9G82
 
 | | CTX-M-14 mixed with piperacillin at 3s delay time - serial crystallography temperature series; 37C, 310K | | Descriptor: | Beta-lactamase, Hydrolyzed piperacillin, Piperacillin, ... | | Authors: | Prester, A, von Stetten, D, Mehrabi, P, Schulz, E.C. | | Deposit date: | 2024-07-22 | | Release date: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Probing the modulation of enzyme kinetics by multi-temperature, time-resolved serial crystallography.
Nat Commun, 16, 2025
|
|
1M75
 
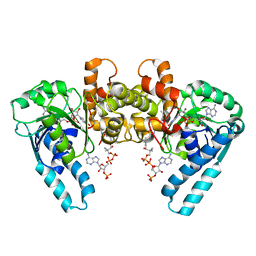 | |
7S14
 
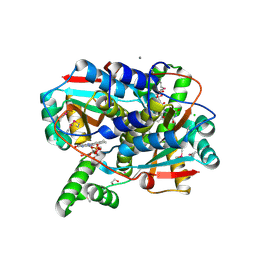 | | Crystal structure of putative NAD(P)H-flavin oxidoreductase from Haemophilus influenzae 86-028NP | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Kim, Y, Maltseva, N, Endres, M, Crofts, T, Joachimiak, A, Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2021-08-31 | | Release date: | 2021-10-06 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Functional and Structural Characterization of Diverse NfsB Chloramphenicol Reductase Enzymes from Human Pathogens.
Microbiol Spectr, 10, 2022
|
|
9G5J
 
 | | Structure of the PRO-PRO endopeptidase (PPEP-3) E153A Y189F in complex with substrate peptide Ac-EPLPPPP-NH2 from Geobacillus thermodenitrificans | | Descriptor: | ACE-GLU-PRO-LEU-PRO-PRO-PRO-PRO-NH2, ATLF-like domain-containing protein, TETRAETHYLENE GLYCOL, ... | | Authors: | Claushuis, B, Wojtalla, F, van Leeuwen, H, Corver, J, Baumann, U, Hensbergen, P. | | Deposit date: | 2024-07-17 | | Release date: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of the PRO-PRO endopeptidase (PPEP-3) E153A Y189F in complex with substrate peptide Ac-EPLPPPP-NH2 from Geobacillus thermodenitrificans
To Be Published
|
|
7SB8
 
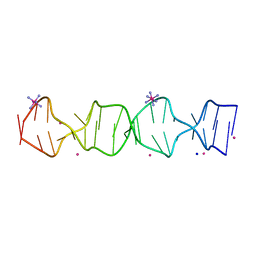 | | d(GA(CGA)5) parallel-stranded homo-duplex | | Descriptor: | COBALT HEXAMMINE(III), GA(CGA)5, SODIUM ION, ... | | Authors: | Luteran, E.M, Paukstelis, P.J. | | Deposit date: | 2021-09-24 | | Release date: | 2021-10-06 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.317 Å) | | Cite: | The parallel-stranded d(CGA) duplex is a highly predictable structural motif with two conformationally distinct strands.
Acta Crystallogr D Struct Biol, 78, 2022
|
|
9G6W
 
 | | L-SIGN CRD in complex with Man96. | | Descriptor: | (2~{R},3~{S},4~{R},5~{S},6~{S})-6-(2-chloroethyloxy)-2-(hydroxymethyl)-5-[4-[(imidazolidin-2-ylideneamino)methyl]-1,2,3-triazol-1-yl]oxane-3,4-diol, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CALCIUM ION, ... | | Authors: | Thepaut, M, Cavazzoli, G, Bernardi, A, Fieschi, F. | | Deposit date: | 2024-07-19 | | Release date: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | L-SIGN CRD in complex with Man96.
To be published
|
|
1M7Z
 
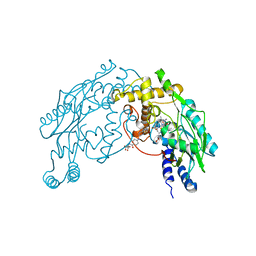 | | Structure of Nitric Oxide Synthase Heme Protein from Bacillus Subtilis with N-Hydroxy-Arginine and Tetrahydrofolate Bound | | Descriptor: | (6S)-5,6,7,8-TETRAHYDROFOLATE, N-OMEGA-HYDROXY-L-ARGININE, Nitric oxide synthase, ... | | Authors: | Pant, K, Bilwes, A.M, Adak, S, Stuehr, D.J, Crane, B.R. | | Deposit date: | 2002-07-23 | | Release date: | 2002-10-30 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Structure of a nitric oxide synthase heme protein from Bacillus subtilis.
Biochemistry, 41, 2002
|
|
9G5N
 
 | | Xylose Isomerase collected at 20C using serial fixed-target crystallography | | Descriptor: | MAGNESIUM ION, Xylose isomerase | | Authors: | Schulz, E.C, Prester, A, Stetten, D.V, Gore, G, Hatton, C.E, Bartels, K, Leimkohl, J.P, Schikora, H, Ginn, H.M, Tellkamp, F, Mehrabi, P. | | Deposit date: | 2024-07-17 | | Release date: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Xylose Isomerase collected at 20C using serial fixed-target crystallography
To Be Published
|
|
1M8F
 
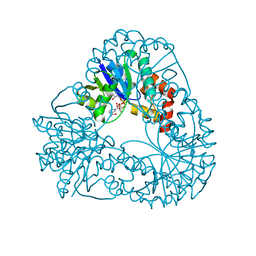 | |
7S2H
 
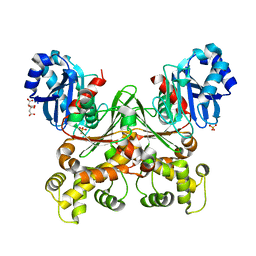 | | Crystal structure of Trypanosoma cruzi glucokinase in the apo form (open conformation) | | Descriptor: | CITRATE ANION, Glucokinase 1, putative, ... | | Authors: | Kearns, S.P, Daneshian, L, Swartz, P.D, Carey, S.M, Chruszcz, M, D'Antonio, E.L. | | Deposit date: | 2021-09-03 | | Release date: | 2021-10-06 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of Trypanosoma cruzi glucokinase in the apo form (open conformation)
To Be Published
|
|
9G5X
 
 | | Xylose Isomerase collected at 45C using serial fixed-target crystallography | | Descriptor: | MAGNESIUM ION, Xylose isomerase | | Authors: | Schulz, E.C, Prester, A, Stetten, D.V, Gore, G, Hatton, C.E, Bartels, K, Leimkohl, J.P, Schikora, H, Ginn, H.M, Tellkamp, F, Mehrabi, P. | | Deposit date: | 2024-07-17 | | Release date: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Probing the modulation of enzyme kinetics by multi-temperature, time-resolved serial crystallography.
Nat Commun, 16, 2025
|
|
2PE5
 
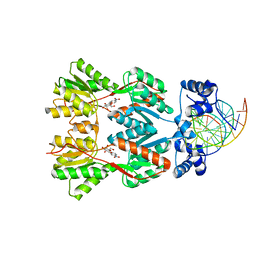 | | Crystal Structure of the Lac Repressor bound to ONPG in repressed state | | Descriptor: | 2-nitrophenyl beta-D-galactopyranoside, DNA (5'-D(*DAP*DAP*DTP*DTP*DGP*DTP*DGP*DAP*DGP*DCP*DGP*DCP*DTP*DCP*DAP*DCP*DAP*DAP*DTP*DT)-3'), Lactose operon repressor | | Authors: | Daber, R, Stayrook, S.E, Rosenberg, A, Lewis, M. | | Deposit date: | 2007-04-02 | | Release date: | 2008-03-18 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Structural analysis of lac repressor bound to allosteric effectors
J.Mol.Biol., 370, 2007
|
|
1MEQ
 
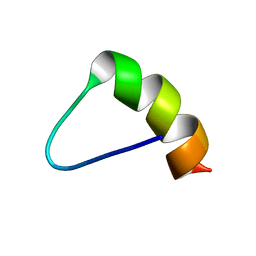 | | HIV gp120 C5 | | Descriptor: | Exterior Membrane Glycoprotein (GP120) | | Authors: | Caffrey, M, Jacobs, A, Guilhaudis, L. | | Deposit date: | 2002-08-08 | | Release date: | 2002-12-11 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the HIV gp120 C5 Domain
Eur.J.Biochem., 269, 2002
|
|
1MES
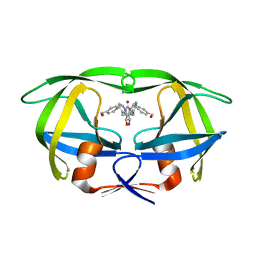 
 | | HIV-1 MUTANT (I84V) PROTEASE COMPLEXED WITH DMP323 | | Descriptor: | HIV-1 PROTEASE, [4-R-(-4-ALPHA,5-ALPHA,6-BETA,7-BETA)]-HEXAHYDRO-5,6-BIS(HYDROXY)-[1,3-BIS([4-HYDROXYMETHYL-PHENYL]METHYL)-4,7-BIS(PHEN YLMETHYL)]-2H-1,3-DIAZEPINONE | | Authors: | Ala, P, Chang, C.-H. | | Deposit date: | 1997-04-11 | | Release date: | 1998-04-15 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Molecular basis of HIV-1 protease drug resistance: structural analysis of mutant proteases complexed with cyclic urea inhibitors.
Biochemistry, 36, 1997
|
|
1MF0
 
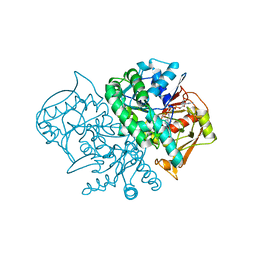 | | Structure of the Recombinant Mouse-Muscle Adenylosuccinate Synthetase Complexed with AMP, GDP, HPO4(2-), and Mg(2+) | | Descriptor: | ADENOSINE MONOPHOSPHATE, Adenylosuccinate Synthetase, GUANOSINE-5'-DIPHOSPHATE, ... | | Authors: | Iancu, C.V, Borza, T, Fromm, H.J, Honzatko, R.B. | | Deposit date: | 2002-08-09 | | Release date: | 2002-10-30 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Feedback inhibition and product complexes of recombinant mouse muscle adenylosuccinate synthetase.
J.Biol.Chem., 277, 2002
|
|
9G7V
 
 | | CTX-M-14 apo serial crystallography temperature series; 10C, 283K | | Descriptor: | Beta-lactamase, SULFATE ION | | Authors: | Prester, A, von Stetten, D, Mehrabi, P, Schulz, E.C. | | Deposit date: | 2024-07-22 | | Release date: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Probing the modulation of enzyme kinetics by multi-temperature, time-resolved serial crystallography.
Nat Commun, 16, 2025
|
|
1M8N
 
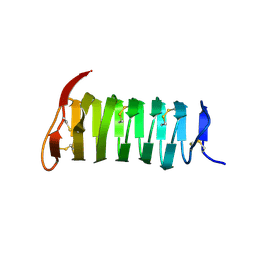 | |
9G7Y
 
 | | CTX-M-14 apo serial crystallography temperature series; 40C, 313K | | Descriptor: | Beta-lactamase, SULFATE ION | | Authors: | Prester, A, von Stetten, D, Mehrabi, P, Schulz, E.C. | | Deposit date: | 2024-07-22 | | Release date: | 2025-07-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Probing the modulation of enzyme kinetics by multi-temperature, time-resolved serial crystallography.
Nat Commun, 16, 2025
|
|
9FXO
 
 | |
1M94
 
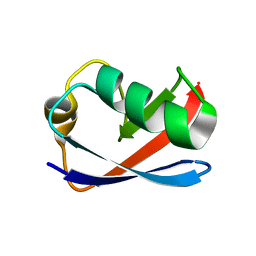 | | Solution Structure of the Yeast Ubiquitin-Like Modifier Protein Hub1 | | Descriptor: | Protein YNR032c-a | | Authors: | Ramelot, T.A, Cort, J.R, Yee, A.A, Semesi, A, Edwards, A.M, Arrowsmith, C.H, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2002-07-26 | | Release date: | 2002-12-11 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the Yeast Ubiquitin-Like Modifier Protein Hub1
J.STRUCT.FUNCT.GENOM., 4, 2003
|
|
7S2N
 
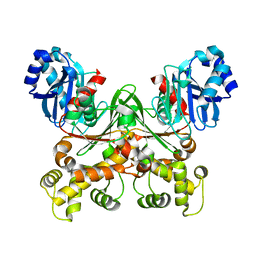 | |
1MA0
 
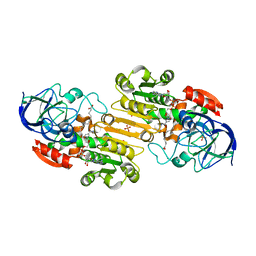 | | Ternary complex of Human glutathione-dependent formaldehyde dehydrogenase with NAD+ and dodecanoic acid | | Descriptor: | Glutathione-dependent formaldehyde dehydrogenase, LAURIC ACID, NICOTINAMIDE-ADENINE-DINUCLEOTIDE, ... | | Authors: | Sanghani, P.C, Robinson, H, Bosron, W.F, Hurley, T.D. | | Deposit date: | 2002-07-30 | | Release date: | 2002-08-02 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Human glutathione-dependent formaldehyde dehydrogenase. Structures of apo, binary, and inhibitory ternary complexes.
Biochemistry, 41, 2002
|
|
7SBC
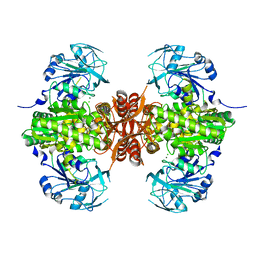 
 | |
7S2P
 
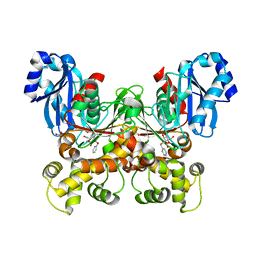 | | Crystal structure of the F337L mutation of Trypanosoma cruzi glucokinase in complex with inhibitor CBZ-GlcN | | Descriptor: | 2-{[(benzyloxy)carbonyl]amino}-2-deoxy-beta-D-glucopyranose, Glucokinase 1, SULFATE ION | | Authors: | Carey, S.M, Nettles, R.B, Daneshian, L, Chruszcz, M, D'Antonio, E.L. | | Deposit date: | 2021-09-03 | | Release date: | 2021-10-06 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Crystal structure of the F337L mutation of Trypanosoma cruzi glucokinase in complex with inhibitor CBZ-GlcN
To Be Published
|
|
1MG6
 
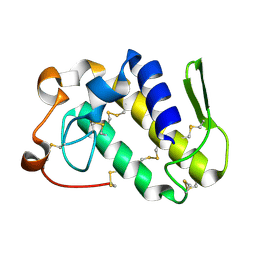 | |
