1S4A
 
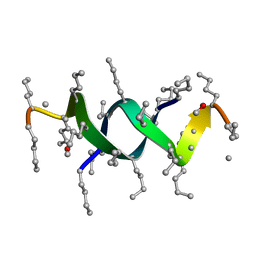 | |
1S1O
 
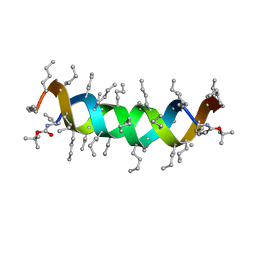 | |
1OKJ
 
 | |
1RIK
 
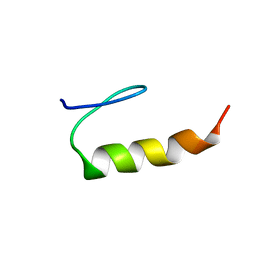 | | E6-binding zinc finger (E6apc1) | | Descriptor: | E6apc1 peptide | | Authors: | Liu, Y, Liu, Z, Androphy, E, Chen, J, Baleja, J.D. | | Deposit date: | 2003-11-17 | | Release date: | 2004-08-03 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Design and characterization of helical peptides that inhibit the E6 protein of papillomavirus.
Biochemistry, 43, 2004
|
|
1RVS
 
 | | STRUCTURE OF TRANSTHYRETIN IN AMYLOID FIBRILS DETERMINED BY SOLID-STATE MAGIC ANGLE SPINNING NMR | | Descriptor: | Transthyretin | | Authors: | Jaroniec, C.P, Macphee, C.E, Bajaj, V.S, Mcmahon, M.T, Dobson, C.M, Griffin, R.G. | | Deposit date: | 2003-12-14 | | Release date: | 2004-01-20 | | Last modified: | 2024-05-01 | | Method: | SOLID-STATE NMR | | Cite: | High-resolution molecular structure of a peptide in an amyloid fibril determined by magic angle spinning NMR spectroscopy
Proc.Natl.Acad.Sci.USA, 101, 2004
|
|
1SN9
 
 | |
1SNA
 
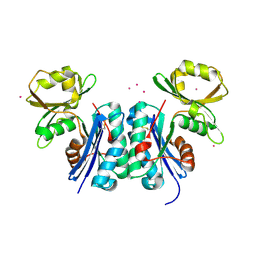
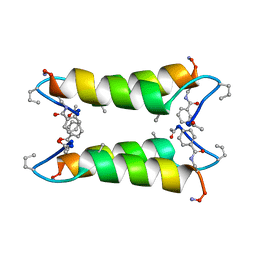
 | | An Oligomeric Domain-Swapped Beta-Beta-Alpha Mini-Protein | | Descriptor: | ISOPROPYL ALCOHOL, tetrameric beta-beta-alpha mini-protein | | Authors: | Ali, M.H, Peisach, E, Allen, K.N, Imperiali, B. | | Deposit date: | 2004-03-10 | | Release date: | 2004-08-17 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | X-ray structure analysis of a designed oligomeric miniprotein reveals a discrete quaternary architecture.
Proc.Natl.Acad.Sci.Usa, 101, 2004
|
|
1SNE
 
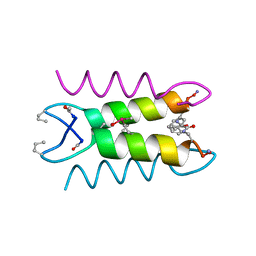
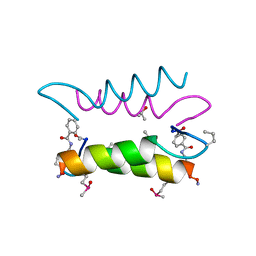 | | An Oligomeric Domain-Swapped Beta-Beta-Alpha Mini-Protein | | Descriptor: | ISOPROPYL ALCOHOL, tetrameric beta-beta-alpha mini-protein | | Authors: | Ali, M.H, Peisach, E, Allen, K.N, Imperiali, B. | | Deposit date: | 2004-03-10 | | Release date: | 2004-08-17 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | X-ray structure analysis of a designed oligomeric miniprotein reveals a discrete quaternary architecture.
Proc.Natl.Acad.Sci.Usa, 101, 2004
|
|
1SVX
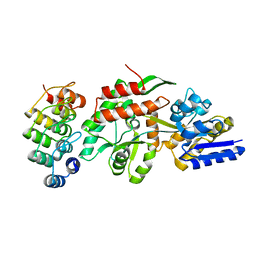 
 | | Crystal structure of a designed selected Ankyrin Repeat protein in complex with the Maltose Binding Protein | | Descriptor: | Ankyrin Repeat Protein off7, Maltose-binding periplasmic protein | | Authors: | Binz, H.K, Amstutz, P, Kohl, A, Stumpp, M.T, Briand, C, Forrer, P, Gruetter, M.G, Plueckthun, A. | | Deposit date: | 2004-03-30 | | Release date: | 2004-05-25 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.24 Å) | | Cite: | High-affinity binders selected from designed ankyrin repeat protein libraries
NAT.BIOTECHNOL., 22, 2004
|
|
6D37
 
 | |
6CFA
 
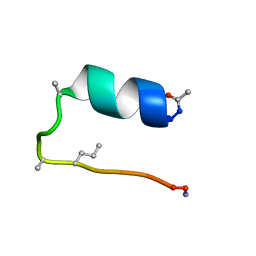
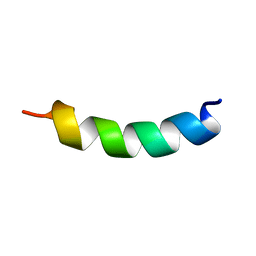 | | peptide PaAMP1R3 | | Descriptor: | peptide PaAMP1R3 | | Authors: | Alves, E.S.F, Liao, L.M. | | Deposit date: | 2018-02-14 | | Release date: | 2019-03-06 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Synthetic peptide PaAMP1R3
To be Published
|
|
7U4P
 
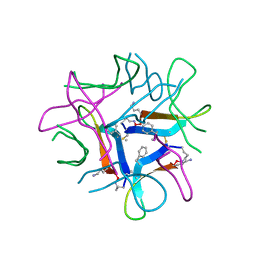 | |
6DMA
 
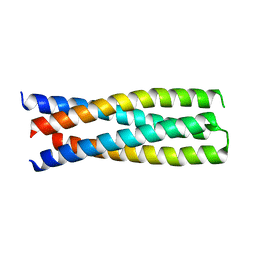 | | DHD15_closed | | Descriptor: | DHD15_closed_A, DHD15_closed_B | | Authors: | Bick, M.J, Chen, Z, Baker, D. | | Deposit date: | 2018-06-04 | | Release date: | 2018-12-19 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (3.363 Å) | | Cite: | Programmable design of orthogonal protein heterodimers.
Nature, 565, 2019
|
|
6DR4
 
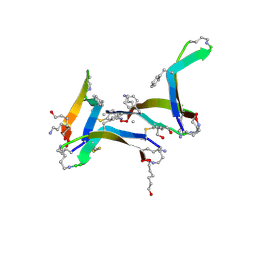 | |
6DKM
 
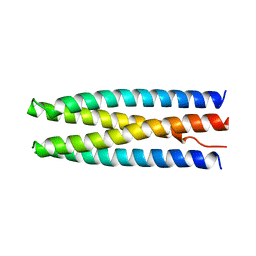 | | DHD131 | | Descriptor: | DHD131_A, DHD131_B | | Authors: | Bick, M.J, Chen, Z, Baker, D. | | Deposit date: | 2018-05-29 | | Release date: | 2018-12-19 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | Programmable design of orthogonal protein heterodimers.
Nature, 565, 2019
|
|
6D6X
 
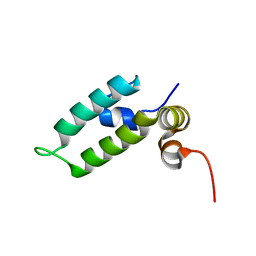 | | HSP40 co-chaperone Sis1 J-domain | | Descriptor: | Type II HSP40 co-chaperone | | Authors: | Pinheiro, G.M.S, Amorim, G.C, Iqbal, A, Ramos, C.H.I, Almeida, F.C.L. | | Deposit date: | 2018-04-23 | | Release date: | 2019-05-01 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Solution NMR investigation on the structure and function of the isolated J-domain from Sis1: Evidence of transient inter-domain interactions in the full-length protein.
Arch.Biochem.Biophys., 669, 2019
|
|
6DM9
 | | DHD15_extended | | Descriptor: | DHD15_extended_A, DHD15_extended_B, SULFATE ION | | Authors: | Bick, M.J, Chen, Z, Baker, D. | | Deposit date: | 2018-06-04 | | Release date: | 2018-12-19 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Programmable design of orthogonal protein heterodimers.
Nature, 565, 2019
|
|
6E5J
 
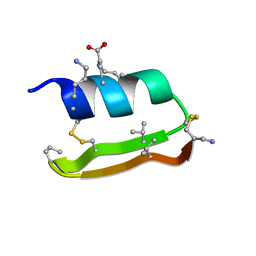 | | Heterogeneous-Backbone Mimics of a Designed Disulfide-Rich Protein: Aib turn, beta3 helix, N-methyl hairpin | | Descriptor: | Designed peptide NC_HEE_D1: Aib turn, beta3 helix, N-methyl hairpin mutant | | Authors: | Cabalteja, C.C, Mihalko, D.S, Horne, W.S. | | Deposit date: | 2018-07-20 | | Release date: | 2018-11-21 | | Last modified: | 2023-11-15 | | Method: | SOLUTION NMR | | Cite: | Heterogeneous-Backbone Foldamer Mimics of a Computationally Designed, Disulfide-Rich Miniprotein.
Chembiochem, 20, 2019
|
|
6E5I
 | |
6EE5
 
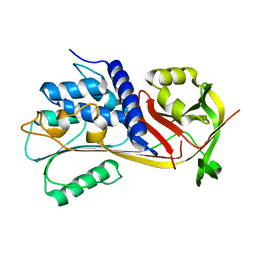 | |
6DR6
 
 | |
6DLC
 | |
6F5E
 
 | | Crystal structure of DARPin-DARPin rigid fusion, variant DD_D12_10_47 in complex JNK1a1 and JIP1 peptide | | Descriptor: | C-Jun-amino-terminal kinase-interacting protein 1, DD_D12_10_47, Mitogen-activated protein kinase 8 | | Authors: | Wu, Y, Mittl, P.R, Honegger, A, Batyuk, A, Plueckthun, A. | | Deposit date: | 2017-12-01 | | Release date: | 2017-12-13 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of DARPin-DARPin rigid fusion, variant DD_D12_10_47 in complex JNK1a1 and JIP1 peptide
To be published
|
|
7URJ
 | |
6DR5
 
 | |
