6ACS
 
 | | poly-cis-prenyltransferase | | Descriptor: | CITRIC ACID, Ditrans,polycis-undecaprenyl-diphosphate synthase ((2E,6E)-farnesyl-diphosphate specific), GLYCEROL, ... | | Authors: | Ko, T.-P, Chen, Y. | | Deposit date: | 2018-07-27 | | Release date: | 2018-12-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Structure of undecaprenyl pyrophosphate synthase from Acinetobacter baumannii
Acta Crystallogr F Struct Biol Commun, 74, 2018
|
|
7CM8
 
 | | High resolution crystal structure of M92A mutant of O-acetyl-L-serine sulfhydrylase from Haemophilus influenzae | | Descriptor: | Cysteine synthase, GLYCEROL, SODIUM ION | | Authors: | Kaushik, A, Rahisuddin, R, Saini, N, Kumaran, S. | | Deposit date: | 2020-07-25 | | Release date: | 2020-08-19 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Molecular mechanism of selective substrate engagement and inhibitor disengagement of cysteine synthase.
J.Biol.Chem., 296, 2020
|
|
7NYS
 
 | | monomeric acetyl-CoA synthase in closed conformation with carbon monoxide bound to the Ni proximal of cluster A | | Descriptor: | $l^{3}-oxidanylidynemethylnickel, 2-[3,8,8,12,12-pentakis(2-hydroxy-2-oxoethyl)-2,7,11-tris(oxidanylidene)-1,4,6,9,10,13-hexaoxa-5$l^{6}-titanaspiro[4.4^{5}.4^{5}]tridecan-3-yl]ethanoic acid, CHLORIDE ION, ... | | Authors: | Kreibich, J, Jeoung, J.H, Dobbek, H. | | Deposit date: | 2021-03-23 | | Release date: | 2022-04-13 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Ligand binding at the Ni,Ni-[4Fe-4S] cluster of acetyl-CoA synthase
To Be Published
|
|
7NYP
 
 | | monomeric acetyl-CoA synthase in closed conformation | | Descriptor: | 2-[3,8,8,12,12-pentakis(2-hydroxy-2-oxoethyl)-2,7,11-tris(oxidanylidene)-1,4,6,9,10,13-hexaoxa-5$l^{6}-titanaspiro[4.4^{5}.4^{5}]tridecan-3-yl]ethanoic acid, CO-methylating acetyl-CoA synthase, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Kreibich, J, Jeoung, J.H, Dobbek, H. | | Deposit date: | 2021-03-23 | | Release date: | 2022-04-13 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Ligand binding at the Ni,Ni-[4Fe-4S] cluster of acetyl-CoA synthase
To Be Published
|
|
7NZ5
 
 | | monomeric acetyl-CoA synthase with Zn at the proximal position of cluster A | | Descriptor: | (R,R)-2,3-BUTANEDIOL, 2-[3,8,8,12,12-pentakis(2-hydroxy-2-oxoethyl)-2,7,11-tris(oxidanylidene)-1,4,6,9,10,13-hexaoxa-5$l^{6}-titanaspiro[4.4^{5}.4^{5}]tridecan-3-yl]ethanoic acid, CITRIC ACID, ... | | Authors: | Kreibich, J, Jeoung, J.H, Dobbek, H. | | Deposit date: | 2021-03-23 | | Release date: | 2022-04-13 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Ligand binding at the Ni,Ni-[4Fe-4S] cluster of acetyl-CoA synthase
To Be Published
|
|
7C35
 
 | |
1Q8J
 
 | | Cobalamin-dependent methionine synthase (1-566) from Thermotoga maritima (Cd2+, Hcy, methyltetrahydrofolate complex) | | Descriptor: | 2-AMINO-4-MERCAPTO-BUTYRIC ACID, 5-METHYL-5,6,7,8-TETRAHYDROFOLIC ACID, 5-methyltetrahydrofolate S-homocysteine methyltransferase, ... | | Authors: | Evans, J.C, Huddler, D.P, Hilgers, M.T, Romanchuk, G, Matthews, R.G, Ludwig, M.L. | | Deposit date: | 2003-08-21 | | Release date: | 2004-03-23 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structures of the N-terminal modules imply large domain motions during catalysis by methionine synthase.
Proc.Natl.Acad.Sci.Usa, 101, 2004
|
|
1Q7M
 
 | | Cobalamin-dependent methionine synthase (MetH) from Thermotoga maritima (Oxidized, Monoclinic) | | Descriptor: | 5-methyltetrahydrofolate S-homocysteine methyltransferase | | Authors: | Evans, J.C, Huddler, D.P, Hilgers, M.T, Romanchuk, G, Matthews, R.G, Ludwig, M.L. | | Deposit date: | 2003-08-19 | | Release date: | 2004-03-23 | | Last modified: | 2017-10-11 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structures of the N-terminal modules imply large domain motions during catalysis by methionine synthase.
Proc.Natl.Acad.Sci.Usa, 101, 2004
|
|
1Q85
 
 | | Cobalamin-dependent methionine synthase (1-566) from Thermotoga maritima (Cd2+ complex, Se-Met) | | Descriptor: | 5-methyltetrahydrofolate S-homocysteine methyltransferase, CADMIUM ION | | Authors: | Evans, J.C, Huddler, D.P, Hilgers, M.T, Romanchuk, G, Matthews, R.G, Ludwig, M.L. | | Deposit date: | 2003-08-20 | | Release date: | 2004-03-23 | | Last modified: | 2017-10-11 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structures of the N-terminal modules imply large domain motions during catalysis by methionine synthase.
Proc.Natl.Acad.Sci.Usa, 101, 2004
|
|
1Q7Z
 
 | | Cobalamin-dependent methionine synthase (1-566) from Thermotoga maritima (Cd2+ complex) | | Descriptor: | 5-methyltetrahydrofolate S-homocysteine methyltransferase, CADMIUM ION | | Authors: | Evans, J.C, Huddler, D.P, Hilgers, M.T, Romanchuk, G, Matthews, R.G, Ludwig, M.L. | | Deposit date: | 2003-08-20 | | Release date: | 2004-03-23 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structures of the N-terminal modules imply large domain motions during catalysis by methionine synthase.
Proc.Natl.Acad.Sci.Usa, 101, 2004
|
|
1Q7Q
 
 | | Cobalamin-dependent methionine synthase (1-566) from T. maritima (Oxidized, Orthorhombic) | | Descriptor: | 5-methyltetrahydrofolate S-homocysteine methyltransferase | | Authors: | Evans, J.C, Huddler, D.P, Hilgers, M.T, Romanchuk, G, Matthews, R.G, Ludwig, M.L. | | Deposit date: | 2003-08-19 | | Release date: | 2004-03-23 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structures of the N-terminal modules imply large domain motions during catalysis by methionine synthase.
Proc.Natl.Acad.Sci.Usa, 101, 2004
|
|
1Q8A
 
 | | Cobalamin-dependent methionine synthase (1-566) from Thermotoga maritima (Cd2+:L-Hcy complex, Se-Met) | | Descriptor: | 2-AMINO-4-MERCAPTO-BUTYRIC ACID, 5-methyltetrahydrofolate S-homocysteine methyltransferase, CADMIUM ION | | Authors: | Evans, J.C, Huddler, D.P, Hilgers, M.T, Romanchuk, G, Matthews, R.G, Ludwig, M.L. | | Deposit date: | 2003-08-20 | | Release date: | 2004-03-23 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structures of the N-terminal modules imply large domain motions during catalysis by methionine synthase.
Proc.Natl.Acad.Sci.Usa, 101, 2004
|
|
5XCW
 
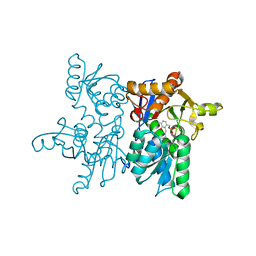 | |
5ILZ
 
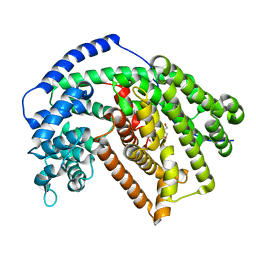 | | Tobacco 5-epi-aristolochene synthase with BIS-TRIS propane (BTP) buffer molecule | | Descriptor: | 2-[3-(2-HYDROXY-1,1-DIHYDROXYMETHYL-ETHYLAMINO)-PROPYLAMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 5-epi-aristolochene synthase | | Authors: | Koo, H.J, Louie, G.V, Xu, Y, Bowman, M, Noel, J.P. | | Deposit date: | 2016-03-05 | | Release date: | 2017-03-08 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Small-molecule buffer components can directly affect terpene-synthase activity by interacting with the substrate-binding site of the enzyme
To Be Published
|
|
4CX7
 | | Structure of human iNOS heme domain in complex with (R)-6-(3-AMINO-2-(5-(2-(6-AMINO-4- METHYLPYRIDIN-2-YL)ETHYL)PYRIDIN-3-YL)PROPYL)-4- METHYLPYRIDIN-2-AMINE | | Descriptor: | (R)-6-(3-amino-2-(5-(2-(6-amino-4-methylpyridin-2-yl)ethyl)pyridin-3-yl)propyl)-4-methylpyridin-2-amine, 5,6,7,8-TETRAHYDROBIOPTERIN, GLYCEROL, ... | | Authors: | Li, H, Poulos, T.L. | | Deposit date: | 2014-04-03 | | Release date: | 2014-08-13 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (3.16 Å) | | Cite: | Mobility of a Conserved Tyrosine Residue Controls Isoform-Dependent Enzyme-Inhibitor Interactions in Nitric Oxide Synthases.
Biochemistry, 53, 2014
|
|
8QN5
 
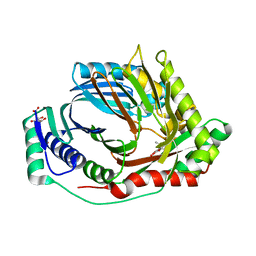 | | M. tuberculosis salicylate synthase MbtI in complex with methyl-AMT (new crystal form) | | Descriptor: | 3-{[(1Z)-1-carboxyprop-1-en-1-yl]oxy}-2-hydroxybenzoic acid, AMMONIUM ION, CITRATE ANION, ... | | Authors: | Mori, M, Villa, S, Meneghetti, M, Bellinzoni, M. | | Deposit date: | 2023-09-25 | | Release date: | 2023-11-15 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.544 Å) | | Cite: | Structural Study of a New MbtI-Inhibitor Complex: Towards an Optimized Model for Structure-Based Drug Discovery.
Pharmaceuticals, 16, 2023
|
|
8E0Z
 
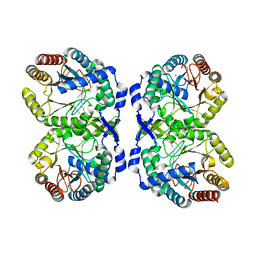 | |
8E0V
 
 | | DAHP (3-deoxy-D-arabinoheptulosonate-7-phosphate) Synthase complexed with Mn(II), PEP, and Pi in unbound:(bound)2:other Conformations | | Descriptor: | MANGANESE (II) ION, PHOSPHATE ION, PHOSPHOENOLPYRUVATE, ... | | Authors: | Berti, P.J, Junop, M.S, Grainger, R. | | Deposit date: | 2022-08-09 | | Release date: | 2022-10-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Role of Half-of-Sites Reactivity and Inter-Subunit Communications in DAHP Synthase Catalysis and Regulation.
Biochemistry, 61, 2022
|
|
8E0X
 
 | | DAHP (3-deoxy-D-arabinoheptulosonate-7-phosphate) Synthase complexed with Mn(II), PEP, and Pi in unbound:(bound)2:other Conformations | | Descriptor: | MANGANESE (II) ION, PHOSPHATE ION, PHOSPHOENOLPYRUVATE, ... | | Authors: | Berti, P.J, Junop, M.S, Grainger, R. | | Deposit date: | 2022-08-09 | | Release date: | 2022-10-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Role of Half-of-Sites Reactivity and Inter-Subunit Communications in DAHP Synthase Catalysis and Regulation.
Biochemistry, 61, 2022
|
|
8E0T
 
 | |
8E0S
 
 | | DAHP (3-deoxy-D-arabinoheptulosonate-7-phosphate) Synthase complexed with DAHP Oxime in unbound:(bound)2:unbound conformations | | Descriptor: | DAHP Oxime, Phospho-2-dehydro-3-deoxyheptonate aldolase, Phe-sensitive, ... | | Authors: | Berti, P.J, Junop, M.S, Grainger, R. | | Deposit date: | 2022-08-09 | | Release date: | 2022-10-12 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Role of Half-of-Sites Reactivity and Inter-Subunit Communications in DAHP Synthase Catalysis and Regulation.
Biochemistry, 61, 2022
|
|
5YPW
 
 | | Crystal structure of IlvN.Val-1b | | Descriptor: | Acetolactate synthase isozyme 1 small subunit, VALINE | | Authors: | Sarma, S.P, Bansal, A, Schindelin, H, Demeler, B. | | Deposit date: | 2017-11-03 | | Release date: | 2018-09-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystallographic Structures of IlvN·Val/Ile Complexes: Conformational Selectivity for Feedback Inhibition of Aceto Hydroxy Acid Synthases.
Biochemistry, 58, 2019
|
|
5YPY
 
 | | Crystal structure of IlvN. Val-1c | | Descriptor: | Acetolactate synthase isozyme 1 small subunit, VALINE | | Authors: | Sarma, S.P, Bansal, A, Schindelin, H, Demeler, B. | | Deposit date: | 2017-11-04 | | Release date: | 2018-09-19 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.966 Å) | | Cite: | Crystallographic Structures of IlvN·Val/Ile Complexes: Conformational Selectivity for Feedback Inhibition of Aceto Hydroxy Acid Synthases.
Biochemistry, 58, 2019
|
|
5XCP
 
 | |
5XCN
 
 | |
