8UZL
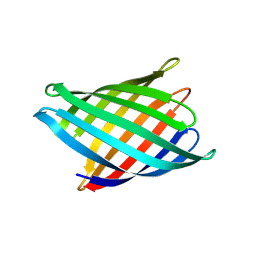 
 | | Designed Transmembrane beta-barrel- TMB10_163 | | Descriptor: | Designed Transmembrane beta-barrel TMB10_163, HEXANE-1,6-DIOL | | Authors: | Bera, A.K, Lemma, S.B, Kang, A, Baker, D. | | Deposit date: | 2023-11-15 | | Release date: | 2024-07-17 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Sculpting conducting nanopore size and shape through de novo protein design
Science, 2024
|
|
6VDP
 
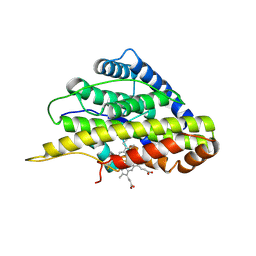 | | Crystal structure of SfmD truncated variant | | Descriptor: | 3-methyl-L-tyrosine peroxygenase, HEME C | | Authors: | Shin, I, Liu, A. | | Deposit date: | 2019-12-27 | | Release date: | 2021-03-10 | | Last modified: | 2021-07-14 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A novel catalytic heme cofactor in SfmD with a single thioether bond and a bis -His ligand set revealed by a de novo crystal structural and spectroscopic study.
Chem Sci, 12, 2021
|
|
6VDZ
 
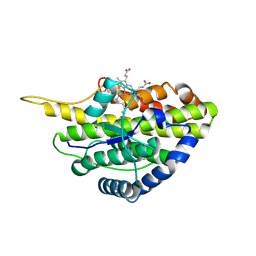 | | Crystal structure of reduced SfmD by soaking with sodium hydrosulfite | | Descriptor: | 3-methyl-L-tyrosine peroxygenase, HEME C | | Authors: | Shin, I, Liu, A. | | Deposit date: | 2019-12-27 | | Release date: | 2021-03-10 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | A novel catalytic heme cofactor in SfmD with a single thioether bond and a bis -His ligand set revealed by a de novo crystal structural and spectroscopic study.
Chem Sci, 12, 2021
|
|
6VE0
 
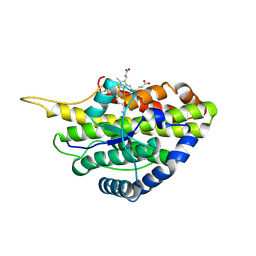 | | Crystal structure of reduced SfmD by soaking with sodium hydrosulfite | | Descriptor: | 3-methyl-L-tyrosine peroxygenase, HEME C | | Authors: | Shin, I, Liu, A. | | Deposit date: | 2019-12-27 | | Release date: | 2021-03-10 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3.15 Å) | | Cite: | A novel catalytic heme cofactor in SfmD with a single thioether bond and a bis -His ligand set revealed by a de novo crystal structural and spectroscopic study.
Chem Sci, 12, 2021
|
|
6VDQ
 
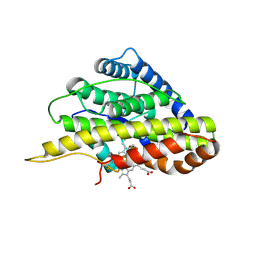 | | Crystal structure of SfmD | | Descriptor: | 3-methyl-L-tyrosine peroxygenase, HEME C | | Authors: | Shin, I, Liu, A. | | Deposit date: | 2019-12-27 | | Release date: | 2021-03-10 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | A novel catalytic heme cofactor in SfmD with a single thioether bond and a bis -His ligand set revealed by a de novo crystal structural and spectroscopic study.
Chem Sci, 12, 2021
|
|
8QAA
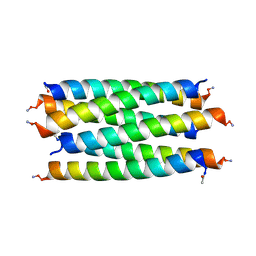 
 | |
8QAB
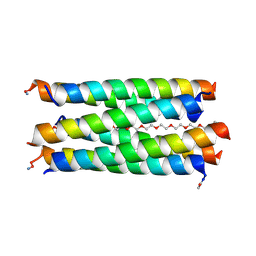 
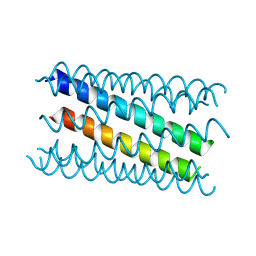
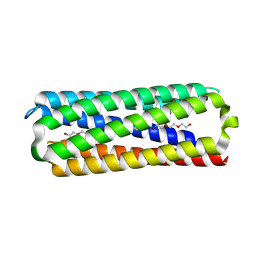
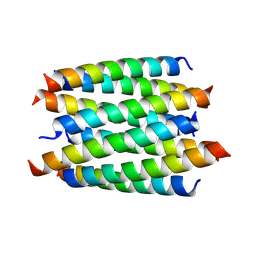
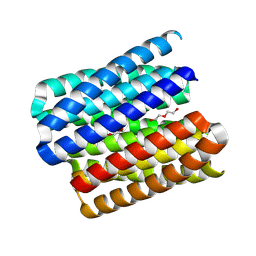
 | | X-ray crystal structure of a de novo designed antiparallel coiled-coil hexameric alpha-helical barrel with 4 heptad repeats, apCCHex | | Descriptor: | 1-methoxy-2-[2-[2-[2-[2-[2-[2-[2-[2-[2-(2-methoxyethoxy)ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethoxy]ethane, apCCHex | | Authors: | Naudin, E.N, Albanese, K.I, Petrenas, R, Woolfson, D.N. | | Deposit date: | 2023-08-22 | | Release date: | 2024-07-03 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Rationally seeded computational protein design of ɑ-helical barrels.
Nat.Chem.Biol., 2024
|
|
8QAI
 | |
8QAH
 | |
8QKD
 | |
8QAD
 | |
8QAF
 | |
8QAE
 | |
8QAC
 | |
8QAG
 | |
4D8H
 
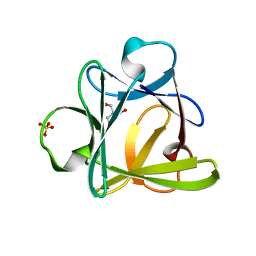 | | Crystal structure of Symfoil-4P/PV2: de novo designed beta-trefoil architecture with symmetric primary structure, primitive version 2 (6xLeu / PV1) | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, SULFATE ION, de novo protein | | Authors: | Blaber, M, Longo, L. | | Deposit date: | 2012-01-10 | | Release date: | 2013-01-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.901 Å) | | Cite: | Simplified protein design biased for prebiotic amino acids yields a foldable, halophilic protein.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
7UBF
 
 | |
7UBD
 
 | |
7UBI
 
 | |
7UZL
 
 | |
7UBC
 
 | |
7UBE
 
 | |
7UBG
 
 | |
7UBH
 
 | |
1KYC
 
 | |
