1Y0Y
 
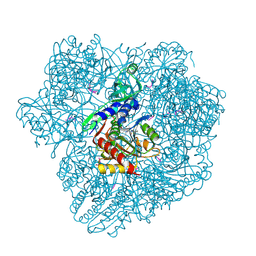 | |
1YF1
 
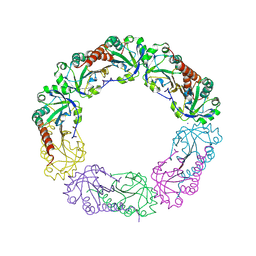 | | Structural and biochemical analysis of the link between enzymatic activity and oligomerization in AhpC, a bacterial peroxiredoxin. | | Descriptor: | Alkyl hydroperoxide reductase subunit C, SODIUM ION | | Authors: | Parsonage, D, Youngblood, D.S, Sarma, G.N, Wood, Z.A, Karplus, P.A, Poole, L.B. | | Deposit date: | 2004-12-29 | | Release date: | 2005-08-16 | | Last modified: | 2024-11-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Analysis of the Link between Enzymatic Activity and Oligomeric State in AhpC, a Bacterial Peroxiredoxin.
Biochemistry, 44, 2005
|
|
1YF9
 
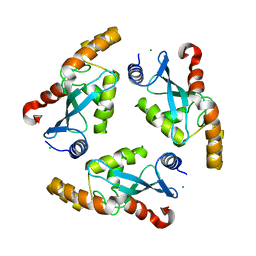 | |
1YHU
 
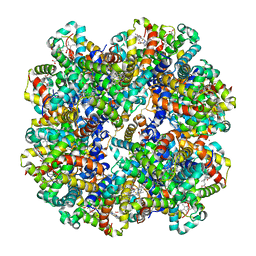 | | Crystal structure of Riftia pachyptila C1 hemoglobin reveals novel assembly of 24 subunits. | | Descriptor: | Giant hemoglobins B chain, OXYGEN MOLECULE, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Flores, J.F, Fisher, C.R, Carney, S.L, Green, B.N, Freytag, J.K, Schaeffer, S.W, Royer, W.E. | | Deposit date: | 2005-01-10 | | Release date: | 2005-02-08 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (3.15 Å) | | Cite: | Sulfide binding is mediated by zinc ions discovered in the crystal structure of a hydrothermal vent tubeworm hemoglobin.
Proc.Natl.Acad.Sci.Usa, 102, 2005
|
|
1Y18
 
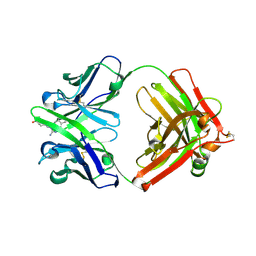 | | Fab fragment of catalytic elimination antibody 34E4 E(H50)D mutant in complex with hapten | | Descriptor: | 2-AMINO-5,6-DIMETHYL-BENZIMIDAZOLE-1-PENTANOIC ACID, CHLORIDE ION, Catalytic antibody 34E4 heavy chain, ... | | Authors: | Debler, E.W, Ito, S, Heine, A, Wilson, I.A. | | Deposit date: | 2004-11-17 | | Release date: | 2005-04-05 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural origins of efficient proton abstraction from carbon by a catalytic antibody
Proc.Natl.Acad.Sci.USA, 102, 2005
|
|
1Y20
 
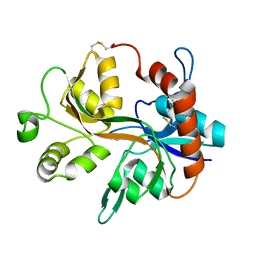 | |
1Y28
 
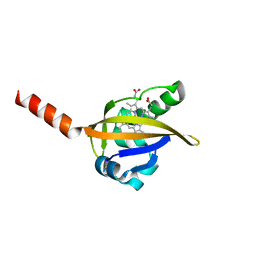 | | Crystal structure of the R220A metBJFIXL HEME domain | | Descriptor: | PROTOPORPHYRIN IX CONTAINING FE, Sensor protein fixL | | Authors: | Dunham, C.M, Dioum, E.M, Tuckerman, J.R, Gonzalez, G, Scott, W.G, Gilles-Gonzalez, M.A. | | Deposit date: | 2004-11-21 | | Release date: | 2004-12-07 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | A distal arginine in the oxygen-sensing heme-PAS domains is essential to ligand binding, signal transduction, and structure
Biochemistry, 42, 2003
|
|
1Y0E
 
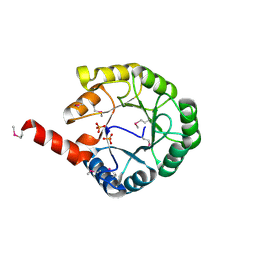 | |
4V6D
 
 | | Crystal structure of the E. coli 70S ribosome in an intermediate state of ratcheting | | Descriptor: | 16S rRNA, 23S rRNA, 30S ribosomal protein S10, ... | | Authors: | Zhang, W, Dunkle, J.A, Cate, J.H.D. | | Deposit date: | 2009-06-27 | | Release date: | 2014-07-09 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.814 Å) | | Cite: | Structures of the ribosome in intermediate States of ratcheting.
Science, 325, 2009
|
|
1Y12
 
 | |
1Y1O
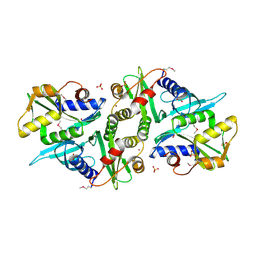 
 | | X-ray crystal Structure of Penicillin-binding protein-related factor A from Bacillus stearothermophilus | | Descriptor: | NICKEL (II) ION, Penicillin-binding protein-related factor A, SULFATE ION | | Authors: | Osipiuk, J, Li, H, Moy, S, Collart, F, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2004-11-19 | | Release date: | 2004-12-07 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | X-ray crystal Structure of Penicillin-binding protein-related factor A from Bacillus stearothermophilus
To be Published
|
|
4V7S
 
 | | Crystal structure of the E. coli ribosome bound to telithromycin. | | Descriptor: | 16S rRNA, 23S rRNA, 30S ribosomal protein S10, ... | | Authors: | Dunkle, J.A, Xiong, L, Mankin, A.S, Cate, J.H.D. | | Deposit date: | 2010-08-05 | | Release date: | 2014-07-09 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (3.2547 Å) | | Cite: | Structures of the Escherichia coli ribosome with antibiotics bound near the peptidyl transferase center explain spectra of drug action.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
1Y21
 
 | | Crystal Structure of Cimex Nitrophorin NO Complex | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, NITRIC OXIDE, PROTOPORPHYRIN IX CONTAINING FE, ... | | Authors: | Weichsel, A, Maes, E.M, Andersen, J.F, Valenzuela, J.G, Shokhireva, T.K, Walker, F.A, Montfort, W.R. | | Deposit date: | 2004-11-19 | | Release date: | 2004-11-30 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Heme-assisted S-nitrosation of a proximal thiolate in a nitric oxide transport protein.
Proc.Natl.Acad.Sci.Usa, 102, 2005
|
|
1Y2A
 
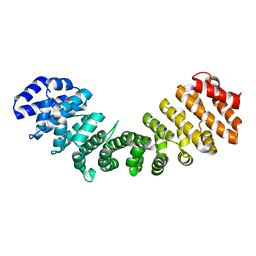 | | Structure of mammalian importin bound to the non-classical PLSCR1-NLS | | Descriptor: | Importin alpha-2 Subunit, decamer fragment of Phospholipid scramblase 1 | | Authors: | Chen, M.-H, Ben-Efraim, I, Mitrousis, G, Walker-Kopp, N, Sims, P.J, Cingolani, G. | | Deposit date: | 2004-11-22 | | Release date: | 2005-02-01 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Phospholipid Scramblase 1 Contains a Nonclassical Nuclear Localization Signal with Unique Binding Site in Importin alpha
J.Biol.Chem., 280, 2005
|
|
1Y2B
 
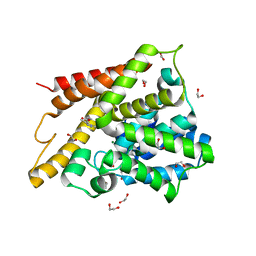 | | Catalytic Domain Of Human Phosphodiesterase 4D In Complex With 3,5-dimethyl-1H-pyrazole-4-carboxylic acid ethyl ester | | Descriptor: | 1,2-ETHANEDIOL, 3,5-DIMETHYL-1H-PYRAZOLE-4-CARBOXYLIC ACID ETHYL ESTER, MAGNESIUM ION, ... | | Authors: | Card, G.L, Blasdel, L, England, B.P, Zhang, C, Suzuki, Y, Gillette, S, Fong, D, Ibrahim, P.N, Artis, D.R, Bollag, G, Milburn, M.V, Kim, S.-H, Schlessinger, J, Zhang, K.Y.J. | | Deposit date: | 2004-11-22 | | Release date: | 2005-03-01 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | A family of phosphodiesterase inhibitors discovered by cocrystallography and scaffold-based drug design
Nat.Biotechnol., 23, 2005
|
|
1Y2O
 
 | | Structure of N-terminal domain IRSp53/BAIAP2 | | Descriptor: | BAI1-associated protein 2 isoform 1 | | Authors: | Millard, T.H, Bompard, G, Heung, M.-Y, Dafforn, T.R, Scott, D.J, Machesky, L.M, Futterer, K. | | Deposit date: | 2004-11-23 | | Release date: | 2005-02-15 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structural basis of filopodia formation induced by the IRSp53/MIM homology domain of human IRSp53
Embo J., 24, 2005
|
|
1Y30
 
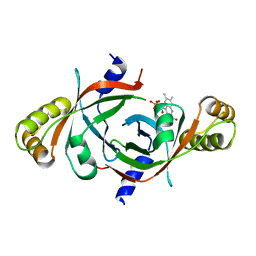 | | X-ray crystal structure of mycobacterium tuberculosis pyridoxine 5'-phosphate oxidase complexed with flavin mononucleotide at 2.2 a resolution | | Descriptor: | FLAVIN MONONUCLEOTIDE, hypothetical protein Rv1155 | | Authors: | Biswal, B.K, Cherney, M.M, Wang, M, Garen, C, James, M.N, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2004-11-23 | | Release date: | 2005-08-30 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structures of Mycobacterium tuberculosispyridoxine 5'-phosphate oxidase and its complexes with flavin mononucleotide and pyridoxal 5'-phosphate.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
1Y3A
 
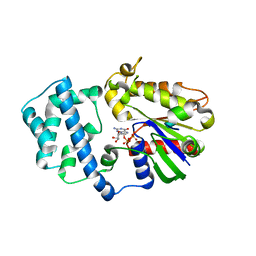 | | Structure of G-Alpha-I1 bound to a GDP-selective peptide provides insight into guanine nucleotide exchange | | Descriptor: | GUANOSINE-5'-DIPHOSPHATE, Guanine nucleotide-binding protein G(i), alpha-1 subunit, ... | | Authors: | Johnston, C.A, Willard, F.S, Jezyk, M.R, Fredericks, Z, Bodor, E.T, Jones, M.B, Blaesius, R, Harden, T.K, Sondek, J, Watts, V.J, Ramer, J.K, Siderovski, D.P. | | Deposit date: | 2004-11-24 | | Release date: | 2005-07-12 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of G-Alpha-I1 bound to a GDP-selective peptide provides insight into guanine nucleotide exchange
Structure, 7, 2005
|
|
1Y2T
 
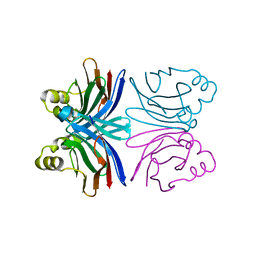 | | Crystal structure of the common edible mushroom (Agaricus bisporus) lectin | | Descriptor: | lectin | | Authors: | Carrizo, M.E, Capaldi, S, Perduca, M, Irazoqui, F.J, Nores, G.A, Monaco, H.L. | | Deposit date: | 2004-11-23 | | Release date: | 2004-12-21 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The Antineoplastic Lectin of the Common Edible Mushroom (Agaricus bisporus) Has Two Binding Sites, Each Specific for a Different Configuration at a Single Epimeric Hydroxyl
J.Biol.Chem., 280, 2005
|
|
1Y31
 
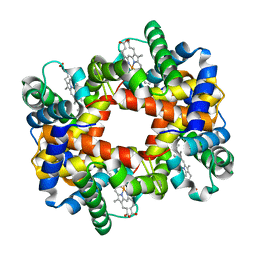 | |
1Y3U
 
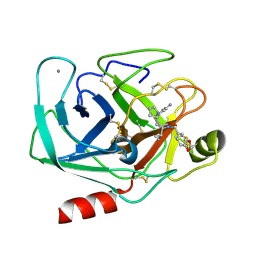 | | TRYPSIN INHIBITOR COMPLEX | | Descriptor: | (1R,3AS,4R,8AS,8BR)-4-(2-BENZO[1,3]DIOXOL-5-YLMETHYL-1-ISOPROPYL-3-OXO-DECAHYDRO-PYRROLO[3,4-A]PYRROLIZIN-4-YL)-BENZAMIDINE, CALCIUM ION, Trypsin, ... | | Authors: | Fokkens, J, Obst, U, Heine, A, Diederich, F, Klebe, G. | | Deposit date: | 2004-11-26 | | Release date: | 2005-12-06 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural and Thermodynamic Characterisation of Protein-Inhibitor-Complexes of Thrombin and Trypsin Inhibitors
To be Published
|
|
1Y45
 
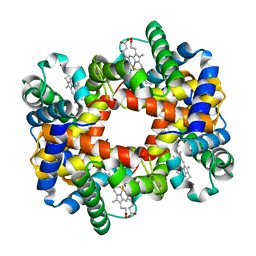 | |
1Y4P
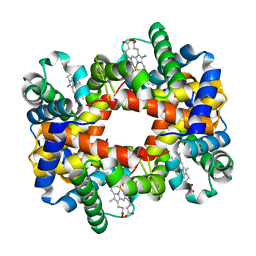 
 | |
1Y43
 
 | | crystal structure of aspergilloglutamic peptidase from Aspergillus niger | | Descriptor: | Aspergillopepsin II heavy chain, Aspergillopepsin II light chain, SULFATE ION | | Authors: | Sasaki, H, Nakagawa, A, Iwata, S, Muramatsu, T, Suganuma, M, Sawano, Y, Kojima, M, Kubota, K, Takahashi, K. | | Deposit date: | 2004-11-30 | | Release date: | 2005-12-13 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | The three-dimensional structure of aspergilloglutamic peptidase from Aspergillus niger
Proc.Jpn.Acad.,Ser.B, 80, 2004
|
|
1Y5C
 
 | |
