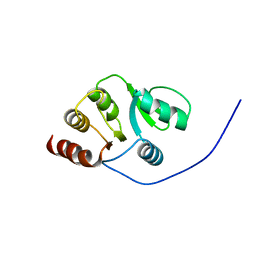 | | NMR Conformer closest to the mean coordinates of the domain 513-651 of the SARS-CoV nonstructural protein nsp3 | | Descriptor: | Replicase polyprotein 1ab | | Authors: | Chatterjee, A, Johnson, M.A, Serrano, P, Pedrini, B, Joseph, J, Saikatendu, K, Neuman, B, Stevens, R.C, Wilson, I.A, Buchmeier, M.J, Kuhn, P, Wuthrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2008-01-04 | | Release date: | 2008-02-05 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Nuclear magnetic resonance structure shows that the severe acute respiratory syndrome coronavirus-unique domain contains a macrodomain fold.
J.Virol., 83, 2009
|
|
2JZH
 
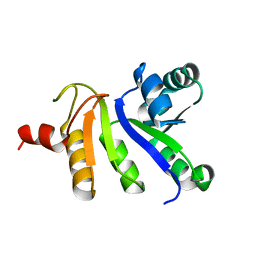 | | structure of IIB domain of the mannose transporter of E. coli | | Descriptor: | PTS system mannose-specific EIIAB component | | Authors: | Komlosh, M, Williams Jr, D.C. | | Deposit date: | 2008-01-08 | | Release date: | 2008-02-19 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structures of Productive and Non-productive Complexes between the A and B Domains of the Cytoplasmic Subunit of the Mannose Transporter of the Escherichia coli Phosphotransferase System.
J.Biol.Chem., 283, 2008
|
|
2JZI
 
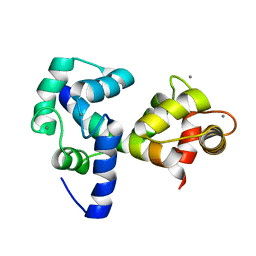 | |
2JZJ
 | | Structure of CrCVNH (C. richardii CVNH) | | Descriptor: | Cyanovirin-N homolog | | Authors: | Koharudin, L.M.I, Viscomi, A.R, Jee, J, Ottonello, S, Gronenborn, A.M. | | Deposit date: | 2008-01-09 | | Release date: | 2008-03-25 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | The evolutionarily conserved family of cyanovirin-N homologs: structures and carbohydrate specificity.
Structure, 16, 2008
|
|
2JZK
 | | Structure of TbCVNH (T. BORCHII CVNH) | | Descriptor: | Cyanovirin-N homolog | | Authors: | Koharudin, L.M.I, Viscomi, A.R, Jee, J, Ottonello, S, Gronenborn, A.M. | | Deposit date: | 2008-01-09 | | Release date: | 2008-03-25 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | The evolutionarily conserved family of cyanovirin-N homologs: structures and carbohydrate specificity.
Structure, 16, 2008
|
|
2JZL
 | | Structure of NcCVNH (N. CRASSA CVNH) | | Descriptor: | Cyanovirin-N homolog | | Authors: | Koharudin, L.M.I, Viscomi, A.R, Jee, J, Ottonello, S, Gronenborn, A.M. | | Deposit date: | 2008-01-09 | | Release date: | 2008-03-25 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | The evolutionarily conserved family of cyanovirin-N homologs: structures and carbohydrate specificity.
Structure, 16, 2008
|
|
2JZM
 
 | |
2JZN
 
 | |
2JZO
 
 | |
2JZP
 | | NMR solution structure of Kx5Q ProtL mutant | | Descriptor: | Protein L | | Authors: | Lopez-Mendez, N, Tadeo, X, Castano, D, Pons, M, Millet Aguilar-Galindo, O. | | Deposit date: | 2008-01-11 | | Release date: | 2009-01-13 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Protein stabilization and the hofmeister effect: the role of hydrophobic solvation
Biophys.J., 97, 2009
|
|
2JZQ
 
 | | Design of an Active Ultra-Stable Single-Chain Insulin Analog 20 Structures | | Descriptor: | Insulin | | Authors: | Hua, Q.X, Nakarawa, S, Jia, W.H, Huang, K, Philips, N.F, Hu, S.Q, Weiss, M.A. | | Deposit date: | 2008-01-11 | | Release date: | 2008-02-26 | | Last modified: | 2021-10-20 | | Method: | SOLUTION NMR | | Cite: | Design of an active ultrastable single-chain insulin analog: synthesis, structure, and therapeutic implications.
J.Biol.Chem., 283, 2008
|
|
2JZR
 
 | | Solution structure of the oxidized form (Cys67-Cys70) of the N-terminal domain of PilB from N. meningitidis. | | Descriptor: | Peptide methionine sulfoxide reductase msrA/msrB | | Authors: | Quinternet, M, Tsan, P, Neiers, F, Beaufils, C, Boschi-Muller, S, Averlant-Petit, M, Branlant, G, Cung, M. | | Deposit date: | 2008-01-15 | | Release date: | 2008-07-29 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Solution structure and dynamics of the reduced and oxidized forms of the N-terminal domain of PilB from Neisseria meningitidis.
Biochemistry, 47, 2008
|
|
2JZS
 
 | | Solution structure of the reduced form of the N-terminal domain of PilB from N. meningitidis. | | Descriptor: | Peptide methionine sulfoxide reductase msrA/msrB | | Authors: | Quinternet, M, Tsan, P, Neiers, F, Beaufils, C, Boschi-Muller, S, Averlant-Petit, M, Branlant, G, Cung, M. | | Deposit date: | 2008-01-15 | | Release date: | 2008-07-29 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure and dynamics of the reduced and oxidized forms of the N-terminal domain of PilB from Neisseria meningitidis.
Biochemistry, 47, 2008
|
|
2JZT
 
 | | Solution NMR structure of Q8ZP25_SALTY from Salmonella typhimurium. Northeast Structural Genomics Consortium target StR70 | | Descriptor: | Putative thiol-disulfide isomerase and thioredoxin | | Authors: | Parish, D, Liu, G, Shen, Y, Ho, C, Cunningham, K, Xiao, R, Swapna, G.V.T, Acton, T, Bansal, S, Prestegard, J.H, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-01-16 | | Release date: | 2008-02-19 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Protein chaperones Q8ZP25_SALTY from Salmonella typhimurium and HYAE_ECOLI from Escherichia coli exhibit thioredoxin-like structures despite lack of canonical thioredoxin active site sequence motif.
J.STRUCT.FUNCT.GENOM., 9, 2008
|
|
2JZV
 
 | | Solution structure of S. aureus PrsA-PPIase | | Descriptor: | Foldase protein prsA | | Authors: | Seppala, R, Tossavainen, H, Heikkinen, S, Koskela, H, Kontinen, V, Permi, P. | | Deposit date: | 2008-01-21 | | Release date: | 2009-01-20 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the parvulin-type PPIase domain of Staphylococcus aureus PrsA - Implications for the catalytic mechanism of parvulins.
Bmc Struct.Biol., 9, 2009
|
|
