6BGY
 
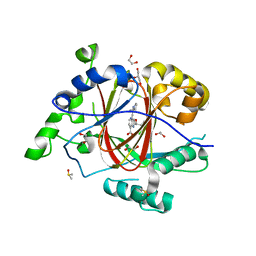 | | LINKED KDM5A JMJ DOMAIN BOUND TO THE INHIBITOR 2-((2-chlorophenyl)(2-(1-methylpyrrolidin-2-yl)ethoxy)methyl)-1H-pyrrolo[3,2-b]pyridine-7-carboxylic acid(Compound 46) | | Descriptor: | 1,2-ETHANEDIOL, 2-[(S)-(2-chlorophenyl){2-[(2R)-1-methylpyrrolidin-2-yl]ethoxy}methyl]-1H-pyrrolo[3,2-b]pyridine-7-carboxylic acid, DIMETHYL SULFOXIDE, ... | | Authors: | Horton, J.R, Cheng, X. | | Deposit date: | 2017-10-29 | | Release date: | 2018-03-28 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.22 Å) | | Cite: | Insights into the Action of Inhibitor Enantiomers against Histone Lysine Demethylase 5A.
J. Med. Chem., 61, 2018
|
|
8W5L
 
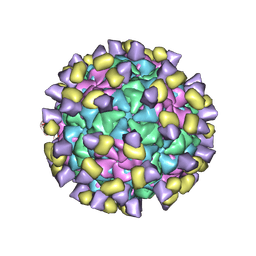 | | Cryo-EM structure of Qb-Ab16 | | Descriptor: | Heavy chain of Ab16, Light chain of Ab16, Minor capsid protein A1 | | Authors: | Bao, K.Y, Li, R.H, Hua, Z.L, Hou, B.D, Zhu, P. | | Deposit date: | 2023-08-27 | | Release date: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | Competition propels, rather than limits, the success of low-affinity B cells in the germinal center response.
Cell Rep, 44, 2025
|
|
8W5R
 
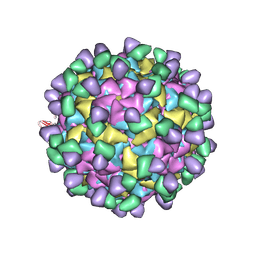 | | Cryo-EM structure of Qb-Ab53 | | Descriptor: | Heavy chain of Ab53, Light chain of Ab53, Minor capsid protein A1 | | Authors: | Bao, K.Y, Li, R.H, Hua, Z.L, Hou, B.D, Zhu, P. | | Deposit date: | 2023-08-27 | | Release date: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Competition propels, rather than limits, the success of low-affinity B cells in the germinal center response.
Cell Rep, 44, 2025
|
|
8W5G
 
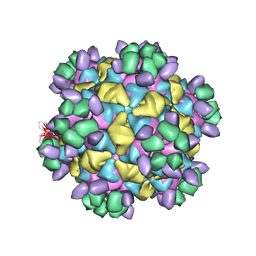 | | Cryo-EM structure of Qb-Ab7 | | Descriptor: | Heavy chain of Ab7, Light chain of Ab7, Minor capsid protein A1 | | Authors: | Bao, K.Y, Li, R.H, Hua, Z.L, Hou, B.D, Zhu, P. | | Deposit date: | 2023-08-26 | | Release date: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (4 Å) | | Cite: | Competition propels, rather than limits, the success of low-affinity B cells in the germinal center response.
Cell Rep, 44, 2025
|
|
8W5H
 
 | | Cryo-EM structure of Qb-Ab10 | | Descriptor: | Heavy chain of Ab10, Light chain of Ab10, Minor capsid protein A1 | | Authors: | Bao, K.Y, Li, R.H, Hua, Z.L, Hou, B.D, Zhu, P. | | Deposit date: | 2023-08-26 | | Release date: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | Competition propels, rather than limits, the success of low-affinity B cells in the germinal center response.
Cell Rep, 44, 2025
|
|
8W5W
 
 | | Cryo-EM structure of Qb-Ab8 | | Descriptor: | Heavy chain of Ab8, Light chain of Ab8, Minor capsid protein A1 | | Authors: | Bao, K.Y, Li, R.H, Hua, Z.L, Hou, B.D, Zhu, P. | | Deposit date: | 2023-08-27 | | Release date: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Competition propels, rather than limits, the success of low-affinity B cells in the germinal center response.
Cell Rep, 44, 2025
|
|
8W5M
 
 | | Cryo-EM structure of Qb-Ab17 | | Descriptor: | Heavy chain of Ab17, Light chain of Ab17, Minor capsid protein A1 | | Authors: | Bao, K.Y, Li, R.H, Hua, Z.L, Hou, B.D, Zhu, P. | | Deposit date: | 2023-08-27 | | Release date: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Competition propels, rather than limits, the success of low-affinity B cells in the germinal center response.
Cell Rep, 44, 2025
|
|
5LHI
 
 | | Structure of the KDM1A/CoREST complex with the inhibitor N-[3-(ethoxymethyl)-2-[[4-[[(3R)-pyrrolidin-3-yl]methoxy]phenoxy]methyl]phenyl]-4-methylthieno[3,2-b]pyrrole-5-carboxamide | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, GLYCEROL, LYSINE-SPECIFIC HISTONE DEMETHYLASE 1, ... | | Authors: | Cecatiello, V, Pasqualato, S. | | Deposit date: | 2016-07-12 | | Release date: | 2017-02-22 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Thieno[3,2-b]pyrrole-5-carboxamides as New Reversible Inhibitors of Histone Lysine Demethylase KDM1A/LSD1. Part 2: Structure-Based Drug Design and Structure-Activity Relationship.
J. Med. Chem., 60, 2017
|
|
8W5O
 
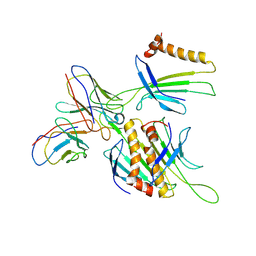 | | Cryo-EM structure of Qb-Ab31 | | Descriptor: | Heavy chain of Ab31, Light chain of Ab31, Minor capsid protein A1 | | Authors: | Bao, K.Y, Li, R.H, Hua, Z.L, Hou, B.D, Zhu, P. | | Deposit date: | 2023-08-27 | | Release date: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Competition propels, rather than limits, the success of low-affinity B cells in the germinal center response.
Cell Rep, 44, 2025
|
|
8W5N
 
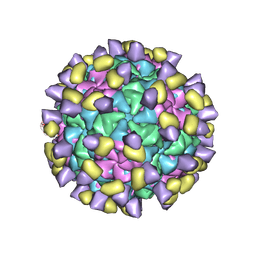 | | Cryo-EM structure of Qb-Ab21 | | Descriptor: | Heavy chain of Ab21, Light chain of Ab21, Minor capsid protein A1 | | Authors: | Bao, K.Y, Li, R.H, Hua, Z.L, Hou, B.D, Zhu, P. | | Deposit date: | 2023-08-27 | | Release date: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Competition propels, rather than limits, the success of low-affinity B cells in the germinal center response.
Cell Rep, 44, 2025
|
|
8W5I
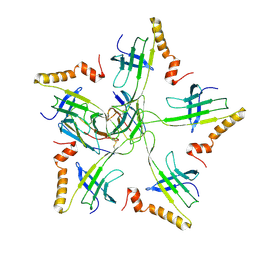 
 | | Cryo-EM structure of Qb-Ab14 | | Descriptor: | Heavy chain of Ab14, Light chain of Ab14, Minor capsid protein A1 | | Authors: | Bao, K.Y, Li, R.H, Hua, Z.L, Hou, B.D, Zhu, P. | | Deposit date: | 2023-08-26 | | Release date: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Competition propels, rather than limits, the success of low-affinity B cells in the germinal center response.
Cell Rep, 44, 2025
|
|
6E0R
 
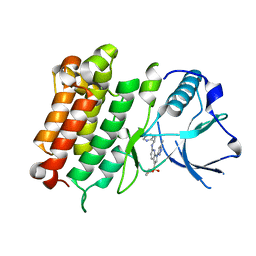 | | hALK in complex with compound 7 N-((1S)-1-(5-fluoropyridin-2-yl)ethyl)-1-(5-methyl-1H-pyrazol-3-yl)-3-(oxetan-3-ylsulfonyl)-1H-pyrrolo[2,3-b]pyridin-6-amine | | Descriptor: | ALK tyrosine kinase receptor, N-[(1S)-1-(5-fluoropyridin-2-yl)ethyl]-1-(5-methyl-1H-pyrazol-3-yl)-3-[(oxetan-3-yl)sulfonyl]-1H-pyrrolo[2,3-b]pyridin-6-amine | | Authors: | Lane, W, Saikatendu, K. | | Deposit date: | 2018-07-06 | | Release date: | 2019-05-01 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.303 Å) | | Cite: | Discovery of Potent, Selective, and Brain-Penetrant 1 H-Pyrazol-5-yl-1 H-pyrrolo[2,3- b]pyridines as Anaplastic Lymphoma Kinase (ALK) Inhibitors.
J.Med.Chem., 62, 2019
|
|
6EDL
 
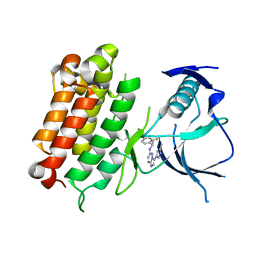 | | hALK in complex with compound 1 (S)-N-(1-(2,4-difluorophenyl)ethyl)-3-(3-methyl-1H-pyrazol-5-yl)imidazo[1,2-b]pyridazin-6-amine | | Descriptor: | ALK tyrosine kinase receptor, N-[(1S)-1-(2,4-difluorophenyl)ethyl]-3-(5-methyl-1H-pyrazol-3-yl)imidazo[1,2-b]pyridazin-6-amine | | Authors: | Lane, W, Saikatendu, K. | | Deposit date: | 2018-08-09 | | Release date: | 2019-05-01 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.799 Å) | | Cite: | Discovery of Potent, Selective, and Brain-Penetrant 1 H-Pyrazol-5-yl-1 H-pyrrolo[2,3- b]pyridines as Anaplastic Lymphoma Kinase (ALK) Inhibitors.
J.Med.Chem., 62, 2019
|
|
9DFD
 
 | | Crystal structure of the wild-type Thermus thermophilus 70S ribosome in complex with lasso peptide lariocidin B, mRNA, aminoacylated A-site Phe-tRNAphe, aminoacylated P-site fMet-tRNAmet, and deacylated E-site tRNAphe at 2.60A resolution | | Descriptor: | 16S Ribosomal RNA, 23S Ribosomal RNA, 30S ribosomal protein S10, ... | | Authors: | Aleksandrova, E.V, Travin, D.Y, Jangra, M, Kaur, M, Darwish, L, Koteva, K, Klepacki, D, Wang, W, Tiffany, M, Sokaribo, A, Coombes, B.K, Vazquez-Laslop, N, Wright, G.D, Mankin, A.S, Polikanov, Y.S. | | Deposit date: | 2024-08-29 | | Release date: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | A Broad Spectrum Lasso Peptide Antibiotic Targeting the Bacterial Ribosome.
Res Sq, 2024
|
|
6OQ5
 
 | | Structure of the full-length Clostridium difficile toxin B in complex with 3 VHHs | | Descriptor: | 5D, 7F, E3, ... | | Authors: | Chen, P, Lam, K, Jin, R. | | Deposit date: | 2019-04-25 | | Release date: | 2019-07-10 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (3.87 Å) | | Cite: | Structure of the full-length Clostridium difficile toxin B.
Nat.Struct.Mol.Biol., 26, 2019
|
|
6EBW
 
 | | hALK in complex with compound 9 (6-(((1S)-1-(5-Fluoropyridin-2-yl)ethyl)amino)-1-(3-methyl-1H-pyrazol-5-yl)-1H-pyrrolo[2,3-b]pyridin-3-yl)(morpholin-4-yl)methanone | | Descriptor: | ALK tyrosine kinase receptor, [6-{[(1S)-1-(5-fluoropyridin-2-yl)ethyl]amino}-1-(5-methyl-1H-pyrazol-3-yl)-1H-pyrrolo[2,3-b]pyridin-3-yl](morpholin-4-yl)methanone | | Authors: | Lane, W, Saikatendu, K. | | Deposit date: | 2018-08-07 | | Release date: | 2019-05-01 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.455 Å) | | Cite: | Discovery of Potent, Selective, and Brain-Penetrant 1 H-Pyrazol-5-yl-1 H-pyrrolo[2,3- b]pyridines as Anaplastic Lymphoma Kinase (ALK) Inhibitors.
J.Med.Chem., 62, 2019
|
|
5MNS
 
 | | Structural and functional characterization of OleP in complex with 6DEB in sodium formate | | Descriptor: | 6-DEOXYERYTHRONOLIDE B, Cytochrome P-450, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Parisi, G, Savino, C, Montemiglio, L.C, Vallone, B. | | Deposit date: | 2016-12-13 | | Release date: | 2018-02-28 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.62 Å) | | Cite: | Substrate-induced conformational change in cytochrome P450 OleP.
FASEB J., 33, 2019
|
|
2N96
 
 | | An unexpected mode of small molecule DNA binding provides the structural basis for DNA cleavage by the potent antiproliferative agent (-)-lomaiviticin A | | Descriptor: | (1R,1'R,2S,2'S,3R,3'R,5aR,10aR,11a'S)-2'-[(2,6-dideoxy-3-O-methyl-alpha-L-arabino-hexopyranosyl)oxy]-2,2'-diethyl-11,11 '-dihydrazinyl-6,6',9,9'-tetrahydroxy-4,4',5,5',10,10'-hexaoxo-1,1'-bis{[2,4,6-trideoxy-4-(dimethylamino)-beta-L-arabino -hexopyranosyl]oxy}[2,2',3,3',4,4',5,5',5a,8,10,10',10a,11a'-tetradecahydro-1H,1'H-[3,3'-bibenzo[b]fluorene]]-2-yl 2,6-dideoxy-3-O-methyl-alpha-L-arabino-hexopyranoside, DNA (5'-D(*GP*CP*TP*AP*TP*AP*GP*C)-3') | | Authors: | Woo, C.M, Li, Z, Paulson, E, Herzon, S.B. | | Deposit date: | 2015-11-07 | | Release date: | 2016-06-01 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural basis for DNA cleavage by the potent antiproliferative agent (-)-lomaiviticin A.
Proc.Natl.Acad.Sci.USA, 113, 2016
|
|
5DLX
 
 | | FIRST DOMAIN OF HUMAN BROMODOMAIN BRD4 IN COMPLEX WITH INHIBITOR N-{3-[4-(3-chlorophenyl)piperazin-1-yl]propyl}-1-{3-methyl-[1,2,4]triazolo[4,3-b]pyridazin-6-yl}piperidine-4-carboxamide | | Descriptor: | Bromodomain-containing protein 4, N-{3-[4-(3-chlorophenyl)piperazin-1-yl]propyl}-1-(3-methyl[1,2,4]triazolo[4,3-b]pyridazin-6-yl)piperidine-4-carboxamide | | Authors: | Raux, B, Rebuffet, E, Betzi, S, Morelli, X. | | Deposit date: | 2015-09-07 | | Release date: | 2016-06-01 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Protein-Protein Interaction Inhibition (2P2I)-Oriented Chemical Library Accelerates Hit Discovery.
Acs Chem.Biol., 11, 2016
|
|
3I0C
 
 | | Crystal structure of GTB C80S/C196S unliganded | | Descriptor: | ABO glycosyltransferase | | Authors: | Schuman, B, Persson, M, Landry, R.C, Polakowski, R, Weadge, J.T, Seto, N.O.L, Borisova, S, Palcic, M.M, Evans, S.V. | | Deposit date: | 2009-06-25 | | Release date: | 2010-08-11 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Cysteine-to-serine mutants dramatically reorder the active site of human ABO(H) blood group B glycosyltransferase without affecting activity: structural insights into cooperative substrate binding
J.Mol.Biol., 402, 2010
|
|
3I0H
 
 | | Crystal structure of GTB C80S/C196S/C209S unliganded | | Descriptor: | ABO glycosyltransferase | | Authors: | Schuman, B, Persson, M, Landry, R.C, Polakowski, R, Weadge, J.T, Seto, N.O.L, Borisova, S, Palcic, M.M, Evans, S.V. | | Deposit date: | 2009-06-25 | | Release date: | 2010-08-11 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Cysteine-to-serine mutants dramatically reorder the active site of human ABO(H) blood group B glycosyltransferase without affecting activity: structural insights into cooperative substrate binding
J.Mol.Biol., 402, 2010
|
|
3I0K
 
 | | Crystal structure of GTB C80S/C196S/C209S + UDP + H antigen | | Descriptor: | ABO glycosyltransferase, URIDINE-5'-DIPHOSPHATE, alpha-L-fucopyranose-(1-2)-hexyl beta-D-galactopyranoside | | Authors: | Schuman, B, Persson, M, Landry, R.C, Polakowski, R, Weadge, J.T, Seto, N.O.L, Borisova, S, Palcic, M.M, Evans, S.V. | | Deposit date: | 2009-06-25 | | Release date: | 2010-08-11 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Cysteine-to-serine mutants dramatically reorder the active site of human ABO(H) blood group B glycosyltransferase without affecting activity: structural insights into cooperative substrate binding
J.Mol.Biol., 402, 2010
|
|
5KQY
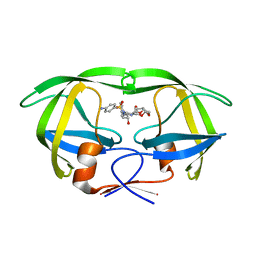 
 | | Protease E35D-DRV | | Descriptor: | (3R,3AS,6AR)-HEXAHYDROFURO[2,3-B]FURAN-3-YL(1S,2R)-3-[[(4-AMINOPHENYL)SULFONYL](ISOBUTYL)AMINO]-1-BENZYL-2-HYDROXYPROPYLCARBAMATE, Protease E35D-DRV | | Authors: | Liu, Z, Poole, K.M, Mahon, B.P, McKenna, R, Fanucci, G.E. | | Deposit date: | 2016-07-06 | | Release date: | 2016-09-21 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Effects of Hinge-region Natural Polymorphisms on Human Immunodeficiency Virus-Type 1 Protease Structure, Dynamics, and Drug Pressure Evolution.
J.Biol.Chem., 291, 2016
|
|
3I0F
 
 | | Crystal structure of GTB C80S/C196S + UDP + H antigen | | Descriptor: | ABO glycosyltransferase, MANGANESE (II) ION, URIDINE-5'-DIPHOSPHATE, ... | | Authors: | Schuman, B, Persson, M, Landry, R.C, Polakowski, R, Weadge, J.T, Seto, N.O.L, Borisova, S, Palcic, M.M, Evans, S.V. | | Deposit date: | 2009-06-25 | | Release date: | 2010-08-11 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Cysteine-to-serine mutants dramatically reorder the active site of human ABO(H) blood group B glycosyltransferase without affecting activity: structural insights into cooperative substrate binding
J.Mol.Biol., 402, 2010
|
|
3I0J
 
 | | Crystal structure of GTB C80S/C196S/C209S + H antigen | | Descriptor: | ABO glycosyltransferase, alpha-L-fucopyranose-(1-2)-hexyl beta-D-galactopyranoside | | Authors: | Schuman, B, Persson, M, Landry, R.C, Polakowski, R, Weadge, J.T, Seto, N.O.L, Borisova, S, Palcic, M.M, Evans, S.V. | | Deposit date: | 2009-06-25 | | Release date: | 2010-08-11 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | Cysteine-to-serine mutants dramatically reorder the active site of human ABO(H) blood group B glycosyltransferase without affecting activity: structural insights into cooperative substrate binding.
J.Mol.Biol., 402, 2010
|
|
