4OSO
 
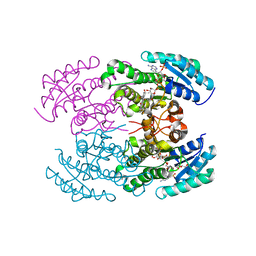 | | The crystal structure of landomycin C-6 ketoreductase LanV with bound NADP and rabelomycin | | Descriptor: | DI(HYDROXYETHYL)ETHER, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Reductase homolog, ... | | Authors: | Paananen, P, Niiranen, L, Patrikainen, P, Metsa-Ketela, M. | | Deposit date: | 2014-02-13 | | Release date: | 2014-10-01 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure-based engineering of angucyclinone 6-ketoreductases.
Chem.Biol., 21, 2014
|
|
6FV6
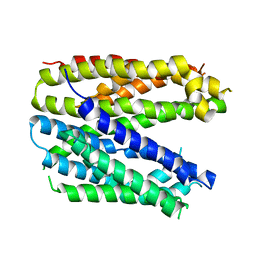 
 | | Monomer structure of the MATE family multidrug resistance transporter Aq_128 from Aquifex aeolicus in the outward-facing state | | Descriptor: | Aq128 | | Authors: | Zhao, J, Safarian, S, Thielmann, Y, Xie, H, Wang, J, Michel, H. | | Deposit date: | 2018-03-01 | | Release date: | 2019-03-20 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3.8 Å) | | Cite: | Monomer structure of Aq128 in the outward-facing state
To Be Published
|
|
6FV8
 
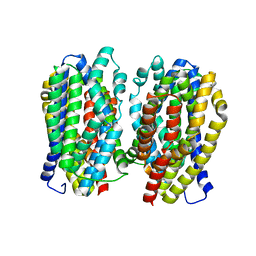 | | Dimer structure of the MATE family multidrug resistance transporter Aq_128 from Aquifex aeolicus in the outward-facing state | | Descriptor: | Aq128 | | Authors: | Zhao, J, Safarian, S, Thielmann, Y, Xie, H, Wang, J, Michel, H. | | Deposit date: | 2018-03-01 | | Release date: | 2019-03-20 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Dimer structure of the MATE family multidrug resistance transporter Aq128
in the outward-facing state
To Be Published
|
|
4OVI
 
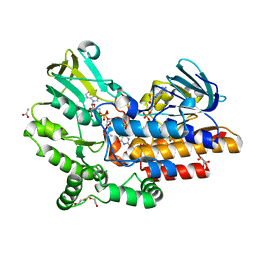 | | Phenylacetone monooxygenase: oxidised enzyme in complex with APADP | | Descriptor: | 3-ACETYLPYRIDINE ADENINE DINUCLEOTIDE, ACETATE ION, CHLORIDE ION, ... | | Authors: | Martinoli, C, Fraaije, M.W, Mattevi, A. | | Deposit date: | 2013-09-19 | | Release date: | 2014-05-28 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Beyond the Protein Matrix: Probing Cofactor Variants in a Baeyer-Villiger Oxygenation Reaction.
Acs Catalysis, 3, 2013
|
|
6FV7
 
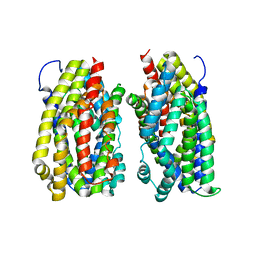 | | Dimer structure of the MATE family multidrug resistance transporter Aq_128 from Aquifex aeolicus in the outward-facing state | | Descriptor: | Aq128 | | Authors: | Zhao, J, Safarian, S, Thielmann, Y, Xie, H, Wang, J, Michel, H. | | Deposit date: | 2018-03-01 | | Release date: | 2019-03-20 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3.7 Å) | | Cite: | Dimer structure of Aq128 in the outward-facing state
To Be Published
|
|
1I1I
 
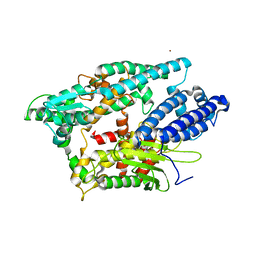 | | NEUROLYSIN (ENDOPEPTIDASE 24.16) CRYSTAL STRUCTURE | | Descriptor: | NEUROLYSIN, ZINC ION | | Authors: | Brown, C.K, Madauss, K, Lian, W, Tolbert, W.D, Beck, M.R, Rodgers, D.W. | | Deposit date: | 2001-02-01 | | Release date: | 2001-02-28 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of neurolysin reveals a deep channel that limits substrate access.
Proc.Natl.Acad.Sci.USA, 98, 2001
|
|
4TV7
 
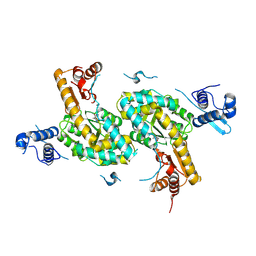 | |
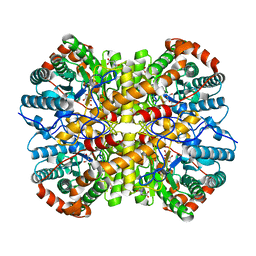
5NHC
 | |
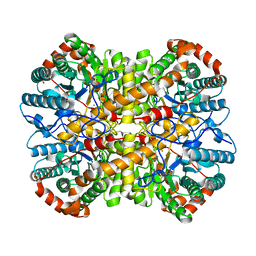
5NHB
 | |
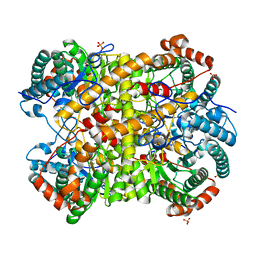
5NHA
 | |
5N7J
 
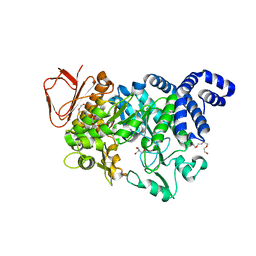 | |
5NH9
 
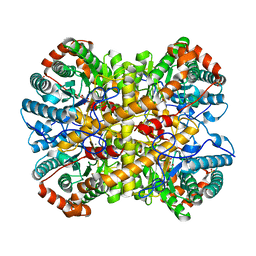 | |
5N6V
 
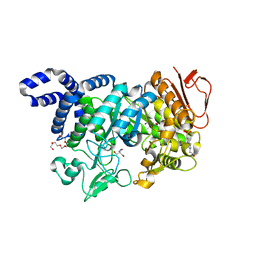 | | Crystal structure of Neisseria polysaccharea amylosucrase mutant derived from Neutral genetic Drift-based engineering | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, Amylosucrase, ... | | Authors: | Daude, D, Verges, A, Tranier, S. | | Deposit date: | 2017-02-16 | | Release date: | 2018-03-07 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Neutral Genetic Drift-Based Engineering of a Sucrose-Utilizing Enzyme toward Glycodiversification.
Acs Catalysis, 2019
|
|
1PZJ
 
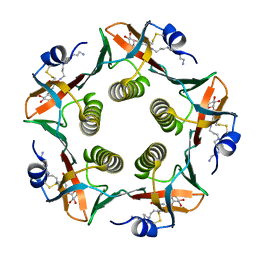 | | Cholera Toxin B-Pentamer Complexed With Nitrophenyl Galactoside 5 | | Descriptor: | Cholera Toxin B Subunit, N-{3-[4-(3-AMINO-PROPYL)-PIPERAZIN-1-YL]-PROPYL}-3-NITRO-5-(GALACTOPYRANOSYL)-ALPHA-BENZAMIDE, N-{3-[4-(3-AMINO-PROPYL)-PIPERAZIN-1-YL]-PROPYL}-3-NITRO-5-(GALACTOPYRANOSYL)-BETA-BENZAMIDE | | Authors: | Mitchell, D.D, Pickens, J.C, Korotkov, K, Fan, E, Hol, W.G.J. | | Deposit date: | 2003-07-11 | | Release date: | 2004-03-09 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.46 Å) | | Cite: | 3,5-Substituted phenyl galactosides as leads in designing effective cholera toxin antagonists; synthesis and crystallographic studies
Bioorg.Med.Chem., 12, 2004
|
|
1NX9
 
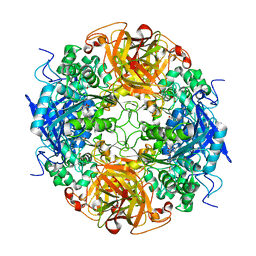 | | Acetobacter turbidans alpha-amino acid ester hydrolase S205A mutant complexed with ampicillin | | Descriptor: | (2S,5R,6R)-6-{[(2R)-2-AMINO-2-PHENYLETHANOYL]AMINO}-3,3-DIMETHYL-7-OXO-4-THIA-1-AZABICYCLO[3.2.0]HEPTANE-2-CARBOXYLIC ACID, GLYCEROL, alpha-amino acid ester hydrolase | | Authors: | Barends, T.R.M, Polderman-Tijmes, J.J, Jekel, P.A, Janssen, D.B, Dijkstra, B.W. | | Deposit date: | 2003-02-10 | | Release date: | 2004-03-09 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Acetobacter turbidans {alpha}-Amino Acid Ester Hydrolase: HOW A SINGLE MUTATION IMPROVES AN ANTIBIOTIC-PRODUCING ENZYME.
J.Biol.Chem., 281, 2006
|
|
5OEI
 
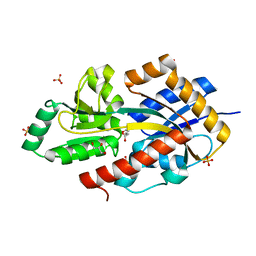 | | R. palustris Rpa4515 with oxoadipate | | Descriptor: | 2-OXOADIPIC ACID, GLYCEROL, SULFATE ION, ... | | Authors: | Rosa, L, Rafferty, J, Kelly, D, Dix, S.R. | | Deposit date: | 2017-07-07 | | Release date: | 2017-11-08 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Structural basis for high-affinity adipate binding to AdpC (RPA4515), an orphan periplasmic-binding protein from the tripartite tricarboxylate transporter (TTT) family in Rhodopseudomonas palustris.
FEBS J., 284, 2017
|
|
1O7T
 
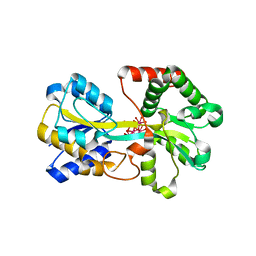 | | Metal nanoclusters bound to the Ferric Binding Protein from Neisseria gonorrhoeae. | | Descriptor: | HF OXO CLUSTER HF5, HF-OXO-PHOSPHATE CLUSTER PHF, IRON BINDING PROTEIN, ... | | Authors: | Alexeev, D, Zu, H, Guo, M, Zhong, W, Hunter, D.J.B, Yang, W, Campopiano, D.J, Sadler, P.J. | | Deposit date: | 2002-11-12 | | Release date: | 2003-02-27 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | A novel protein-mineral interface.
Nat. Struct. Biol., 10, 2003
|
|
5NH7
 
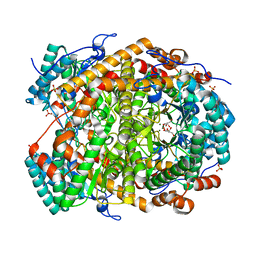 | |
5NH4
 
 | |
5NHD
 
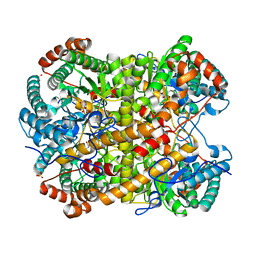 | |
1PZK
 
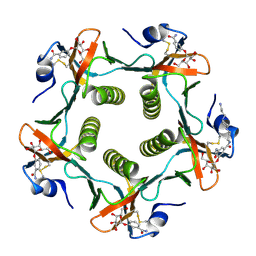 | | Cholera Toxin B-Pentamer Complexed With N-Acyl Phenyl Galactoside 9h | | Descriptor: | Cholera Toxin B Subunit, N-{3-[4-(3-AMINO-PROPYL)-PIPERAZIN-1-YL]-PROPYL}-3-(2-THIOPHEN-2-YL-ACETYLAMINO)-5-(3,4,5-TRIHYDROXY-6-HYDROXYMETHYL-TETRAHYDRO-PYRAN-2-YLOXY)-BENZAMIDE | | Authors: | Mitchell, D.D, Pickens, J.C, Korotkov, K, Fan, E, Hol, W.G.J. | | Deposit date: | 2003-07-11 | | Release date: | 2004-03-09 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | 3,5-Substituted phenyl galactosides as leads in designing effective cholera toxin antagonists; synthesis and crystallographic studies
Bioorg.Med.Chem., 12, 2004
|
|
2YM1
 
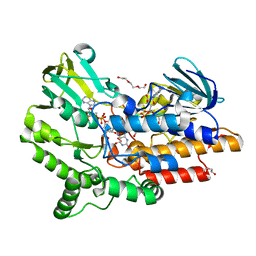 | | SNAPSHOTS OF ENZYMATIC BAEYER-VILLIGER CATALYSIS: OXYGEN ACTIVATION AND INTERMEDIATE STABILIZATION: Arg337Lys MUTANT IN COMPLEX WITH NADP | | Descriptor: | DI(HYDROXYETHYL)ETHER, FLAVIN-ADENINE DINUCLEOTIDE, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Orru, R, Dudek, H.M, Martinoli, C, Torres Pazmino, D.E, Royant, A, Weik, M, Fraaije, M.W, Mattevi, A. | | Deposit date: | 2011-06-06 | | Release date: | 2011-06-22 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.28 Å) | | Cite: | Snapshots of Enzymatic Baeyer-Villiger Catalysis: Oxygen Activation and Intermediate Stabilization.
J.Biol.Chem., 286, 2011
|
|
2YLS
 
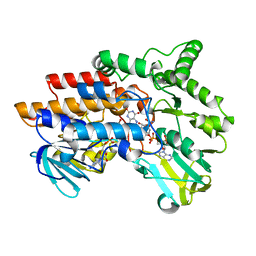 | | SNAPSHOTS OF ENZYMATIC BAEYER-VILLIGER CATALYSIS: OXYGEN ACTIVATION AND INTERMEDIATE STABILIZATION: REDUCED ENZYME BOUND TO NADP | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, PHENYLACETONE MONOOXYGENASE | | Authors: | Orru, R, Dudek, H.M, Martinoli, C, Torres Pazmino, D.E, Royant, A, Weik, M, Fraaije, M.W, Mattevi, A. | | Deposit date: | 2011-06-06 | | Release date: | 2011-06-22 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Snapshots of Enzymatic Baeyer-Villiger Catalysis: Oxygen Activation and Intermediate Stabilization.
J.Biol.Chem., 286, 2011
|
|
2YM2
 
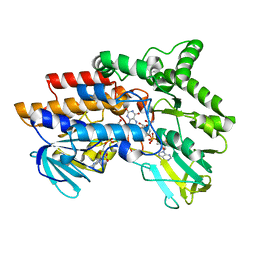 | | SNAPSHOTS OF ENZYMATIC BAEYER-VILLIGER CATALYSIS: OXYGEN ACTIVATION AND INTERMEDIATE STABILIZATION: Arg337Lys MUTANT REDUCED STATE WITH NADP | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, PHENYLACETONE MONOOXYGENASE | | Authors: | Orru, R, Dudek, H.M, Martinoli, C, Torres Pazmino, D.E, Royant, A, Weik, M, Fraaije, M.W, Mattevi, A. | | Deposit date: | 2011-06-06 | | Release date: | 2011-06-22 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Snapshots of Enzymatic Baeyer-Villiger Catalysis: Oxygen Activation and Intermediate Stabilization.
J.Biol.Chem., 286, 2011
|
|
1WZA
 
 | |
