4U07
 
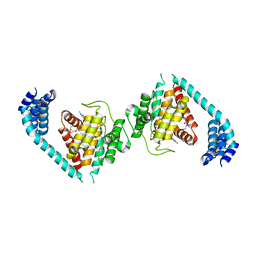 | | ATP bound to eukaryotic FIC domain containing protein | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Adenosine monophosphate-protein transferase FICD, MAGNESIUM ION, ... | | Authors: | Cole, A.R, Bunney, T.D, Katan, M. | | Deposit date: | 2014-07-11 | | Release date: | 2014-12-10 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.64 Å) | | Cite: | Crystal structure of the human, FIC-domain containing protein HYPE and implications for its functions.
Structure, 22, 2014
|
|
4U0Z
 
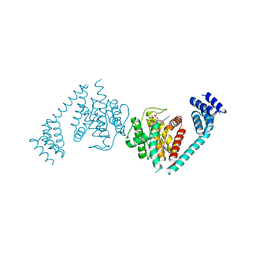 | | Eukaryotic Fic Domain containing protein with bound APCPP | | Descriptor: | Adenosine monophosphate-protein transferase FICD, DIPHOSPHOMETHYLPHOSPHONIC ACID ADENOSYL ESTER, MAGNESIUM ION | | Authors: | Cole, A.R, Bunney, T.D, Katan, M. | | Deposit date: | 2014-07-14 | | Release date: | 2014-12-10 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Crystal structure of the human, FIC-domain containing protein HYPE and implications for its functions.
Structure, 22, 2014
|
|
6K5O
 
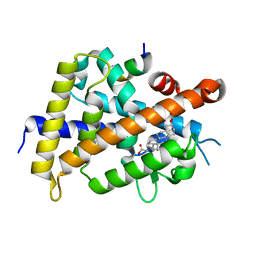 | | Development of Novel Lithocholic Acid Derivatives as Vitamin D Receptor Agonists | | Descriptor: | (4~{R})-4-[(3~{R},5~{R},8~{R},9~{S},10~{S},13~{R},14~{S},17~{R})-10,13-dimethyl-3-methylsulfonyloxy-2,3,4,5,6,7,8,9,11,12,14,15,16,17-tetradecahydro-1~{H}-cyclopenta[a]phenanthren-17-yl]pentanoic acid, Mediator of RNA polymerase II transcription subunit 1, Vitamin D3 receptor | | Authors: | Masuno, H, Kagechika, H, Ito, N. | | Deposit date: | 2019-05-29 | | Release date: | 2019-07-24 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Development of novel lithocholic acid derivatives as vitamin D receptor agonists.
Bioorg.Med.Chem., 27, 2019
|
|
4OJ2
 
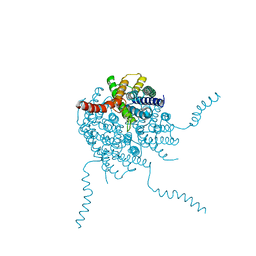 | |
4U0S
 
 | | Structure of Eukaryotic fic domain containing protein with ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Adenosine monophosphate-protein transferase FICD, MAGNESIUM ION, ... | | Authors: | Cole, A.R, Katan, M, Bunney, T.D. | | Deposit date: | 2014-07-14 | | Release date: | 2014-12-10 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | Crystal structure of the human, FIC-domain containing protein HYPE and implications for its functions.
Structure, 22, 2014
|
|
2FQF
 
 | | Crystal Structures of E. coli Laccase CueO under different copper binding situations | | Descriptor: | Blue copper oxidase cueO, CITRIC ACID, COPPER (II) ION, ... | | Authors: | Li, X, Wei, Z, Zhang, M, Teng, M, Gong, W. | | Deposit date: | 2006-01-18 | | Release date: | 2007-01-30 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structures of E. coli laccase CueO at different copper concentrations.
Biochem.Biophys.Res.Commun., 354, 2007
|
|
6KE6
 
 | |
5DIY
 
 | | Thermobaculum terrenum O-GlcNAc hydrolase mutant - D120N | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Hyaluronidase, TGF-beta-activated kinase 1 and MAP3K7-binding protein 1 | | Authors: | Ostrowski, A, Gundogdu, M, Ferenbach, A.T, Lebedev, A, van Aalten, D.M.F. | | Deposit date: | 2015-09-01 | | Release date: | 2015-10-28 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.06 Å) | | Cite: | Evidence for a Functional O-Linked N-Acetylglucosamine (O-GlcNAc) System in the Thermophilic Bacterium Thermobaculum terrenum.
J.Biol.Chem., 290, 2015
|
|
6KR4
 
 | | Crystal structure of the liprin-alpha3_SAM123/LAR_D1D2 complex | | Descriptor: | HEXAETHYLENE GLYCOL, Liprin-alpha-3, PHOSPHATE ION, ... | | Authors: | Xie, X, Liang, M, Luo, L, Wei, Z. | | Deposit date: | 2019-08-20 | | Release date: | 2020-01-15 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Structural basis of liprin-alpha-promoted LAR-RPTP clustering for modulation of phosphatase activity.
Nat Commun, 11, 2020
|
|
6DWM
 
 | | Structure of Human Cytochrome P450 1A1 with Bergamottin | | Descriptor: | 3-[(3-CHOLAMIDOPROPYL)DIMETHYLAMMONIO]-1-PROPANESULFONATE, 4-{[(2E)-3,7-dimethylocta-2,6-dien-1-yl]oxy}-7H-furo[3,2-g][1]benzopyran-7-one, Cytochrome P450 1A1, ... | | Authors: | Bart, A.G, Scott, E.E. | | Deposit date: | 2018-06-26 | | Release date: | 2018-10-03 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Structures of human cytochrome P450 1A1 with bergamottin and erlotinib reveal active-site modifications for binding of diverse ligands.
J. Biol. Chem., 293, 2018
|
|
6L54
 
 | | Structure of SMG189 | | Descriptor: | GUANOSINE-5'-TRIPHOSPHATE, MAGNESIUM ION, Protein SMG8, ... | | Authors: | Xu, Y, Qi, Y. | | Deposit date: | 2019-10-22 | | Release date: | 2020-04-15 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (3.43 Å) | | Cite: | Cryo-EM structure of SMG1-SMG8-SMG9 complex.
Cell Res., 29, 2019
|
|
4PYP
 
 | | Crystal structure of the human glucose transporter GLUT1 | | Descriptor: | Solute carrier family 2, facilitated glucose transporter member 1, nonyl beta-D-glucopyranoside | | Authors: | Deng, D, Yan, C.Y, Xu, C, Wu, J.P, Sun, P.C, Hu, M.X, Yan, N. | | Deposit date: | 2014-03-27 | | Release date: | 2014-05-21 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (3.166 Å) | | Cite: | Crystal structure of the human glucose transporter GLUT1
Nature, 510, 2014
|
|
4Q21
 
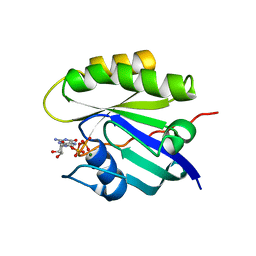 | |
2C9O
 
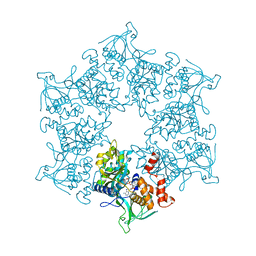 | | 3D Structure of the human RuvB-like helicase RuvBL1 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, RUVB-LIKE 1 | | Authors: | Matias, P.M, Gorynia, S, Donner, P, Carrondo, M.A. | | Deposit date: | 2005-12-14 | | Release date: | 2006-10-23 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of the human AAA+ protein RuvBL1.
J. Biol. Chem., 281, 2006
|
|
4QH7
 
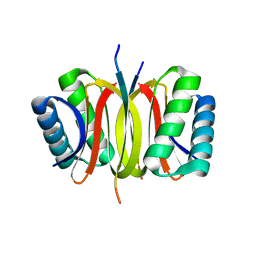 | | LC8 - Ana2 (159-168) Complex | | Descriptor: | Anastral spindle 2, Dynein light chain 1, cytoplasmic | | Authors: | Slevin, L.K, Romes, E.R, Slep, K.C. | | Deposit date: | 2014-05-27 | | Release date: | 2014-06-11 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.829 Å) | | Cite: | The Mechanism of Dynein Light Chain LC8-mediated Oligomerization of the Ana2 Centriole Duplication Factor.
J.Biol.Chem., 289, 2014
|
|
4QQP
 
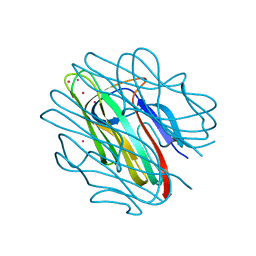 | | Crystal structure of C1QL3 mutant D207A | | Descriptor: | CADMIUM ION, Complement C1q-like protein 3 | | Authors: | Ressl, S, Brunger, A.T. | | Deposit date: | 2014-06-27 | | Release date: | 2015-04-15 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.464 Å) | | Cite: | Structures of C1q-like Proteins Reveal Unique Features among the C1q/TNF Superfamily.
Structure, 23, 2015
|
|
4QQO
 
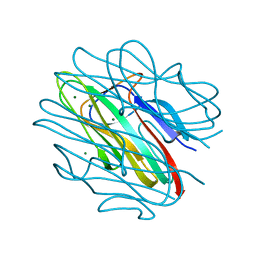 | | Crystal structure of C1QL3 mutant D205A | | Descriptor: | Complement C1q-like protein 3, MAGNESIUM ION | | Authors: | Ressl, S, Brunger, A.T. | | Deposit date: | 2014-06-27 | | Release date: | 2015-04-22 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.031 Å) | | Cite: | Structures of C1q-like Proteins Reveal Unique Features among the C1q/TNF Superfamily.
Structure, 23, 2015
|
|
4QQL
 
 | | Crystal structure of C1QL3 in P1 space group | | Descriptor: | Complement C1q-like protein 3, MAGNESIUM ION | | Authors: | Ressl, S, Brunger, A.T. | | Deposit date: | 2014-06-27 | | Release date: | 2015-04-22 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Structures of C1q-like Proteins Reveal Unique Features among the C1q/TNF Superfamily.
Structure, 23, 2015
|
|
4QPY
 
 | | Crystal structure of C1QL2 | | Descriptor: | Complement C1q-like protein 2 | | Authors: | Ressl, S, Brunger, A.T. | | Deposit date: | 2014-06-25 | | Release date: | 2015-04-15 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.384 Å) | | Cite: | Structures of C1q-like Proteins Reveal Unique Features among the C1q/TNF Superfamily.
Structure, 23, 2015
|
|
4QQH
 
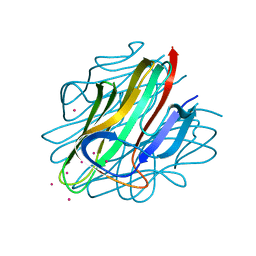 | | Crystal structure of C1QL3 in space group H32 | | Descriptor: | CADMIUM ION, Complement C1q-like protein 3 | | Authors: | Ressl, S, Brunger, A.T. | | Deposit date: | 2014-06-27 | | Release date: | 2015-04-15 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Structures of C1q-like Proteins Reveal Unique Features among the C1q/TNF Superfamily.
Structure, 23, 2015
|
|
4QT7
 
 | |
2Z75
 
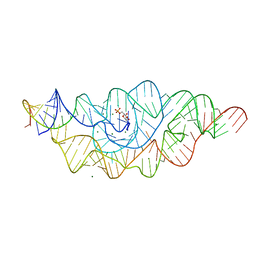 | | T. tengcongensis glmS ribozyme bound to glucosamine-6-phosphate | | Descriptor: | 2-amino-2-deoxy-6-O-phosphono-alpha-D-glucopyranose, MAGNESIUM ION, glmS ribozyme RNA, ... | | Authors: | Klein, D.J, Wilkinson, S.R, Been, M.D, Ferre-D'Amare, A.R. | | Deposit date: | 2007-08-15 | | Release date: | 2007-09-04 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Requirement of helix P2.2 and nucleotide G1 for positioning the cleavage site and cofactor of the glmS ribozyme
J.Mol.Biol., 373, 2007
|
|
2Z74
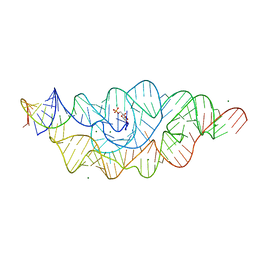 
 | | T. tengcongensis glmS ribozyme bound to glucose-6-phosphate | | Descriptor: | 6-O-phosphono-alpha-D-glucopyranose, MAGNESIUM ION, glmS ribozyme RNA, ... | | Authors: | Klein, D.J, Wilkinson, S.R, Been, M.D, Ferre-D'Amare, A.R. | | Deposit date: | 2007-08-15 | | Release date: | 2007-09-04 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Requirement of helix P2.2 and nucleotide G1 for positioning the cleavage site and cofactor of the glmS ribozyme
J.Mol.Biol., 373, 2007
|
|
4QQ2
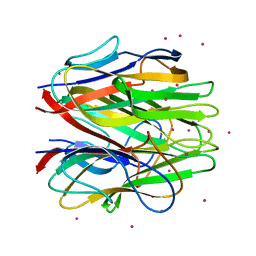 
 | | Crystal structure of C1QL1 | | Descriptor: | C1q-related factor, CADMIUM ION | | Authors: | Ressl, S, Brunger, A.T. | | Deposit date: | 2014-06-26 | | Release date: | 2015-04-15 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.801 Å) | | Cite: | Structures of C1q-like Proteins Reveal Unique Features among the C1q/TNF Superfamily.
Structure, 23, 2015
|
|
7TQ7
 
 | | Structure of MERS 3CL protease in complex with the cyclopropane based inhibitor 13c | | Descriptor: | N-{(2S)-1-oxo-3-[(3S)-2-oxopyrrolidin-3-yl]propan-2-yl}-N~2~-({[(1R,2R)-2-propylcyclopropyl]methoxy}carbonyl)-L-leucinamide, Orf1a protein, TETRAETHYLENE GLYCOL | | Authors: | Lovell, S, Liu, L, Battaile, K.P, Nguyen, H.N, Chamandi, S.D, Picard, H.R, Madden, T.K, Thruman, H.A, Kim, Y, Groutas, W.C, Chang, K.O. | | Deposit date: | 2022-01-26 | | Release date: | 2022-02-09 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Broad-Spectrum Cyclopropane-Based Inhibitors of Coronavirus 3C-like Proteases: Biochemical, Structural, and Virological Studies.
Acs Pharmacol Transl Sci, 6, 2023
|
|
