1PHH
 
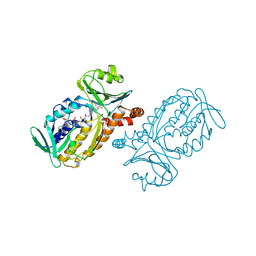 | | CRYSTAL STRUCTURE OF P-HYDROXYBENZOATE HYDROXYLASE COMPLEXED WITH ITS REACTION PRODUCT 3,4-DIHYDROXYBENZOATE | | Descriptor: | 3,4-DIHYDROXYBENZOIC ACID, FLAVIN-ADENINE DINUCLEOTIDE, P-HYDROXYBENZOATE HYDROXYLASE | | Authors: | Schreuder, H.A, Drenth, J. | | Deposit date: | 1987-11-04 | | Release date: | 1988-04-16 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of p-hydroxybenzoate hydroxylase complexed with its reaction product 3,4-dihydroxybenzoate.
J.Mol.Biol., 199, 1988
|
|
2LNA
 
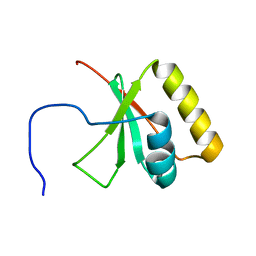 | | Solution NMR Structure of the mitochondrial inner membrane domain (residues 164-251), FtsH_ext, from the paraplegin-like protein AFG3L2 from Homo sapiens, Northeast Structural Genomics Consortium Target HR6741A | | Descriptor: | AFG3-like protein 2 | | Authors: | Ramelot, T.A, Yang, Y, Lee, H, Janua, H, Kohan, E, Shastry, R, Acton, T.B, Xiao, R, Everett, J.K, Prestegard, J.H, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG), Mitochondrial Protein Partnership (MPP) | | Deposit date: | 2011-12-20 | | Release date: | 2012-01-25 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | NMR structure and MD simulations of the AAA protease intermembrane space domain indicates peripheral membrane localization within the hexaoligomer.
Febs Lett., 587, 2013
|
|
2QW8
 
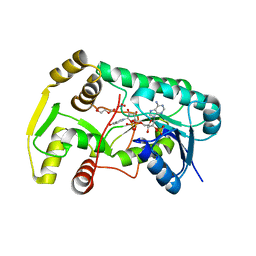 | | Structure of Eugenol Synthase from Ocimum basilicum | | Descriptor: | DI(HYDROXYETHYL)ETHER, Eugenol synthase 1, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Louie, G.V, Noel, J.P. | | Deposit date: | 2007-08-09 | | Release date: | 2008-06-10 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structure and reaction mechanism of basil eugenol synthase.
Plos One, 2, 2007
|
|
2FC7
 | | Solution structure of the ZZ domain of ZZZ3 protein | | Descriptor: | ZINC ION, ZZZ3 protein | | Authors: | Dang, W, Muto, Y, Inoue, M, Shirouzu, M, Terada, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-12-12 | | Release date: | 2006-06-12 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the ZZ domain of ZZZ3 protein
To be published
|
|
3LCY
 
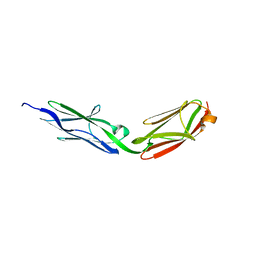 | |
2QYM
 
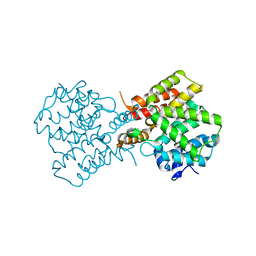 | | crystal structure of unliganded PDE4C2 | | Descriptor: | MAGNESIUM ION, Phosphodiesterase 4C, ZINC ION | | Authors: | Ke, H. | | Deposit date: | 2007-08-15 | | Release date: | 2008-04-08 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structures of the four subfamilies of phosphodiesterase-4 provide insight into the selectivity of their inhibitors.
Biochem.J., 408, 2007
|
|
2R2G
 
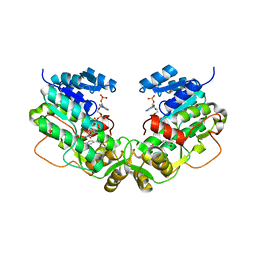 | | Structure of Eugenol Synthase from Ocimum basilicum complexed with EMDF | | Descriptor: | Eugenol synthase 1, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ethyl (1S,2S)-2-(4-hydroxy-3-methoxyphenyl)cyclopropanecarboxylate | | Authors: | Louie, G.V, Noel, J.P, Bowman, M.E. | | Deposit date: | 2007-08-24 | | Release date: | 2008-01-15 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure and reaction mechanism of basil eugenol synthase
Plos One, 2, 2007
|
|
1WVO
 | | Solution structure of RSGI RUH-029, an antifreeze protein like domain in human N-acetylneuraminic acid phosphate synthase gene. | | Descriptor: | Sialic acid synthase | | Authors: | Ito, Y, Hamada, T, Hayashi, F, Yokoyama, S, Hirota, H, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-12-22 | | Release date: | 2006-01-03 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the antifreeze-like domain of human sialic acid synthase
Protein Sci., 15, 2006
|
|
2GBX
 
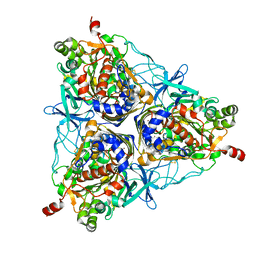 | | Crystal Structure of Biphenyl 2,3-Dioxygenase from Sphingomonas yanoikuyae B1 Bound to Biphenyl | | Descriptor: | BIPHENYL, Biphenyl 2,3-Dioxygenase Alpha Subunit, Biphenyl 2,3-Dioxygenase Beta Subunit, ... | | Authors: | Ferraro, D.J, Brown, E.N, Yu, C, Parales, R.E, Gibson, D.T, Ramaswamy, S. | | Deposit date: | 2006-03-12 | | Release date: | 2007-03-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural investigations of the ferredoxin and terminal oxygenase components of the biphenyl 2,3-dioxygenase from Sphingobium yanoikuyae B1.
Bmc Struct.Biol., 7, 2007
|
|
2LJB
 
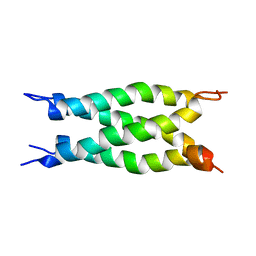 | |
3U46
 
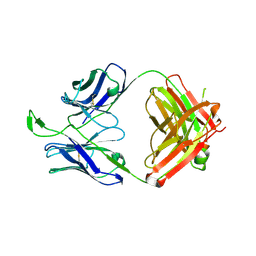 | | CH04H/CH02L P212121 | | Descriptor: | CH02 Light chain Fab, CH04 Heavy chain Fab | | Authors: | Louder, R, Pancera, M, McLellan, J.S, Kwong, P.D. | | Deposit date: | 2011-10-07 | | Release date: | 2011-11-30 | | Last modified: | 2011-12-21 | | Method: | X-RAY DIFFRACTION (2.906 Å) | | Cite: | Structure of HIV-1 gp120 V1/V2 domain with broadly neutralizing antibody PG9.
Nature, 480, 2011
|
|
1QI6
 
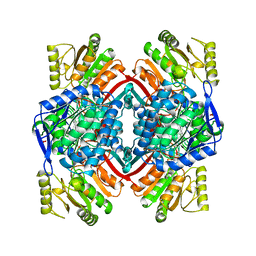 | | SECOND APO FORM OF AN NADP DEPENDENT ALDEHYDE DEHYDROGENASE WITH GLU250 SITUATED 3.7 A FROM CYS284 | | Descriptor: | PROTEIN (NADP DEPENDENT NONPHOSPHORYLATING GLYCERALDEHYDE-3-PHOSPHATE DEHYDROGENASE), SULFATE ION | | Authors: | Cobessi, D, Tete-Favier, F, Marchal, S, Branlant, G, Aubry, A. | | Deposit date: | 1999-06-02 | | Release date: | 2001-01-10 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural and biochemical investigations of the catalytic mechanism of an NADP-dependent aldehyde dehydrogenase from Streptococcus mutans.
J.Mol.Biol., 300, 2000
|
|
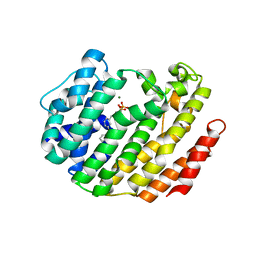
3LEE
 
 | |
3PFT
 
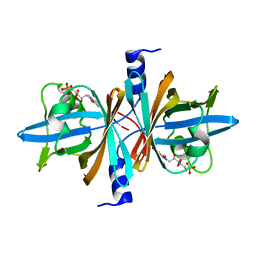 | | Crystal Structure of Untagged C54A Mutant Flavin Reductase (DszD) in Complex with FMN From Mycobacterium goodii | | Descriptor: | FLAVIN MONONUCLEOTIDE, Flavin reductase | | Authors: | Li, Q, Xu, P, Ma, C, Gu, L, Liu, X, Zhang, C, Li, N, Su, J, Li, B, Liu, S. | | Deposit date: | 2010-10-29 | | Release date: | 2011-11-02 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.601 Å) | | Cite: | The flavin reductase DSZD from a desulfurizing mycobacterium goodii strain: systemic manipulation and investigation based on the crystal structure
To be Published
|
|
2QX7
 
 | |
2LIV
 
 | |
2GFB
 
 | |
3LCJ
 
 | | Phosphopantetheine adenylyltransferase from Mycobacterium tuberculosis complexed with CoA | | Descriptor: | COENZYME A, Phosphopantetheine adenylyltransferase | | Authors: | Timofeev, V.I, Smirnova, E.A, Chupova, L.A, Esipov, R.S, Kuranova, I.P. | | Deposit date: | 2010-01-11 | | Release date: | 2010-07-21 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Preparation of the Crystal Complex of Phosphopantetheine
Adenylyltransferase from Mycobacterium tuberculosis
with Coenzyme A and Investigation of Its Three-Dimensional Structure at 2.1-A frame Resolution
CRYSTALLOGRAPHY REPORTS, 55, 2010
|
|
2GR2
 
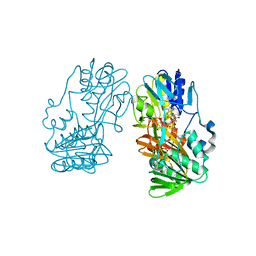 | |
2RLT
 
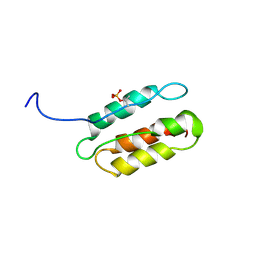 | | phosphorylated CPI-17 (22-120) | | Descriptor: | Protein phosphatase 1 regulatory subunit 14A | | Authors: | Eto, M. | | Deposit date: | 2007-08-11 | | Release date: | 2008-07-15 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Phosphorylation-induced conformational switching of CPI-17 produces a potent myosin phosphatase inhibitor.
Structure, 15, 2007
|
|
2BV7
 
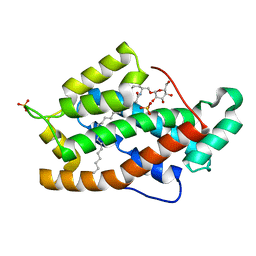 | | Crystal structure of GLTP with bound GM3 | | Descriptor: | GLYCOLIPID TRANSFER PROTEIN, N-{1-[(HEXOPYRANOSYLOXY)METHYL]-2-HYDROXYNONADECYL}TETRACOSANAMIDE, SULFATE ION | | Authors: | Kidron, H, Airenne, T.T, Nymalm, Y, Nylund, M, West, G, Mattjus, P, Salminen, T.A. | | Deposit date: | 2005-06-23 | | Release date: | 2005-12-14 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Structural Evidence for Adaptive Ligand Binding of Glycolipid Transfer Protein.
J.Mol.Biol., 355, 2006
|
|
1WWH
 
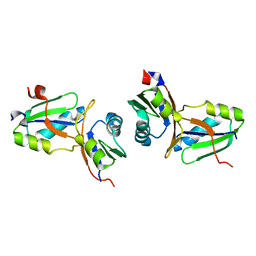 | | Crystal structure of the MPPN domain of mouse Nup35 | | Descriptor: | nucleoporin 35 | | Authors: | Handa, N, Murayama, K, Kukimoto, M, Hamana, H, Uchikubo, T, Takemoto, C, Terada, T, Shirouzu, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-01-05 | | Release date: | 2005-07-05 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | The crystal structure of mouse Nup35 reveals atypical RNP motifs and novel homodimerization of the RRM domain
J.Mol.Biol., 363, 2006
|
|
3CHG
 
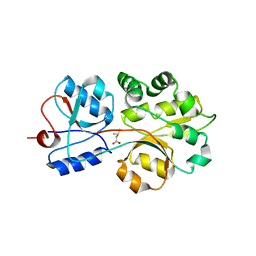 | | The compatible solute-binding protein OpuAC from Bacillus subtilis in complex with DMSA | | Descriptor: | (dimethyl-lambda~4~-sulfanyl)acetic acid, Glycine betaine-binding protein | | Authors: | Smits, S.H.J, Hoing, M, Lecher, J, Jebbar, M, Schmitt, L, Bremer, E. | | Deposit date: | 2008-03-09 | | Release date: | 2008-08-12 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | The Compatible-Solute-Binding Protein OpuAC from Bacillus subtilis: Ligand Binding, Site-Directed Mutagenesis, and Crystallographic Studies
J.Bacteriol., 190, 2008
|
|
7EEY
 
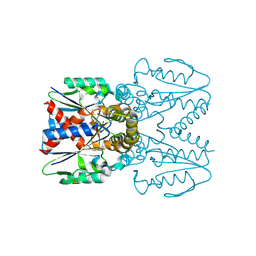 | |
2TPK
 
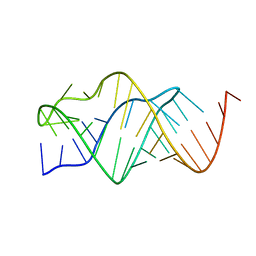 | |
