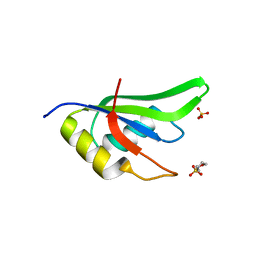
4ZKA
 
 | |
3JCT
 
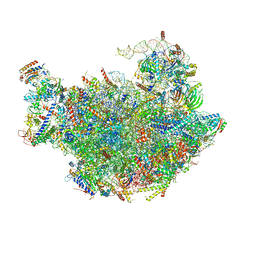 | | Cryo-em structure of eukaryotic pre-60S ribosomal subunits | | 分子名称: | 60S ribosomal protein L11-A, 60S ribosomal protein L13-A, 60S ribosomal protein L14-A, ... | | 著者 | Wu, S, Kumcuoglu, B, Yan, K.G, Brown, H, Zhang, Y.X, Tan, D, Gamalinda, M, Yuan, Y, Li, Z.F, Jakovljevic, J, Ma, C.Y, Lei, J.L, Dong, M.Q, Woolford Jr, J.L, Gao, N. | | 登録日 | 2016-03-09 | | 公開日 | 2016-06-01 | | 最終更新日 | 2024-03-20 | | 実験手法 | ELECTRON MICROSCOPY (3.08 Å) | | 主引用文献 | Diverse roles of assembly factors revealed by structures of late nuclear pre-60S ribosomes
Nature, 534, 2016
|
|
5FJ4
 
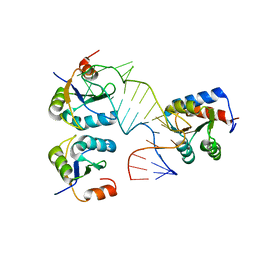 | |
5IQQ
 
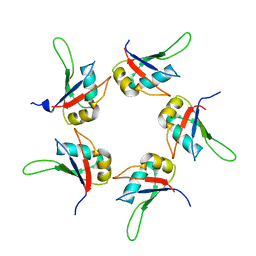 | |
2N82
 
 | |
5EV1
 
 | | Structure I of Intact U2AF65 Recognizing a 3' Splice Site Signal | | 分子名称: | DI(HYDROXYETHYL)ETHER, DNA/RNA (5'-R(*UP*UP*U)-D(P*UP*UP*(BRU)P*U)-R(P*UP*U)-3'), SODIUM ION, ... | | 著者 | Agrawal, A.A, Jenkins, J.L, Kielkopf, C.L. | | 登録日 | 2015-11-19 | | 公開日 | 2016-02-24 | | 最終更新日 | 2023-09-27 | | 実験手法 | X-RAY DIFFRACTION (2.037 Å) | | 主引用文献 | An extended U2AF(65)-RNA-binding domain recognizes the 3' splice site signal.
Nat Commun, 7, 2016
|
|
5EV2
 
 | | Structure II of Intact U2AF65 Recognizing the 3' Splice Site Signal | | 分子名称: | 1,4-DIETHYLENE DIOXIDE, DI(HYDROXYETHYL)ETHER, DNA (5'-R(P*UP*U)-D(P*UP*U)-R(P*U)-D(P*UP*(BRU)P*U)-3'), ... | | 著者 | Agrawal, A.A, Jenkins, J.L, Kielkopf, C.L. | | 登録日 | 2015-11-19 | | 公開日 | 2016-02-24 | | 最終更新日 | 2024-03-06 | | 実験手法 | X-RAY DIFFRACTION (1.86 Å) | | 主引用文献 | An extended U2AF(65)-RNA-binding domain recognizes the 3' splice site signal.
Nat Commun, 7, 2016
|
|
5EV3
 
 | |
5EV4
 
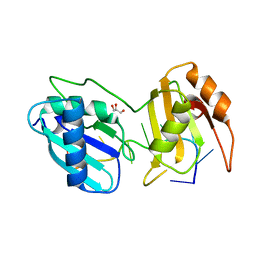 | |
4Y00
 
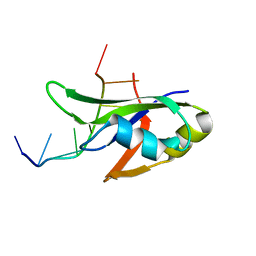 | | Crystal Structure of Human TDP-43 RRM1 Domain with D169G Mutation in Complex with an Unmodified Single-stranded DNA | | 分子名称: | DNA (5'-D(P*TP*TP*GP*AP*GP*CP*GP*T)-3'), TAR DNA-binding protein 43 | | 著者 | Chiang, C.H, Kuo, P.H, Yang, W.Z, Yuan, H.S. | | 登録日 | 2015-02-05 | | 公開日 | 2016-02-10 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (3 Å) | | 主引用文献 | Structural analysis of disease-related TDP-43 D169G mutation: linking enhanced stability and caspase cleavage efficiency to protein accumulation
Sci Rep, 6, 2016
|
|
4Y0F
 | | Crystal Structure of Human TDP-43 RRM1 Domain in Complex with an Unmodified Single-stranded DNA | | 分子名称: | DNA (5'-D(*GP*TP*TP*GP*AP*GP*CP*GP*TP*T)-3'), TAR DNA-binding protein 43 | | 著者 | Chiang, C.H, Kuo, P.H, Doudeva, L.G, Wang, Y.T, Yuan, H.S. | | 登録日 | 2015-02-06 | | 公開日 | 2016-02-10 | | 最終更新日 | 2023-11-08 | | 実験手法 | X-RAY DIFFRACTION (2.648 Å) | | 主引用文献 | Structural analysis of disease-related TDP-43 D169G mutation: linking enhanced stability and caspase cleavage efficiency to protein accumulation
Sci Rep, 6, 2016
|
|
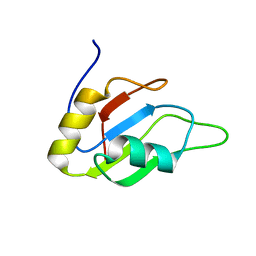
2MYF
 
 | |
5DDP
 
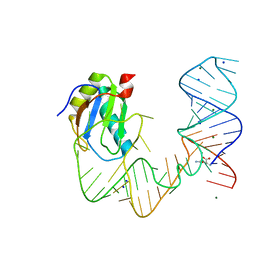 | | L-glutamine riboswitch bound with L-glutamine | | 分子名称: | GLUTAMINE, MAGNESIUM ION, RNA (61-MER), ... | | 著者 | Ren, A, Patel, D.J. | | 登録日 | 2015-08-25 | | 公開日 | 2015-12-23 | | 最終更新日 | 2024-03-06 | | 実験手法 | X-RAY DIFFRACTION (2.302 Å) | | 主引用文献 | Structural and Dynamic Basis for Low-Affinity, High-Selectivity Binding of L-Glutamine by the Glutamine Riboswitch.
Cell Rep, 13, 2015
|
|
5DDO
 
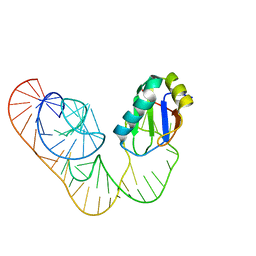 | |
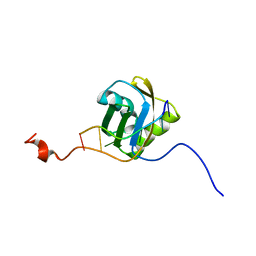
2MQO
 
 | |
5DDQ
 
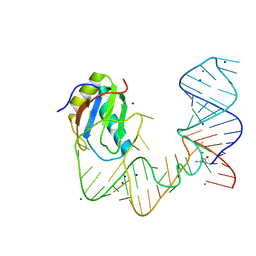 | |
5DDR
 
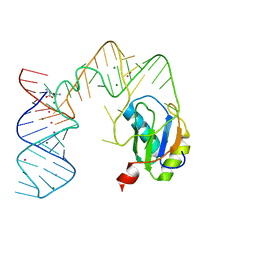 | |
5CA5
 | |
4TLQ
 
 | |
2MZS
 
 | |
2MZQ
 
 | |
2MZR
 
 | |
2MZT
 
 | |
5CYJ
 
 | | X-ray structure of human RBPMS | | 分子名称: | RNA-binding protein with multiple splicing | | 著者 | Teplova, M, Farazi, T.A, Patel, D.J. | | 登録日 | 2015-07-30 | | 公開日 | 2015-09-30 | | 最終更新日 | 2019-12-25 | | 実験手法 | X-RAY DIFFRACTION (1.79 Å) | | 主引用文献 | Structural basis underlying CAC RNA recognition by the RRM domain of dimeric RNA-binding protein RBPMS.
Q. Rev. Biophys., 49, 2016
|
|
5DET
 
 | | X-ray structure of human RBPMS in complex with the RNA | | 分子名称: | RNA (5'-R(*UP*CP*AP*C)-3'), RNA (5'-R(P*UP*CP*AP*CP*U)-3'), RNA-binding protein with multiple splicing, ... | | 著者 | Teplova, M, Farazi, T.A, Tuschl, T, Patel, D.J. | | 登録日 | 2015-08-25 | | 公開日 | 2015-09-23 | | 最終更新日 | 2023-09-27 | | 実験手法 | X-RAY DIFFRACTION (1.95 Å) | | 主引用文献 | Structural basis underlying CAC RNA recognition by the RRM domain of dimeric RNA-binding protein RBPMS.
Q. Rev. Biophys., 49, 2016
|
|
