1HT7
 
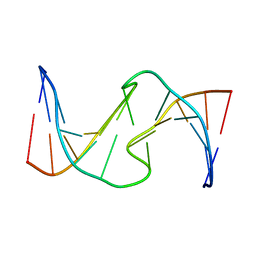 | |
7R0W
 
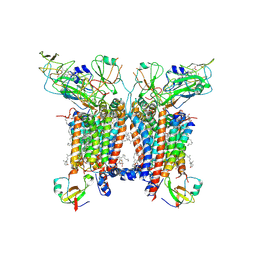 | | 2.8 Angstrom cryo-EM structure of the dimeric cytochrome b6f-PetP complex from Synechocystis sp. PCC 6803 with natively bound lipids and plastoquinone molecules | | Descriptor: | (1R)-2-{[{[(2S)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL (11E)-OCTADEC-11-ENOATE, (1S,8E)-1-{[(2S)-1-hydroxy-3-{[(1S)-1-hydroxypentadecyl]oxy}propan-2-yl]oxy}heptadec-8-en-1-ol, (4S,7R)-4-HYDROXY-N,N,N-TRIMETHYL-9-OXO-7-[(PALMITOYLOXY)METHYL]-3,5,8-TRIOXA-4-PHOSPHAHEXACOSAN-1-AMINIUM 4-OXIDE, ... | | Authors: | Farmer, D.F, Proctor, M.S, Malone, L.A, Swainsbury, D.P.K, Hawkings, F.R, Hitchcock, A, Johnson, M.P. | | Deposit date: | 2022-02-02 | | Release date: | 2022-07-06 | | Last modified: | 2024-07-10 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | Cryo-EM structures of the Synechocystis sp. PCC 6803 cytochrome b6f complex with and without the regulatory PetP subunit.
Biochem.J., 479, 2022
|
|
5JIK
 
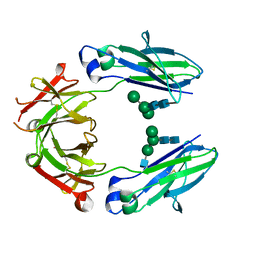 | | Crystal structure of HER2 binding IgG1-Fc (Fcab H10-03-6) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-2)-alpha-D-mannopyranose-(1-6)-[alpha-D-mannopyranose-(1-3)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Ig gamma-1 chain C region, alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Humm, A, Lobner, E, Mlynek, G, Obinger, C, Djinovic-Carugo, K. | | Deposit date: | 2016-04-22 | | Release date: | 2017-04-12 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Fcab-HER2 Interaction: a Menage a Trois. Lessons from X-Ray and Solution Studies.
Structure, 25, 2017
|
|
8A5L
 
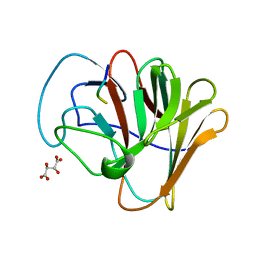 | | TRIM7 PRYSPRY in complex with a 2BC peptide TIEALFQ | | Descriptor: | 2BC peptide TIEALFQ, D(-)-TARTARIC ACID, E3 ubiquitin-protein ligase TRIM7 | | Authors: | Luptak, J. | | Deposit date: | 2022-06-15 | | Release date: | 2022-07-06 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.622 Å) | | Cite: | TRIM7 Restricts Coxsackievirus and Norovirus Infection by Detecting the C-Terminal Glutamine Generated by 3C Protease Processing.
Viruses, 14, 2022
|
|
8YEY
 
 | | TRIP4 ASCH domain in complex with ssDNA-1 | | Descriptor: | Activating signal cointegrator 1, DNA (5'-D(*GP*TP*TP*TP*C)-3') | | Authors: | Ding, J, Yang, H, Hu, C. | | Deposit date: | 2024-02-23 | | Release date: | 2024-06-26 | | Last modified: | 2024-08-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Biochemical and structural characterization of the DNA-binding properties of human TRIP4 ASCH domain reveals insights into its functional role.
Structure, 32, 2024
|
|
8YXW
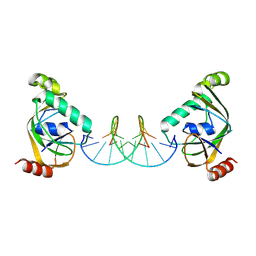 
 | | TRIP4 ASCH domain in complex with a 12bp dsDNA (5'-TGAGGTACCTCC-3') | | Descriptor: | Activating signal cointegrator 1, DNA (5'-D(*GP*GP*AP*GP*GP*TP*AP*CP*CP*TP*CP*A)-3'), DNA (5'-D(*TP*GP*AP*GP*GP*TP*AP*CP*CP*TP*CP*C)-3') | | Authors: | Ding, J, Yang, H, Hu, C. | | Deposit date: | 2024-04-03 | | Release date: | 2024-06-26 | | Last modified: | 2024-08-28 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Biochemical and structural characterization of the DNA-binding properties of human TRIP4 ASCH domain reveals insights into its functional role.
Structure, 32, 2024
|
|
5EV4
 
 | |
8A2B
 
 | | EGFR kinase domain (L858R/V948R) in complex with 2-(6,7-dihydro-5H-pyrrolo[1,2-c]imidazol-1-yl)-2-[6-[2-[4-[[4-(hydroxymethyl)-1-piperidyl]methyl]phenyl]ethynyl]-1-oxo-4-(trifluoromethyl)isoindolin-2-yl]-N-thiazol-2-yl-acetamide | | Descriptor: | (2R)-2-(6,7-dihydro-5H-pyrrolo[1,2-c]imidazol-1-yl)-2-[5-[2-[4-[[4-(hydroxymethyl)piperidin-1-yl]methyl]phenyl]ethynyl]-3-oxidanylidene-7-(trifluoromethyl)-1H-isoindol-2-yl]-N-(1,3-thiazol-2-yl)ethanamide, Epidermal growth factor receptor | | Authors: | Kuglstatter, A, Ehler, A. | | Deposit date: | 2022-06-03 | | Release date: | 2022-10-19 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Discovery of Novel Allosteric EGFR L858R Inhibitors for the Treatment of Non-Small-Cell Lung Cancer as a Single Agent or in Combination with Osimertinib.
J.Med.Chem., 65, 2022
|
|
8A2A
 
 | | EGFR kinase domain in complex with 2-(6,7-dihydro-5H-pyrrolo[1,2-c]imidazol-1-yl)-2-[6-[2-[4-[[4-(hydroxymethyl)-1-piperidyl]methyl]phenyl]ethynyl]-1-oxo-4-(trifluoromethyl)isoindolin-2-yl]-N-thiazol-2-yl-acetamide (form 2) | | Descriptor: | (2R)-2-(6,7-dihydro-5H-pyrrolo[1,2-c]imidazol-1-yl)-2-[5-[2-[4-[[4-(hydroxymethyl)piperidin-1-yl]methyl]phenyl]ethynyl]-3-oxidanylidene-7-(trifluoromethyl)-1H-isoindol-2-yl]-N-(1,3-thiazol-2-yl)ethanamide, Epidermal growth factor receptor, SULFATE ION | | Authors: | Kuglstatter, A, Ehler, A. | | Deposit date: | 2022-06-03 | | Release date: | 2022-10-19 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.43 Å) | | Cite: | Discovery of Novel Allosteric EGFR L858R Inhibitors for the Treatment of Non-Small-Cell Lung Cancer as a Single Agent or in Combination with Osimertinib.
J.Med.Chem., 65, 2022
|
|
5F6D
 
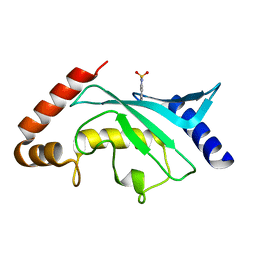 | | Crystal structure of Ubc9 (K48A/K49A/E54A) complexed with Fragment 6 | | Descriptor: | 6~{H}-benzo[c][1,2]benzothiazine 5,5-dioxide, SUMO-conjugating enzyme UBC9 | | Authors: | Lountos, G.T, Hewitt, W.M, Zlotkowski, K, Dahlhauser, S, Saunders, L.B, Needle, D, Tropea, J.E, Zhan, C, Wei, G, Ma, B, Nussinov, R, Schneekloth, J.S.Jr, Waugh, D.S. | | Deposit date: | 2015-12-05 | | Release date: | 2016-04-27 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.553 Å) | | Cite: | Insights Into the Allosteric Inhibition of the SUMO E2 Enzyme Ubc9.
Angew.Chem.Int.Ed.Engl., 55, 2016
|
|
8R3X
 
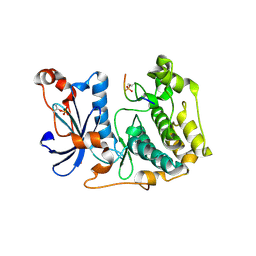 | | Crystal structure of aPKC Iota kinase domain with LLGL2 peptide | | Descriptor: | LLGL scribble cell polarity complex component 2, Protein kinase C iota type | | Authors: | Soriano, E.V, Earl, C.P, Briggs, D.C, McDonald, N.Q. | | Deposit date: | 2023-11-10 | | Release date: | 2024-11-20 | | Last modified: | 2025-04-23 | | Method: | X-RAY DIFFRACTION (2.591 Å) | | Cite: | Capture, mutual inhibition and release mechanism for aPKC-Par6 and its multisite polarity substrate Lgl.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8R3Y
 
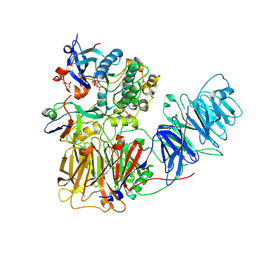 | | Cryo EM structure of a stable LGL/aPKC Iota/Par-6 complex | | Descriptor: | Lethal(2) giant larvae protein homolog 1, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER, Partitioning defective 6 homolog alpha, ... | | Authors: | Earl, C.P, Briggs, D.C, McDonald, N.Q. | | Deposit date: | 2023-11-10 | | Release date: | 2024-11-20 | | Last modified: | 2025-04-23 | | Method: | ELECTRON MICROSCOPY (3.68 Å) | | Cite: | Capture, mutual inhibition and release mechanism for aPKC-Par6 and its multisite polarity substrate Lgl.
Nat.Struct.Mol.Biol., 32, 2025
|
|
8UZ6
 
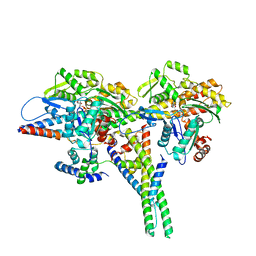 | |
8UWX
 
 | |
8V0I
 
 | |
8UYD
 
 | |
8UZ5
 
 | |
8V0K
 
 | |
8UWY
 
 | |
6Y9N
 
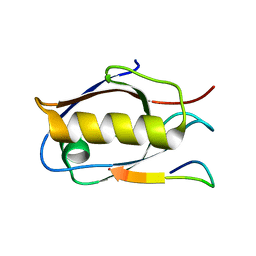 | | Crystal structure of Whirlin PDZ3_C-ter in complex with Myosin 15a C-terminal PDZ binding motif peptide | | Descriptor: | Unconventional myosin-XV, Whirlin | | Authors: | Zhu, Y, Delhommel, F, Haouz, A, Caillet-Saguy, C, Vaney, M, Mechaly, A.E, Wolff, N. | | Deposit date: | 2020-03-10 | | Release date: | 2020-10-07 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Deciphering the Unexpected Binding Capacity of the Third PDZ Domain of Whirlin to Various Cochlear Hair Cell Partners.
J.Mol.Biol., 432, 2020
|
|
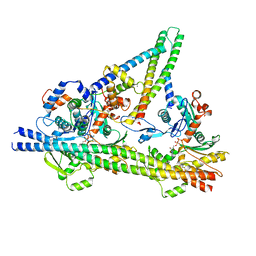
8UZX
 
 | |
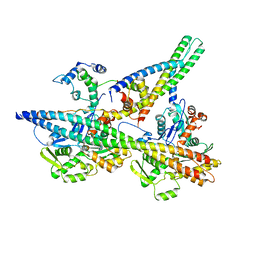
8UWW
 
 | |
8V01
 
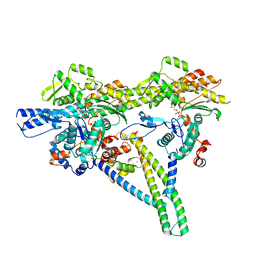 | |
8UZY
 
 | |
8ZM8
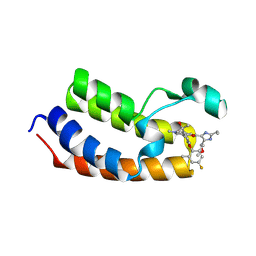 
 | | Crystal Structure of the first bromodomain of human BRD4 BD1 in complex with the inhibitor Y13221 | | Descriptor: | 2-[2-ethyl-4-(methoxymethyl)-1~{H}-imidazol-5-yl]-7-[2-(4-fluoranyl-2,6-dimethyl-phenoxy)-5-(2-oxidanylpropan-2-yl)phenyl]-5-methyl-furo[3,2-c]pyridin-4-one, BRD4_HUMAN | | Authors: | Li, J, Hu, Q, Xu, H, Zhao, X, Zhang, C, Zhu, R, Wu, X, Zhang, Y, Xu, Y. | | Deposit date: | 2024-05-22 | | Release date: | 2024-12-11 | | Last modified: | 2024-12-25 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Discovery of the First BRD4 Second Bromodomain (BD2)-Selective Inhibitors.
J.Med.Chem., 67, 2024
|
|
