1GW4
 
 | |
1GUC
 
 | |
1QHK
 
 | |
1QQV
 
 | |
1HT7
 
 | |
1RVS
 
 | | STRUCTURE OF TRANSTHYRETIN IN AMYLOID FIBRILS DETERMINED BY SOLID-STATE MAGIC ANGLE SPINNING NMR | | Descriptor: | Transthyretin | | Authors: | Jaroniec, C.P, Macphee, C.E, Bajaj, V.S, Mcmahon, M.T, Dobson, C.M, Griffin, R.G. | | Deposit date: | 2003-12-14 | | Release date: | 2004-01-20 | | Last modified: | 2024-05-01 | | Method: | SOLID-STATE NMR | | Cite: | High-resolution molecular structure of a peptide in an amyloid fibril determined by magic angle spinning NMR spectroscopy
Proc.Natl.Acad.Sci.USA, 101, 2004
|
|
1HT4
 
 | |
1RO3
 
 | | New structural insights on short disintegrin echistatin by NMR | | Descriptor: | Disintegrin echistatin | | Authors: | Monleon, D, Esteve, V, Calvete, J.J, Marcinkiewicz, C, Celda, B. | | Deposit date: | 2003-12-01 | | Release date: | 2003-12-09 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Conformation and concerted dynamics of the integrin-binding site and the C-terminal region of echistatin revealed by homonuclear NMR
Biochem.J., 387, 2005
|
|
1SHP
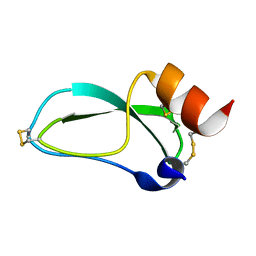 
 | | THE NMR SOLUTION STRUCTURE OF A KUNITZ-TYPE PROTEINASE INHIBITOR FROM THE SEA ANEMONE STICHODACTYLA HELIANTHUS | | Descriptor: | TRYPSIN INHIBITOR | | Authors: | Antuch, W, Berndt, K, Chavez, M, Delfin, J, Wuthrich, K. | | Deposit date: | 1992-11-17 | | Release date: | 1994-01-31 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | The NMR solution structure of a Kunitz-type proteinase inhibitor from the sea anemone Stichodactyla helianthus.
Eur.J.Biochem., 212, 1993
|
|
1RCK
 
 | |
1R73
 
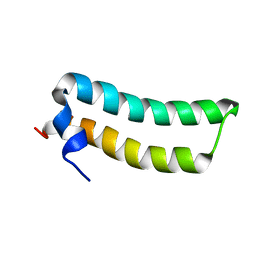 | | Solution Structure of TM1492, the L29 ribosomal protein from Thermotoga maritima | | Descriptor: | 50S ribosomal protein L29 | | Authors: | Peti, W, Etezady-Esfarjani, T, Herrmann, T, Klock, H.E, Lesley, S.A, Wuethrich, K, Joint Center for Structural Genomics (JCSG) | | Deposit date: | 2003-10-17 | | Release date: | 2004-08-10 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | NMR for structural proteomics of Thermotoga maritima: Screening and structure determination
J.STRUCT.FUNCT.GENOM., 5, 2004
|
|
1RCL
 
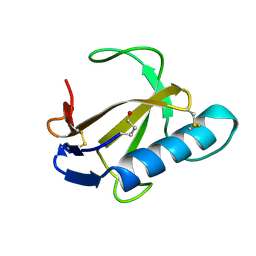 | |
1W0B
 
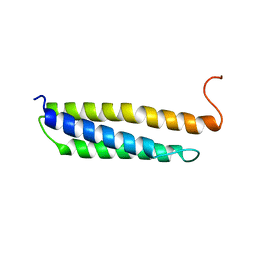 | | Solution structure of the human alpha-hemoglobin stabilizing protein (AHSP) P30A mutant | | Descriptor: | ALPHA-HEMOGLOBIN STABILIZING PROTEIN | | Authors: | Santiveri, C.M, Perez-Canadillas, J.M, Vadivelu, M.K, Allen, M.D, Rutherford, T.J, Watkins, N.A, Bycroft, M. | | Deposit date: | 2004-06-01 | | Release date: | 2004-06-10 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the alpha-hemoglobin stabilizing protein: insights into conformational heterogeneity and binding.
J. Biol. Chem., 279, 2004
|
|
1W0A
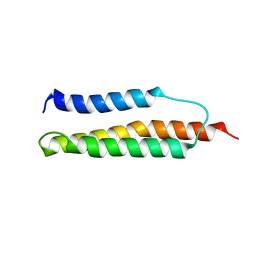 
 | | Solution structure of the trans form of the human alpha-hemoglobin stabilizing protein (AHSP) | | Descriptor: | ALPHA-HEMOGLOBIN STABILIZING PROTEIN | | Authors: | Santiveri, C.M, Perez-Canadillas, J.M, Vadivelu, M.K, Allen, M.D, Rutherford, T.J, Watkins, N.A, Bycroft, M. | | Deposit date: | 2004-06-01 | | Release date: | 2004-06-10 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the alpha-hemoglobin stabilizing protein: insights into conformational heterogeneity and binding.
J. Biol. Chem., 279, 2004
|
|
1DLZ
 
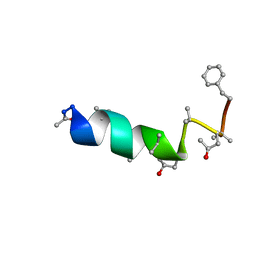 | | SOLUTION STRUCTURE OF THE CHANNEL-FORMER ZERVAMICIN IIB (PEPTAIBOL ANTIBIOTIC) | | Descriptor: | ZERVAMICIN IIB | | Authors: | Balashova, T.A, Shenkarev, Z.O, Tagaev, A.A, Ovchinnikova, T.V, Raap, J, Arseniev, A.S. | | Deposit date: | 1999-12-13 | | Release date: | 2000-02-03 | | Last modified: | 2012-12-12 | | Method: | SOLUTION NMR | | Cite: | NMR Structure of the Channel-Former Zervamicin Iib in Isotropic Solvents.
FEBS Lett., 466, 2000
|
|
1J3G
 
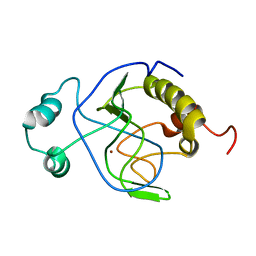 | | Solution structure of Citrobacter Freundii AmpD | | Descriptor: | AmpD protein, ZINC ION | | Authors: | Liepinsh, E, Genereux, C, Dehareng, D, Joris, B, Otting, G. | | Deposit date: | 2003-01-31 | | Release date: | 2003-02-18 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | NMR Structure of Citrobacter freundii AmpD, Comparison with Bacteriophage T7 Lysozyme
and Homology with PGRP Domains
J.Mol.Biol., 327, 2003
|
|
1GM0
 
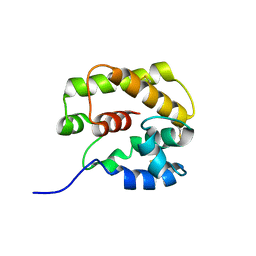 | | A Form of the Pheromone-Binding Protein from Bombyx mori | | Descriptor: | PHEROMONE-BINDING PROTEIN | | Authors: | Horst, R, Damberger, F, Guntert, P, Luginbuhl, P, Nikonova, L, Peng, G, Leal, W.S, Wuthrich, K. | | Deposit date: | 2001-09-05 | | Release date: | 2001-11-30 | | Last modified: | 2018-01-17 | | Method: | SOLUTION NMR | | Cite: | NMR Structure Reveals Novel Intramolecular Regulation Mechanism for Pheromone-Binding and Release
Proc.Natl.Acad.Sci.USA, 98, 2001
|
|
1D8Z
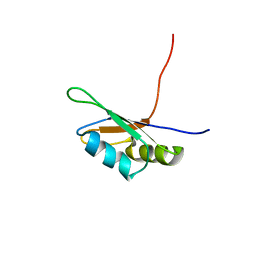 
 | | SOLUTION STRUCTURE OF THE FIRST RNA-BINDING DOMAIN (RBD1) OF HU ANTIGEN C (HUC) | | Descriptor: | HU ANTIGEN C | | Authors: | Inoue, M, Muto, Y, Sakamoto, H, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 1999-10-26 | | Release date: | 2000-04-07 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | NMR studies on functional structures of the AU-rich element-binding domains of Hu antigen C.
Nucleic Acids Res., 28, 2000
|
|
1IMI
 
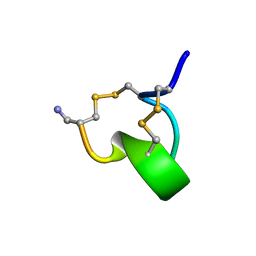 | | SOLUTION STRUCTURE OF ALPHA-CONOTOXIN IM1 | | Descriptor: | PROTEIN (ALPHA-CONOTOXIN IMI) | | Authors: | Maslennikov, I.V, Shenkarev, Z.O, Zhmak, M.N, Tsetlin, V.I, Ivanov, V.T, Arseniev, A.S. | | Deposit date: | 1998-11-27 | | Release date: | 1999-04-23 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | NMR spatial structure of alpha-conotoxin ImI reveals a common scaffold in snail and snake toxins recognizing neuronal nicotinic acetylcholine receptors.
FEBS Lett., 444, 1999
|
|
1HA8
 
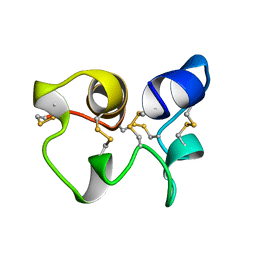 | |
1R3B
 
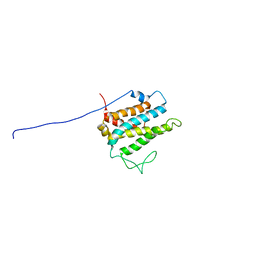 | | Solution structure of xenopus laevis Mob1 | | Descriptor: | MOB1 | | Authors: | Ponchon, L, Dumas, C, Kajava, A.V, Fesquet, D, Padilla, A. | | Deposit date: | 2003-10-01 | | Release date: | 2004-09-28 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | NMR solution structure of Mob1, a mitotic exit network protein and its interaction with an NDR kinase peptide
J.Mol.Biol., 337, 2004
|
|
1RHX
 
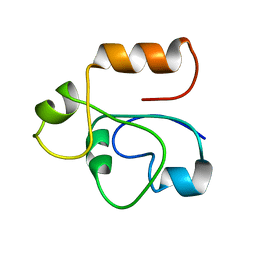 | |
1T5Q
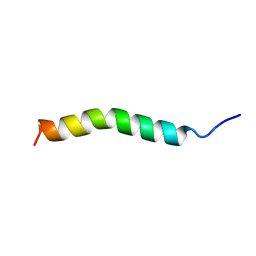 
 | | Solution Structure of GIP(1-30)amide in TFE/Water | | Descriptor: | Gastric inhibitory polypeptide | | Authors: | Alana, I, Hewage, C.M, Malthouse, J.P.G, Parker, J.C, Gault, V.A, O'Harte, F.P.M. | | Deposit date: | 2004-05-05 | | Release date: | 2004-11-16 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the glucose-dependent insulinotropic polypeptide fragment, GIP(1-30)amide.
Biochem.Biophys.Res.Commun., 325, 2004
|
|
1SKH
 
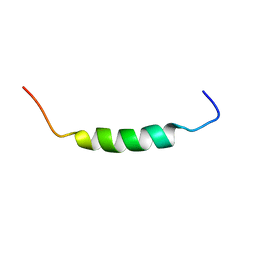 | |
1SMZ
 
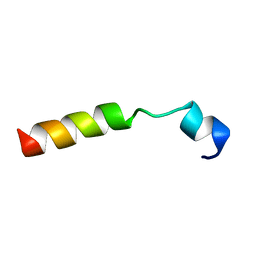 | |
