8ZLZ
 
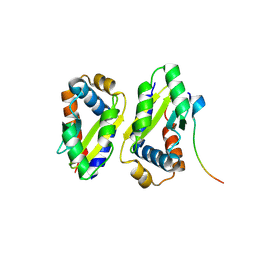 | |
1B6E
 
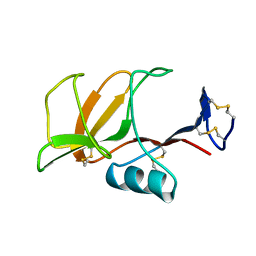 | | HUMAN CD94 | | Descriptor: | CD94 | | Authors: | Boyington, J.C, Riaz, A.N, Patamawenu, A, Coligan, J.E, Brooks, A.G, Sun, P.D. | | Deposit date: | 1999-01-14 | | Release date: | 1999-06-15 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of CD94 reveals a novel C-type lectin fold: implications for the NK cell-associated CD94/NKG2 receptors.
Immunity, 10, 1999
|
|
9DOY
 
 | | Fibrillar assembly of an D-enantiopure, C-alpha methylated, macrocyclic beta-hairpin | | Descriptor: | 5-amino-2,4,6-triiodobenzene-1,3-dicarboxylic acid, D-enantiopure, C-alpha methylated, ... | | Authors: | Samdin, T.D, Lubkowski, J, Anderson, C.F, Schneider, J.P. | | Deposit date: | 2024-09-20 | | Release date: | 2025-04-30 | | Last modified: | 2025-05-07 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | From Hydrogel to Crystal: A Molecular Design Strategy that Chemically Modifies Racemic Gel-Forming Peptides to Furnish Crystalline Fibrils Stabilized by Parallel Rippled beta-Sheets.
J.Am.Chem.Soc., 147, 2025
|
|
8QVZ
 
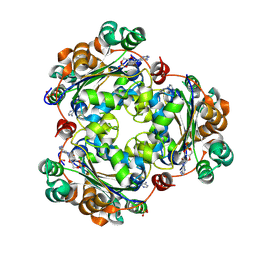 | | Human NDPK-C in complex with cAMP | | Descriptor: | ADENOSINE-3',5'-CYCLIC-MONOPHOSPHATE, Nucleoside diphosphate kinase 3, SULFATE ION | | Authors: | Werten, S, Amjadi, R, Scheffzek, K. | | Deposit date: | 2023-10-18 | | Release date: | 2024-09-18 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.774 Å) | | Cite: | Mechanistic Insights into Substrate Recognition of Human Nucleoside Diphosphate Kinase C Based on Nucleotide-Induced Structural Changes.
Int J Mol Sci, 25, 2024
|
|
8QW3
 
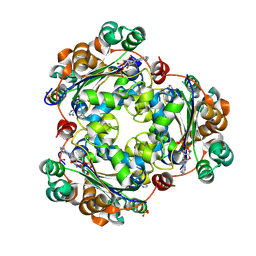 | | Human NDPK-C in complex with ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Nucleoside diphosphate kinase 3, SULFATE ION | | Authors: | Werten, S, Amjadi, R, Scheffzek, K. | | Deposit date: | 2023-10-18 | | Release date: | 2024-09-18 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.255 Å) | | Cite: | Mechanistic Insights into Substrate Recognition of Human Nucleoside Diphosphate Kinase C Based on Nucleotide-Induced Structural Changes.
Int J Mol Sci, 25, 2024
|
|
8QW0
 
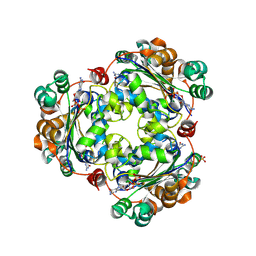 | | Human NDPK-C in complex with GDP | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, Nucleoside diphosphate kinase 3, SULFATE ION | | Authors: | Werten, S, Amjadi, R, Scheffzek, K. | | Deposit date: | 2023-10-18 | | Release date: | 2024-09-18 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.175 Å) | | Cite: | Mechanistic Insights into Substrate Recognition of Human Nucleoside Diphosphate Kinase C Based on Nucleotide-Induced Structural Changes.
Int J Mol Sci, 25, 2024
|
|
9F2H
 
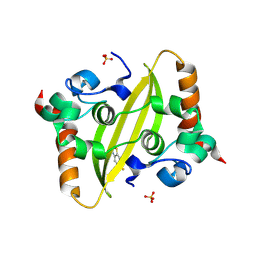 | | Crystal structure of SARS-CoV-2 N-protein C-terminal domain in complex with riluzole | | Descriptor: | 1,2-ETHANEDIOL, 6-(trifluoromethoxy)-1,3-benzothiazol-2-amine, Nucleoprotein, ... | | Authors: | Marquez-Monino, M.A, Gonzalez, B, Perez-Canadillas, J.M. | | Deposit date: | 2024-04-23 | | Release date: | 2024-11-13 | | Last modified: | 2025-01-15 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | The ALS drug riluzole binds to the C-terminal domain of SARS-CoV-2 nucleocapsid protein and has antiviral activity.
Structure, 33, 2025
|
|
6NZ2
 
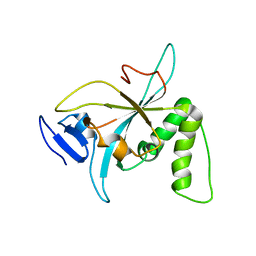 | |
3FD7
 
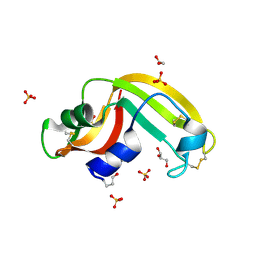 | | Crystal structure of Onconase C87A/C104A-ONC | | Descriptor: | 1,2-ETHANEDIOL, GLYCEROL, Protein P-30, ... | | Authors: | Neumann, P, Schulenburg, C, Arnold, U, Ulbrich-Hofmann, R, Stubbs, M.T. | | Deposit date: | 2008-11-25 | | Release date: | 2009-12-08 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.531 Å) | | Cite: | Impact of the C-terminal disulfide bond on the folding and stability of onconase.
Chembiochem, 11, 2010
|
|
9FBK
 
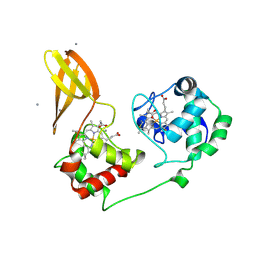 | |
6NSV
 
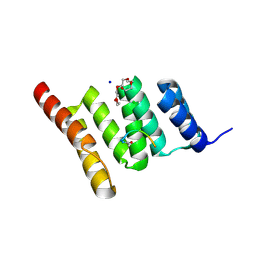 | | Crystal structure of the human CHIP TPR domain in complex with a 5mer acetylated optimized peptide | | Descriptor: | ACE-LEU-TRP-TRP-PRO-ASP, CHLORIDE ION, E3 ubiquitin-protein ligase CHIP, ... | | Authors: | Basu, K, Ravalin, M, Bohn, M.-F, Craik, C.S, Gestwicki, J.E. | | Deposit date: | 2019-01-25 | | Release date: | 2019-07-31 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.305 Å) | | Cite: | Specificity for latent C termini links the E3 ubiquitin ligase CHIP to caspases.
Nat.Chem.Biol., 15, 2019
|
|
9ESA
 
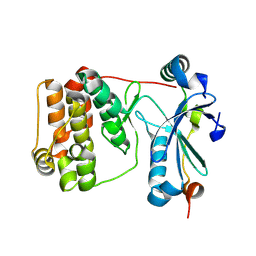 | | Aurora-C with SER mutation in complex with INCENP peptide | | Descriptor: | 1,2-ETHANEDIOL, Aurora kinase C, Inner centromere protein | | Authors: | Hillig, R.C. | | Deposit date: | 2024-03-26 | | Release date: | 2024-09-11 | | Last modified: | 2024-09-25 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Surface-mutagenesis strategies to enable structural biology crystallization platforms.
Acta Crystallogr D Struct Biol, 80, 2024
|
|
6T6J
 
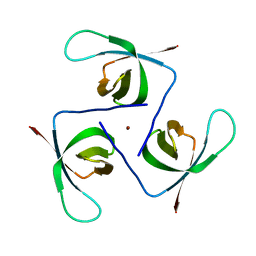 | |
5XDM
 
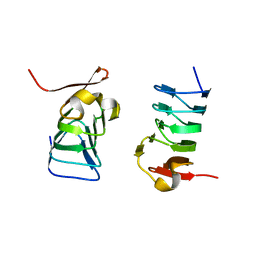 | |
5GU4
 
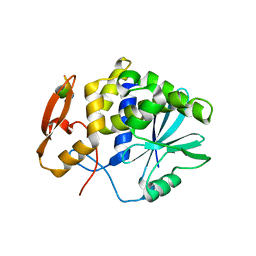 | | rRNA N-glycosylase RTA | | Descriptor: | GLY-PHE-GLY-LEU-PHE-ASP, GLYCEROL, Ricin | | Authors: | Shi, W.W, Tang, Y.S, Sze, S.Y, Zhu, Z.N, Wong, K.B, Shaw, P.C. | | Deposit date: | 2016-08-24 | | Release date: | 2016-11-02 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Crystal Structure of Ribosome-Inactivating Protein Ricin A Chain in Complex with the C-Terminal Peptide of the Ribosomal Stalk Protein P2
Toxins, 8, 2016
|
|
1GDF
 
 | |
6U73
 
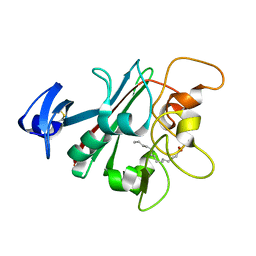 | | Human Angiopoietin-Like 4 C-Terminal Domain (cANGPTL4) with Myristic Acid | | Descriptor: | Angiopoietin-related protein 4, MYRISTIC ACID | | Authors: | Tarver, C.L, Yuan, Q, Singhal, A.J, Ramaker, R, Cooper, S, Pusey, M.L. | | Deposit date: | 2019-08-31 | | Release date: | 2019-10-09 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.38 Å) | | Cite: | Crystal Structures of Angiopoietin-Like 4s C-Terminal Domain (cANGPTL4) Reveal a Binding Site for Various Ligands
To Be Published
|
|
6U0A
 
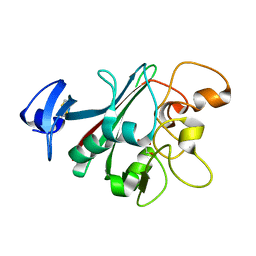 | | Human Angiopoietin-Like 4 C-Terminal Domain (cANGPTL4) with Glycerol | | Descriptor: | Angiopoietin-related protein 4, GLYCEROL | | Authors: | Tarver, C.L, Yuan, Q, Singhal, A.J, Ramaker, R, Cooper, S, Pusey, M.L. | | Deposit date: | 2019-08-14 | | Release date: | 2019-10-09 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | Crystal Structures of Angiopoietin-Like 4 C-Terminal Domain (cANGPTL4) Reveal a Binding Pocket with Multiple Ligands
To Be Published
|
|
5KHI
 
 | | HCN2 CNBD in complex with purine riboside-3', 5'-cyclic monophosphate (cPuMP) | | Descriptor: | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2, Purine riboside-3',5'-cyclic monophosphate | | Authors: | Ng, L.C.T, Putrenko, I, Baronas, V, Van Petegem, F, Accili, E.A. | | Deposit date: | 2016-06-14 | | Release date: | 2016-09-14 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Cyclic Purine and Pyrimidine Nucleotides Bind to the HCN2 Ion Channel and Variably Promote C-Terminal Domain Interactions and Opening.
Structure, 24, 2016
|
|
5KHJ
 
 | | HCN2 CNBD in complex with uridine-3', 5'-cyclic monophosphate (cUMP) | | Descriptor: | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2, Uridine-3',5'-cyclic monophosphate | | Authors: | Ng, L.C.T, Putrenko, I, Baronas, V, Van Petegem, F, Accili, E.A. | | Deposit date: | 2016-06-14 | | Release date: | 2016-09-14 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | Cyclic Purine and Pyrimidine Nucleotides Bind to the HCN2 Ion Channel and Variably Promote C-Terminal Domain Interactions and Opening.
Structure, 24, 2016
|
|
5KHK
 
 | | HCN2 CNBD in complex with 2-aminopurine riboside-3', 5'-cyclic monophosphate (2-NH2-cPuMP) | | Descriptor: | 2-Aminopurine riboside-3',5'-cyclic monophosphate, Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 | | Authors: | Ng, L.C.T, Putrenko, I, Baronas, V, Van Petegem, F, Accili, E.A. | | Deposit date: | 2016-06-14 | | Release date: | 2016-09-14 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.07 Å) | | Cite: | Cyclic Purine and Pyrimidine Nucleotides Bind to the HCN2 Ion Channel and Variably Promote C-Terminal Domain Interactions and Opening.
Structure, 24, 2016
|
|
6U1U
 
 | | Human Angiopoietin-Like 4 C-Terminal Domain (cANGPTL4) with Palmitic Acid | | Descriptor: | Angiopoietin-related protein 4, PALMITIC ACID | | Authors: | Tarver, C.L, Yuan, Q, Singhal, A.J, Ramaker, R, Cooper, S, Pusey, M.L. | | Deposit date: | 2019-08-16 | | Release date: | 2019-10-09 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Crystal Structures of Angiopoietin-Like 4 C-Terminal Domain (cANGPTL4) Reveal a Binding Pocket with Multiple Ligands
To Be Published
|
|
5KHG
 
 | | HCN2 CNBD in complex with cytidine-3', 5'-cyclic monophosphate (cCMP) | | Descriptor: | 4-amino-1-[(2S,4aR,6R,7R,7aS)-2,7-dihydroxy-2-oxidotetrahydro-4H-furo[3,2-d][1,3,2]dioxaphosphinin-6-yl]pyrimidin-2(1H)-one, Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 | | Authors: | Ng, L.C.T, Putrenko, I, Baronas, V, Van Petegem, F, Accili, E.A. | | Deposit date: | 2016-06-14 | | Release date: | 2016-09-14 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.241 Å) | | Cite: | Cyclic Purine and Pyrimidine Nucleotides Bind to the HCN2 Ion Channel and Variably Promote C-Terminal Domain Interactions and Opening.
Structure, 24, 2016
|
|
5E6N
 
 | | Crystal structure of C. elegans LGG-2 | | Descriptor: | Protein lgg-2 | | Authors: | Qi, X, Ren, J.Q, Wu, F, Zhang, H, Feng, W. | | Deposit date: | 2015-10-10 | | Release date: | 2016-01-06 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.102 Å) | | Cite: | Structural Basis of the Differential Function of the Two C. elegans Atg8 Homologs, LGG-1 and LGG-2, in Autophagy
Mol.Cell, 60, 2015
|
|
5KHH
 
 | | HCN2 CNBD in complex with inosine-3', 5'-cyclic monophosphate (cIMP) | | Descriptor: | Inosine-3',5'-cyclic monophosphate, Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 2 | | Authors: | Ng, L.C.T, Putrenko, I, Baronas, V, Van Petegem, F, Accili, E.A. | | Deposit date: | 2016-06-14 | | Release date: | 2016-09-14 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Cyclic Purine and Pyrimidine Nucleotides Bind to the HCN2 Ion Channel and Variably Promote C-Terminal Domain Interactions and Opening.
Structure, 24, 2016
|
|
