1M7P
 
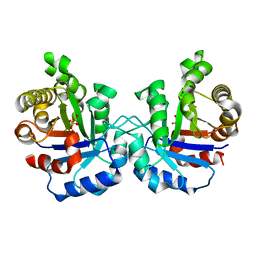 | | Plasmodium Falciparum Triosephosphate isomerase (PfTIM) compled to substrate analog glycerol-3-phosphate (G3P). | | Descriptor: | GLYCERALDEHYDE-3-PHOSPHATE, Triosephosphate Isomerase | | Authors: | Parthasarathy, S, Balaram, H, Balaram, P, Murthy, M.R.N. | | Deposit date: | 2002-07-22 | | Release date: | 2002-11-29 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structures of Plasmodium falciparum triosephosphate isomerase complexed to substrate analogues: observation of the catalytic loop in the open conformation in the ligand-bound state.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
9C0I
 
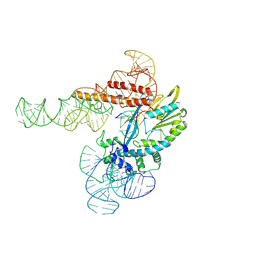 | |
9C0J
 
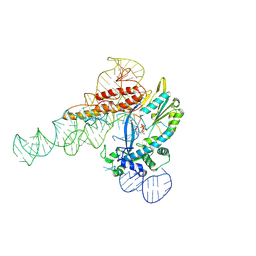 | |
1M8I
 
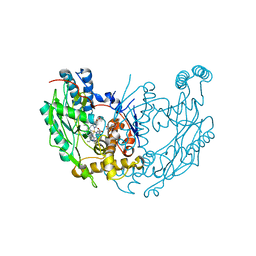 | | inducible nitric oxide synthase with 5-nitroindazole bound | | Descriptor: | 5,6,7,8-TETRAHYDROBIOPTERIN, 5-NITROINDAZOLE, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Rosenfeld, R.J, Garcin, E.D, Panda, K, Andersson, G, Aberg, A, Wallace, A.V, Stuehr, D.J, Tainer, J.A, Getzoff, E.D. | | Deposit date: | 2002-07-24 | | Release date: | 2002-08-07 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Conformational Changes in Nitric Oxide Synthases Induced by Chlorzoxazone and Nitroindazoles: Crystallographic and Computational Analyses of Inhibitor Potency
Biochemistry, 41, 2002
|
|
9C7V
 
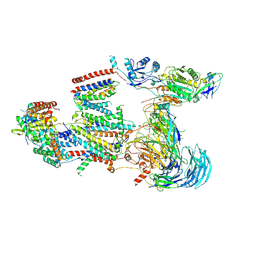 | | Structure of the human BOS:human EMC complex in GDN | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, BOS complex subunit NOMO2, ... | | Authors: | Nguyen, V.N, Tomaleri, G.P, Voorhees, R.M. | | Deposit date: | 2024-06-11 | | Release date: | 2024-08-21 | | Last modified: | 2024-09-25 | | Method: | ELECTRON MICROSCOPY (6.6 Å) | | Cite: | Role of a holo-insertase complex in the biogenesis of biophysically diverse ER membrane proteins.
Mol.Cell, 84, 2024
|
|
9C82
 
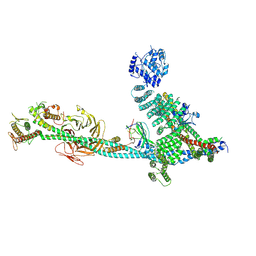 | | Structure of human ULK1C:PI3KC3-C1 supercomplex | | Descriptor: | Beclin 1-associated autophagy-related key regulator, Beclin-1, Phosphatidylinositol 3-kinase catalytic subunit type 3, ... | | Authors: | Chen, M, Hurley, J.H. | | Deposit date: | 2024-06-11 | | Release date: | 2024-07-03 | | Method: | ELECTRON MICROSCOPY (6.84 Å) | | Cite: | Structure and activation of the human autophagy-initiating ULK1C:PI3KC3-C1 supercomplex
bioRxiv, 2023
|
|
9BB7
 
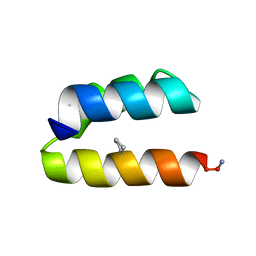 | |
4LTI
 
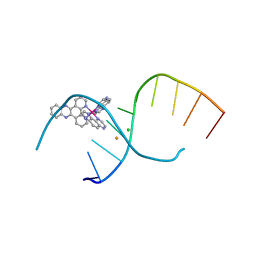 | | Dehydration/Rehydration of a Nucleic Acid system containing a Polypyridyl Ruthenium Complex at 74% relative humidity (4/7) | | Descriptor: | BARIUM ION, CHLORIDE ION, DNA, ... | | Authors: | Hall, J.P, Sanchez-Weatherby, J, Cardin, C.J. | | Deposit date: | 2013-07-23 | | Release date: | 2014-09-24 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.41 Å) | | Cite: | Controlled Dehydration of a Ruthenium Complex-DNA Crystal Induces Reversible DNA Kinking.
J.Am.Chem.Soc., 136, 2014
|
|
6O78
 
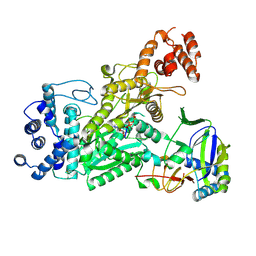 | | Crystal structure of Csm1-Csm4 cassette in complex with pppApApA | | Descriptor: | CRISPR system single-strand-specific deoxyribonuclease Cas10/Csm1 (subtype III-A), Csm4, MANGANESE (II) ION, ... | | Authors: | Jia, N, Patel, D.J. | | Deposit date: | 2019-03-07 | | Release date: | 2019-07-31 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Second Messenger cA4Formation within the Composite Csm1 Palm Pocket of Type III-A CRISPR-Cas Csm Complex and Its Release Path.
Mol.Cell, 75, 2019
|
|
6O7G
 
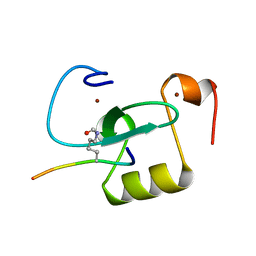 | |
2ETT
 
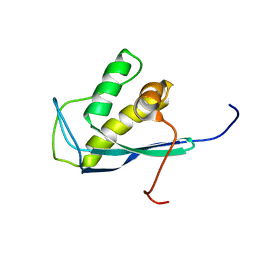 | | Solution Structure of Human Sorting Nexin 22 PX Domain | | Descriptor: | Sorting nexin-22 | | Authors: | Song, J, Zhao, Q, Tyler, R.C, Lee, M.S, Newman, C.L, Markley, J.L, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2005-10-27 | | Release date: | 2005-11-15 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of human sorting nexin 22.
Protein Sci., 16, 2007
|
|
9BB2
 
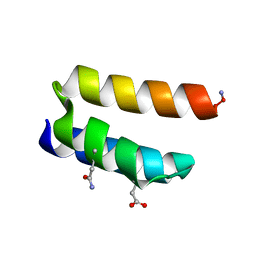 | |
2QB8
 
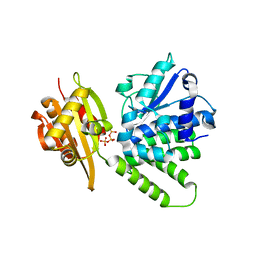 | |
5W7A
 
 | | Rabbit acyloxyacyl hydrolase (AOAH), proteolytically processed, S262A mutant, with LPS (low quality saposin domain) | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 3-HYDROXY-TETRADECANOIC ACID, ... | | Authors: | Gorelik, A, Illes, K, Nagar, B. | | Deposit date: | 2017-06-19 | | Release date: | 2018-01-03 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of the mammalian lipopolysaccharide detoxifier.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
1MGR
 
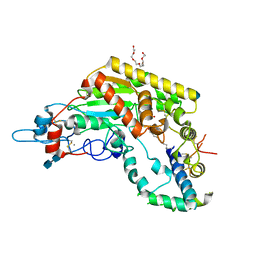
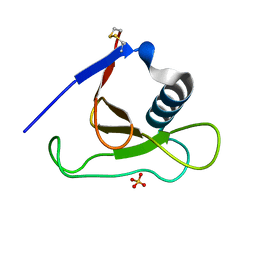 | | Crystal structure of RNase Sa3,cytotoxic microbial ribonuclease | | Descriptor: | Guanyl-specific ribonuclease Sa3, SULFATE ION | | Authors: | Sevcik, J, Urbanikova, L, Leland, P.A, Raines, R.T. | | Deposit date: | 2002-08-16 | | Release date: | 2003-02-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Links X-ray Structure of Two Crystalline Forms of a Streptomycete Ribonuclease with Cytotoxic Activity
J.Biol.Chem., 277, 2002
|
|
6O7Q
 
 | |
2FLE
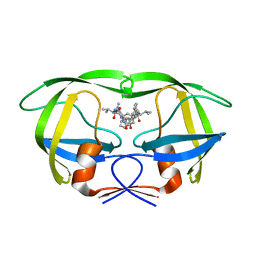 
 | | Structural analysis of asymmetric inhibitor bound to the HIV-1 Protease V82A mutant | | Descriptor: | (2S,2'S)-N,N'-[(2S,3S,4S,5S)-1-CYCLOHEXYL-3,4-DIHYDROXY-6-PHENYLHEXANE-2,5-DIYL]BIS[3-METHYL-2-({[METHYL(PYRIDIN-2-YLMETHYL)AMINO]CARBONYL}AMINO)BUTANAMIDE], GLYCEROL, pol protein | | Authors: | Clemente, J.C, Robbins, A, Dunn, B.M, Sussman, F. | | Deposit date: | 2006-01-05 | | Release date: | 2007-01-16 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Design, synthesis, evaluation, and crystallographic-based structural studies of HIV-1 protease inhibitors with reduced response to the V82A mutation.
J.Med.Chem., 51, 2008
|
|
1MGW
 
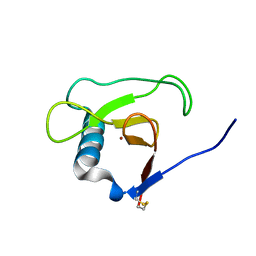 | | Crystal structure of RNase Sa3, cytotoxic microbial ribonuclease | | Descriptor: | Guanyl-specific ribonuclease Sa3, LITHIUM ION | | Authors: | Sevcik, J, Urbanikova, L, Leland, P.A, Raines, R.T. | | Deposit date: | 2002-08-16 | | Release date: | 2003-02-04 | | Last modified: | 2011-11-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Links X-ray Structure of Two Crystalline Forms of a Streptomycete Ribonuclease with Cytotoxic Activity
J.Biol.Chem., 277, 2002
|
|
4LQT
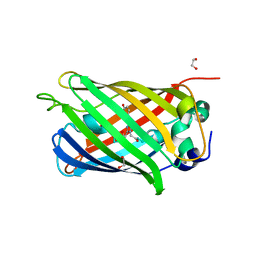 
 | | 1.10A resolution crystal structure of a superfolder green fluorescent protein (W57A) mutant | | Descriptor: | 1,2-ETHANEDIOL, Green fluorescent protein | | Authors: | Lovell, S, Xia, Y, Vo, B, Battaile, K.P, Egan, C, Karanicolas, J. | | Deposit date: | 2013-07-19 | | Release date: | 2013-12-18 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | The designability of protein switches by chemical rescue of structure: mechanisms of inactivation and reactivation.
J.Am.Chem.Soc., 135, 2013
|
|
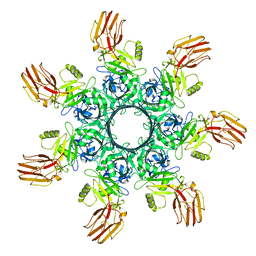
6OKT
 | |
6OLM
 
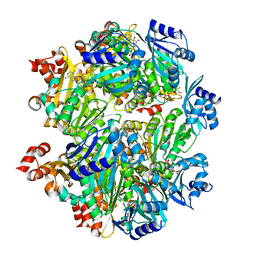 | |
4LR3
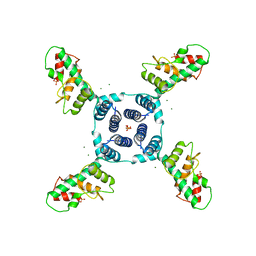 
 | | Crystal structure of E. coli YfbU at 2.5 A resolution | | Descriptor: | MAGNESIUM ION, PHOSPHATE ION, protein YfbU | | Authors: | Wang, J, Wing, R.A. | | Deposit date: | 2013-07-19 | | Release date: | 2014-04-30 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Diamonds in the rough: a strong case for the inclusion of weak-intensity X-ray diffraction data.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
2EU1
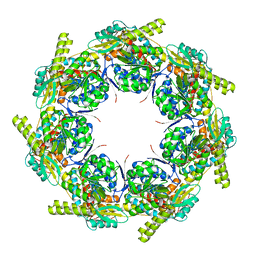 
 | | Crystal structure of the chaperonin GroEL-E461K | | Descriptor: | GROEL | | Authors: | Cabo-Bilbao, A, Spinelli, S, Sot, B, Agirre, J, Mechaly, A.E, Muga, A, Guerin, D.M.A. | | Deposit date: | 2005-10-28 | | Release date: | 2006-08-29 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3.29 Å) | | Cite: | Crystal structure of the temperature-sensitive and allosteric-defective chaperonin GroEL(E461K).
J.Struct.Biol., 155, 2006
|
|
5X50
 
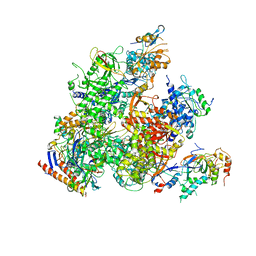 | | RNA Polymerase II from Komagataella Pastoris (Type-2 crystal) | | Descriptor: | DNA-directed RNA polymerase subunit, DNA-directed RNA polymerase subunit beta, RNA polymerase II subunit, ... | | Authors: | Ehara, H, Umehara, T, Sekine, S, Yokoyama, S. | | Deposit date: | 2017-02-14 | | Release date: | 2017-05-17 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (4.293 Å) | | Cite: | Crystal structure of RNA polymerase II from Komagataella pastoris
Biochem. Biophys. Res. Commun., 487, 2017
|
|
5X51
 
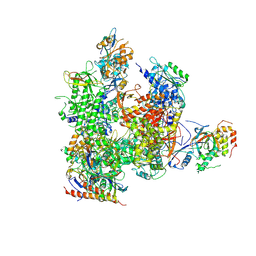 | | RNA Polymerase II from Komagataella Pastoris (Type-3 crystal) | | Descriptor: | DNA-directed RNA polymerase subunit, DNA-directed RNA polymerase subunit beta, RNA polymerase II subunit, ... | | Authors: | Ehara, H, Umehara, T, Sekine, S, Yokoyama, S. | | Deposit date: | 2017-02-14 | | Release date: | 2017-05-17 | | Method: | X-RAY DIFFRACTION (6.996 Å) | | Cite: | Crystal structure of RNA polymerase II from Komagataella pastoris
Biochem. Biophys. Res. Commun., 487, 2017
|
|
