7MQC
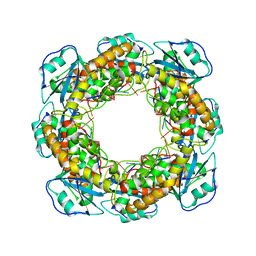 
 | | Bartonella henselae NrnC bound to pGG. C1 reconstruction. | | 分子名称: | NanoRNase C, RNA (5'-R(P*GP*G)-3') | | 著者 | Lormand, J.D, Brownfield, B, Fromme, J.C, Sondermann, H. | | 登録日 | 2021-05-05 | | 公開日 | 2021-09-15 | | 最終更新日 | 2024-05-29 | | 実験手法 | ELECTRON MICROSCOPY (3.64 Å) | | 主引用文献 | Structural characterization of NrnC identifies unifying features of dinucleotidases.
Elife, 10, 2021
|
|
7MQI
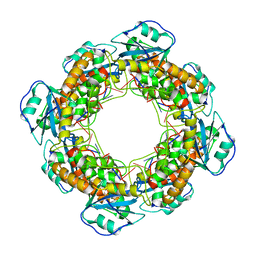 
 | | Bartonella henselae NrnC complexed with pAAAGG in the presence of Ca2+. C1 reconstruction. | | 分子名称: | NanoRNase C, RNA (5'-R(P*AP*AP*AP*GP*G)-3') | | 著者 | Lormand, J.D, Brownfield, B, Fromme, J.C, Sondermann, H. | | 登録日 | 2021-05-05 | | 公開日 | 2021-09-15 | | 最終更新日 | 2024-05-29 | | 実験手法 | ELECTRON MICROSCOPY (3.21 Å) | | 主引用文献 | Structural characterization of NrnC identifies unifying features of dinucleotidases.
Elife, 10, 2021
|
|
7MQE
 
 | | Bartonella henselae NrnC complexed with pAGG. C1 reconstruction. | | 分子名称: | NanoRNase C, RNA (5'-R(P*AP*GP*G)-3') | | 著者 | Lormand, J.D, Brownfield, B, Fromme, J.C, Sondermann, H. | | 登録日 | 2021-05-05 | | 公開日 | 2021-09-15 | | 最終更新日 | 2024-05-29 | | 実験手法 | ELECTRON MICROSCOPY (3.69 Å) | | 主引用文献 | Structural characterization of NrnC identifies unifying features of dinucleotidases.
Elife, 10, 2021
|
|
7MQB
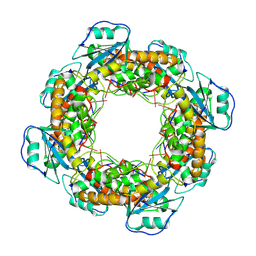 
 | | Bartonella henselae NrnC bound to pGG. D4 Symmetry | | 分子名称: | NanoRNase C, RNA (5'-R(P*GP*G)-3') | | 著者 | Lormand, J.D, Brownfield, B, Fromme, J.C, Sondermann, H. | | 登録日 | 2021-05-05 | | 公開日 | 2021-09-15 | | 最終更新日 | 2024-05-29 | | 実験手法 | ELECTRON MICROSCOPY (3.25 Å) | | 主引用文献 | Structural characterization of NrnC identifies unifying features of dinucleotidases.
Elife, 10, 2021
|
|
7MQG
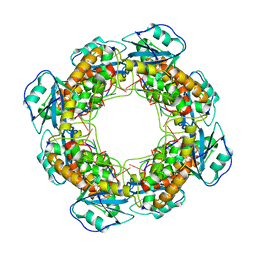 
 | | Bartonella henselae NrnC complexed with pAAAGG. C1 reconstruction. | | 分子名称: | NanoRNase C, RNA (5'-R(P*AP*AP*AP*GP*G)-3') | | 著者 | Lormand, J.D, Brownfield, B, Fromme, J.C, Sondermann, H. | | 登録日 | 2021-05-05 | | 公開日 | 2021-09-15 | | 最終更新日 | 2024-05-29 | | 実験手法 | ELECTRON MICROSCOPY (3.25 Å) | | 主引用文献 | Structural characterization of NrnC identifies unifying features of dinucleotidases.
Elife, 10, 2021
|
|
7MQH
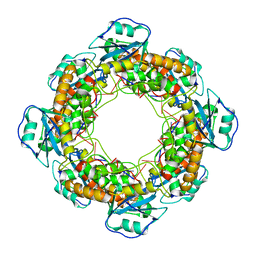 
 | | Bartonella henselae NrnC complexed with pAAAGG in the presence of Ca2+. D4 Symmetry. | | 分子名称: | NanoRNase C, RNA (5'-R(P*AP*AP*AP*GP*G)-3') | | 著者 | Lormand, J.D, Brownfield, B, Fromme, J.C, Sondermann, H. | | 登録日 | 2021-05-05 | | 公開日 | 2021-09-15 | | 最終更新日 | 2024-05-29 | | 実験手法 | ELECTRON MICROSCOPY (3.1 Å) | | 主引用文献 | Structural characterization of NrnC identifies unifying features of dinucleotidases.
Elife, 10, 2021
|
|
7MKJ
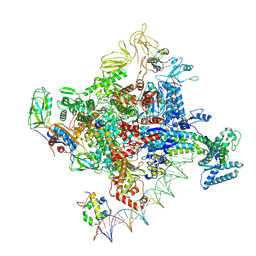 
 | | Cryo-EM structure of Escherichia coli RNA polymerase bound to T7A1 promoter DNA | | 分子名称: | CHAPSO, DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, ... | | 著者 | Saecker, R.M, Darst, S.A, Chen, J. | | 登録日 | 2021-04-23 | | 公開日 | 2021-09-29 | | 最終更新日 | 2024-05-29 | | 実験手法 | ELECTRON MICROSCOPY (2.9 Å) | | 主引用文献 | Structural origins of Escherichia coli RNA polymerase open promoter complex stability.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7MKD
 
 | | Cryo-EM structure of Escherichia coli RNA polymerase bound to lambda PR promoter DNA (class 1) | | 分子名称: | CHAPSO, DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, ... | | 著者 | Saecker, R.M, Darst, S.A, Chen, J. | | 登録日 | 2021-04-23 | | 公開日 | 2021-09-29 | | 最終更新日 | 2021-10-13 | | 実験手法 | ELECTRON MICROSCOPY (3.2 Å) | | 主引用文献 | Structural origins of Escherichia coli RNA polymerase open promoter complex stability.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
6GSJ
 
 | | Structure of T. thermophilus 70S ribosome complex with mRNA, tRNAfMet and cognate tRNAThr in the A-site | | 分子名称: | 16S ribosomal RNA, 23S ribosomal RNA, 30S ribosomal protein S10, ... | | 著者 | Rozov, A, Yusupov, M, Yusupova, G. | | 登録日 | 2018-06-14 | | 公開日 | 2018-07-04 | | 最終更新日 | 2024-01-17 | | 実験手法 | X-RAY DIFFRACTION (2.96 Å) | | 主引用文献 | Tautomeric G•U pairs within the molecular ribosomal grip and fidelity of decoding in bacteria.
Nucleic Acids Res., 46, 2018
|
|
6GZ4
 
 | | tRNA translocation by the eukaryotic 80S ribosome and the impact of GTP hydrolysis, Translocation-intermediate-POST-2 (TI-POST-2) | | 分子名称: | 18S ribosomal RNA, 28S ribosomal RNA, 5.8S ribosomal RNA, ... | | 著者 | Flis, J, Holm, M, Rundlet, E.J, Loerke, J, Hilal, T, Dabrowski, M, Buerger, J, Mielke, T, Blanchard, S.C, Spahn, C.M.T, Budkevich, T.V. | | 登録日 | 2018-07-03 | | 公開日 | 2018-12-05 | | 最終更新日 | 2022-03-30 | | 実験手法 | ELECTRON MICROSCOPY (3.6 Å) | | 主引用文献 | tRNA Translocation by the Eukaryotic 80S Ribosome and the Impact of GTP Hydrolysis.
Cell Rep, 25, 2018
|
|
8CTE
 
 | | Class 2 of erythrocyte ankyrin-1 complex (Composite map) | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Ammonium transporter Rh type A, Ankyrin-1, ... | | 著者 | Vallese, F, Kim, K, Yen, L.Y, Johnston, J.D, Noble, A.J, Cali, T, Clarke, O.B. | | 登録日 | 2022-05-14 | | 公開日 | 2022-07-20 | | 最終更新日 | 2022-07-27 | | 実験手法 | ELECTRON MICROSCOPY (2.9 Å) | | 主引用文献 | Architecture of the human erythrocyte ankyrin-1 complex.
Nat.Struct.Mol.Biol., 29, 2022
|
|
4P0M
 
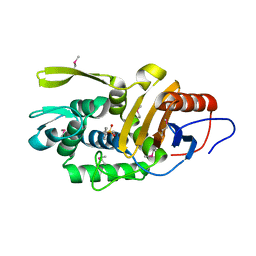 | | Crystal structure of an evolved putative penicillin-binding protein homolog, Rv2911, from Mycobacterium tuberculosis | | 分子名称: | D-alanyl-D-alanine carboxypeptidase | | 著者 | Krieger, I, Yu, M, Bursey, E, Hung, L.-W, Terwilliger, T.C, TB Structural Genomics Consortium (TBSGC) | | 登録日 | 2014-02-21 | | 公開日 | 2014-03-12 | | 最終更新日 | 2023-12-27 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | Subfamily-Specific Adaptations in the Structures of Two Penicillin-Binding Proteins from Mycobacterium tuberculosis.
Plos One, 9, 2014
|
|
6MH7
 
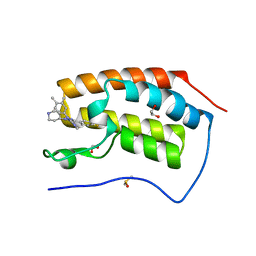 | | Crystal structure of the first bromodomain of human BRD4 in complex with SKT-68, a 1,4,5-trisubstituted imidazole analogue | | 分子名称: | 1,2-ETHANEDIOL, Bromodomain-containing protein 4, DIMETHYL SULFOXIDE, ... | | 著者 | Zhu, J.-Y, Schonbrunn, E. | | 登録日 | 2018-09-17 | | 公開日 | 2019-08-07 | | 最終更新日 | 2023-10-11 | | 実験手法 | X-RAY DIFFRACTION (1.74 Å) | | 主引用文献 | Molecular Basis for the N-Terminal Bromodomain-and-Extra-Terminal-Family Selectivity of a Dual Kinase-Bromodomain Inhibitor.
J.Med.Chem., 61, 2018
|
|
7MKE
 
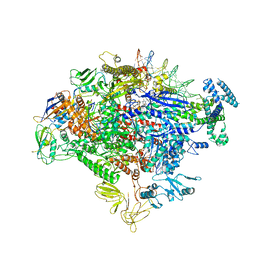 | | Cryo-EM structure of Escherichia coli RNA polymerase bound to lambda PR promoter DNA (class 2) | | 分子名称: | CHAPSO, DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, ... | | 著者 | Saecker, R.M, Darst, S.A, Chen, J. | | 登録日 | 2021-04-23 | | 公開日 | 2021-09-29 | | 最終更新日 | 2021-10-13 | | 実験手法 | ELECTRON MICROSCOPY (3.7 Å) | | 主引用文献 | Structural origins of Escherichia coli RNA polymerase open promoter complex stability.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
7MKI
 
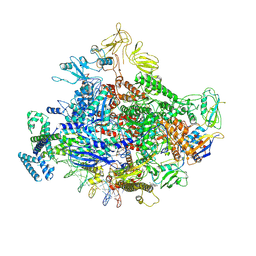 | | Cryo-EM structure of Escherichia coli RNA polymerase bound to lambda PR (-5G to C) promoter DNA | | 分子名称: | CHAPSO, DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, ... | | 著者 | Saecker, R.M, Darst, S.A, Chen, J. | | 登録日 | 2021-04-23 | | 公開日 | 2021-09-29 | | 最終更新日 | 2024-05-29 | | 実験手法 | ELECTRON MICROSCOPY (3.5 Å) | | 主引用文献 | Structural origins of Escherichia coli RNA polymerase open promoter complex stability.
Proc.Natl.Acad.Sci.USA, 118, 2021
|
|
6Y76
 
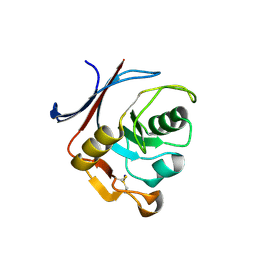 | | AP01 - a redesigned transferrin receptor apical domain | | 分子名称: | SODIUM ION, Transferrin receptor protein 1 | | 著者 | Oberdorfer, G, Berger, S.A, Bjelic, S, Sjostrom, D.J. | | 登録日 | 2020-02-28 | | 公開日 | 2020-07-22 | | 最終更新日 | 2024-10-09 | | 実験手法 | X-RAY DIFFRACTION (1.98 Å) | | 主引用文献 | Computational backbone design enables soluble engineering of transferrin receptor apical domain.
Proteins, 88, 2020
|
|
2HXY
 
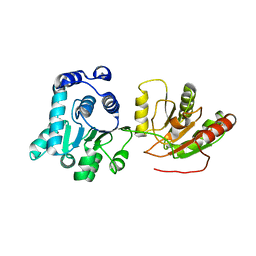 | |
6MH1
 
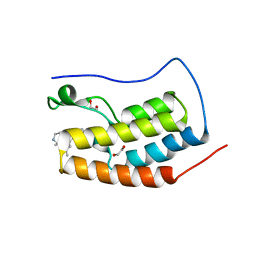 | | CRYSTAL STRUCTURE OF THE FIRST BROMODOMAIN OF HUMAN BRD4 IN COMPLEX WITH HU-10, A 1,4,5-Trisubstituted Imidazole Analogue | | 分子名称: | 1,2-ETHANEDIOL, Bromodomain-containing protein 4, N-(3,5-dimethylphenyl)-4-[4-(4-fluorophenyl)-1-(piperidin-4-yl)-1H-imidazol-5-yl]pyrimidin-2-amine | | 著者 | Zhu, J.-Y, Schonbrunn, E. | | 登録日 | 2018-09-17 | | 公開日 | 2019-08-07 | | 最終更新日 | 2024-03-13 | | 実験手法 | X-RAY DIFFRACTION (1.6 Å) | | 主引用文献 | Molecular Basis for the N-Terminal Bromodomain-and-Extra-Terminal-Family Selectivity of a Dual Kinase-Bromodomain Inhibitor.
J.Med.Chem., 61, 2018
|
|
6ZFB
 
 | | Structure of the B. subtilis RNA POLYMERASE in complex with HelD (dimer) | | 分子名称: | DNA helicase, DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, ... | | 著者 | Pei, H.-P, Hilal, T, Huang, Y.-H, Said, N, Loll, B, Wahl, M.C. | | 登録日 | 2020-06-17 | | 公開日 | 2020-10-14 | | 最終更新日 | 2024-05-15 | | 実験手法 | ELECTRON MICROSCOPY (3.9 Å) | | 主引用文献 | The delta subunit and NTPase HelD institute a two-pronged mechanism for RNA polymerase recycling.
Nat Commun, 11, 2020
|
|
6VFO
 
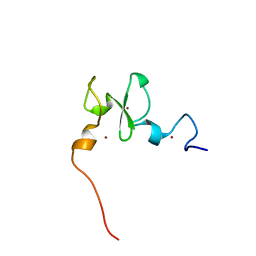 | | Solution structure of the PHD of mouse UHRF1 (NP95) | | 分子名称: | E3 ubiquitin-protein ligase UHRF1, ZINC ION | | 著者 | Lemak, A, Houliston, S, Duan, S, Arrowsmith, C.H. | | 登録日 | 2020-01-06 | | 公開日 | 2020-06-17 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Alternative splicing and allosteric regulation modulate the chromatin binding of UHRF1.
Nucleic Acids Res., 48, 2020
|
|
6VEE
 
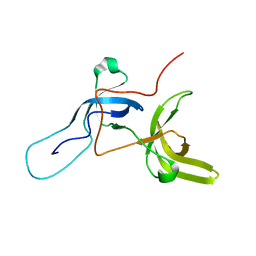 | |
6VED
 
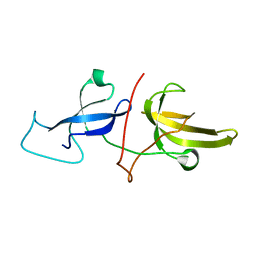 | | Solution structure of the TTD and linker region of UHRF1 | | 分子名称: | E3 ubiquitin-protein ligase UHRF1 | | 著者 | Lemak, A, Houliston, S, Duan, S, Ong, M.S, Arrowsmith, C.H. | | 登録日 | 2019-12-31 | | 公開日 | 2020-06-17 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Alternative splicing and allosteric regulation modulate the chromatin binding of UHRF1.
Nucleic Acids Res., 48, 2020
|
|
6N57
 
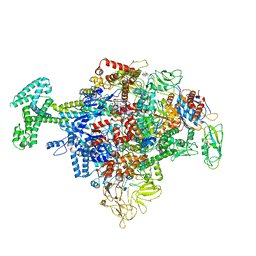 | | Cryo-EM structure of Escherichia coli RNAP polymerase bound with TraR in conformation I | | 分子名称: | CHAPSO, DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, ... | | 著者 | Chen, J, Chiu, C.E, Campbell, E.A, Darst, S.A. | | 登録日 | 2018-11-21 | | 公開日 | 2020-02-26 | | 最終更新日 | 2024-03-20 | | 実験手法 | ELECTRON MICROSCOPY (3.7 Å) | | 主引用文献 | E. coliTraR allosterically regulates transcription initiation by altering RNA polymerase conformation.
Elife, 8, 2019
|
|
4EMG
 
 | |
6XIR
 
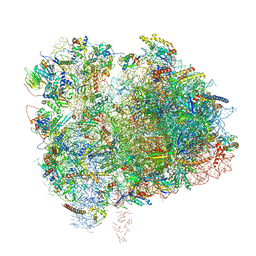 | | Cryo-EM Structure of K63 Ubiquitinated Yeast Translocating Ribosome under Oxidative Stress | | 分子名称: | 18S ribosomal RNA, 35S ribosomal RNA, 40S ribosomal protein S0-A, ... | | 著者 | Zhou, Y, Bartesaghi, A, Silva, G.M. | | 登録日 | 2020-06-21 | | 公開日 | 2020-08-26 | | 最終更新日 | 2020-09-23 | | 実験手法 | ELECTRON MICROSCOPY (3.2 Å) | | 主引用文献 | Structural impact of K63 ubiquitin on yeast translocating ribosomes under oxidative stress.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
