1AQG
 | |
1BLV
 
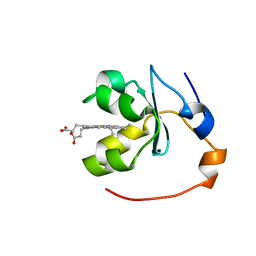 | | SOLUTION STRUCTURE OF OXIDIZED RAT MICROSOMAL CYTOCHROME B5 IN THE PRESENCE OF 2 M GUANIDINIUM CHLORIDE: MONITORING THE EARLY STEPS IN PROTEIN UNFOLDING | | Descriptor: | PROTEIN (CYTOCHROME B5), PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Arnesano, F, Banci, L, Bertini, I, Koulougliotis, D. | | Deposit date: | 1998-07-21 | | Release date: | 1998-07-29 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of oxidized rat microsomal cytochrome b5 in the presence of 2 M guanidinium chloride: monitoring the early steps in protein unfolding.
Biochemistry, 37, 1998
|
|
1FRF
 
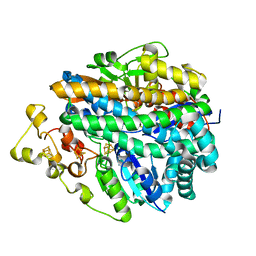 | | CRYSTAL STRUCTURE OF THE NI-FE HYDROGENASE FROM DESULFOVIBRIO FRUCTOSOVORANS | | Descriptor: | FE (III) ION, FE3-S4 CLUSTER, IRON/SULFUR CLUSTER, ... | | Authors: | Montet, Y, Volbeda, A, Piras, C, Hatchikian, E.C, Frey, M, Fontecilla, J.C. | | Deposit date: | 1998-07-21 | | Release date: | 1998-07-29 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | 3Fe-4S] to [4Fe-4S] cluster conversion in Desulfovibrio fructosovorans [NiFe] hydrogenase by site-directed mutagenesis
Proc.Natl.Acad.Sci.USA, 95, 1998
|
|
1PMR
 
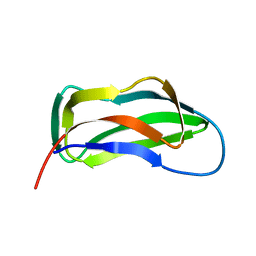 | | LIPOYL DOMAIN FROM THE DIHYDROLIPOYL SUCCINYLTRANSFERASE COMPONENT OF THE 2-OXOGLUTARATE DEHYDROGENASE MULTIENZYME COMPLEX OF ESCHERICHIA COLI, NMR, 25 STRUCTURES | | Descriptor: | DIHYDROLIPOYL SUCCINYLTRANSFERASE | | Authors: | Ricaud, P.M, Howard, M.J, Roberts, E.L, Broadhurst, R.W, Perham, R.N. | | Deposit date: | 1997-07-24 | | Release date: | 1998-07-29 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Three-dimensional structure of the lipoyl domain from the dihydrolipoyl succinyltransferase component of the 2-oxoglutarate dehydrogenase multienzyme complex of Escherichia coli.
J.Mol.Biol., 264, 1996
|
|
1FHE
 
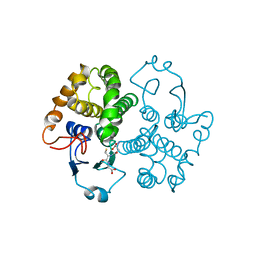 | | GLUTATHIONE TRANSFERASE (FH47) FROM FASCIOLA HEPATICA | | Descriptor: | GLUTATHIONE, GLUTATHIONE TRANSFERASE | | Authors: | Rossjohn, J, Parker, M.W. | | Deposit date: | 1997-07-24 | | Release date: | 1998-07-29 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Crystallization, structural determination and analysis of a novel parasite vaccine candidate: Fasciola hepatica glutathione S-transferase.
J.Mol.Biol., 273, 1997
|
|
2VPF
 | |
1A3Z
 
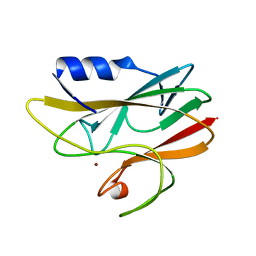 | | REDUCED RUSTICYANIN AT 1.9 ANGSTROMS | | Descriptor: | COPPER (I) ION, RUSTICYANIN | | Authors: | Zhao, D, Shoham, M. | | Deposit date: | 1998-01-27 | | Release date: | 1998-07-29 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Rusticyanin: Extremes in acid stability and redox potential explained by the crystal structure.
Biophys.J., 74, 1998
|
|
1MB1
 
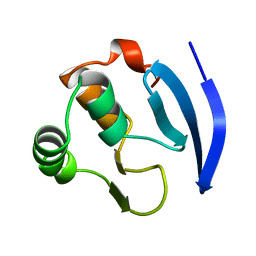 | | MBP1 FROM SACCHAROMYCES CEREVISIAE | | Descriptor: | MLU1-BOX BINDING PROTEIN | | Authors: | Taylor, I.A, Smerdon, S.J. | | Deposit date: | 1997-07-23 | | Release date: | 1998-07-29 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The X-ray structure of the DNA-binding domain from the Saccharomyces cerevisiae cell-cycle transcription factor Mbp1 at 2.1 A resolution.
J.Mol.Biol., 272, 1997
|
|
1NP4
 
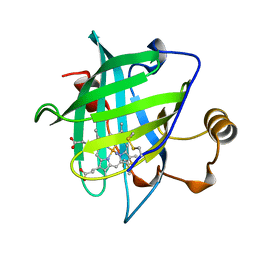 | | CRYSTAL STRUCTURE OF NITROPHORIN 4 FROM RHODNIUS PROLIXUS | | Descriptor: | AMMONIA, PROTEIN (NITROPHORIN 4), PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Andersen, J.F, Weichsel, A, Champagne, D.E, Balfour, C.A, Montfort, W.R. | | Deposit date: | 1998-07-29 | | Release date: | 1998-08-05 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The crystal structure of nitrophorin 4 at 1.5 A resolution: transport of nitric oxide by a lipocalin-based heme protein.
Structure, 6, 1998
|
|
2SPZ
 
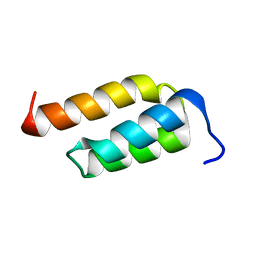 | | STAPHYLOCOCCAL PROTEIN A, Z-DOMAIN, NMR, 10 STRUCTURES | | Descriptor: | IMMUNOGLOBULIN G BINDING PROTEIN A | | Authors: | Montelione, G.T, Tashiro, M, Tejero, R, Lyons, B.A. | | Deposit date: | 1998-07-29 | | Release date: | 1998-08-05 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | High-resolution solution NMR structure of the Z domain of staphylococcal protein A.
J.Mol.Biol., 272, 1997
|
|
1PFO
 
 | | PERFRINGOLYSIN O | | Descriptor: | PERFRINGOLYSIN O | | Authors: | Rossjohn, J, Parker, M.W. | | Deposit date: | 1997-07-31 | | Release date: | 1998-08-05 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of a cholesterol-binding, thiol-activated cytolysin and a model of its membrane form.
Cell(Cambridge,Mass.), 89, 1997
|
|
1BM7
 
 | |
1BMZ
 
 | |
1BN8
 
 | | BACILLUS SUBTILIS PECTATE LYASE | | Descriptor: | CALCIUM ION, PROTEIN (PECTATE LYASE) | | Authors: | Pickersgill, R, Harris, G, Jenkins, J. | | Deposit date: | 1998-07-31 | | Release date: | 1998-08-05 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The structure of Bacillus subtilis pectate lyase in complex with calcium.
Nat.Struct.Biol., 1, 1994
|
|
1BNX
 
 | | STRUCTURAL STUDIES ON THE EFFECTS OF THE DELETION IN THE RED CELL ANION EXCHANGER (BAND3, AE1) ASSOCIATED WITH SOUTH EAST ASIAN OVALOCYTOSIS. | | Descriptor: | PROTEIN (BAND 3) | | Authors: | Chambers, E.J, Bloomberg, G.B, Ring, S.M, Tanner, M.J.A. | | Deposit date: | 1998-07-30 | | Release date: | 1998-08-05 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Studies on the structure of a transmembrane region and a cytoplasmic loop of the human red cell anion exchanger (band 3, AE1).
Biochem.Soc.Trans., 26, 1998
|
|
1BM4
 
 | | MOMLV CAPSID PROTEIN MAJOR HOMOLOGY REGION PEPTIDE ANALOG | | Descriptor: | PROTEIN (MOLONEY MURINE LEUKEMIA VIRUS CAPSID) | | Authors: | Clish, C.B, Peyton, D.H, Barklis, E. | | Deposit date: | 1998-07-28 | | Release date: | 1998-08-05 | | Last modified: | 2024-04-10 | | Method: | SOLUTION NMR | | Cite: | Solution structures of human immunodeficiency virus type 1 (HIV-1) and moloney murine leukemia virus (MoMLV) capsid protein major-homology-region peptide analogs by NMR spectroscopy.
Eur.J.Biochem., 257, 1998
|
|
1A4H
 
 | |
12E8
 
 | | 2E8 FAB FRAGMENT | | Descriptor: | IGG1-KAPPA 2E8 FAB (HEAVY CHAIN), IGG1-KAPPA 2E8 FAB (LIGHT CHAIN) | | Authors: | Rupp, B, Trakhanov, S. | | Deposit date: | 1998-03-14 | | Release date: | 1998-08-05 | | Last modified: | 2023-08-02 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of a monoclonal 2E8 Fab antibody fragment specific for the low-density lipoprotein-receptor binding region of apolipoprotein E refined at 1.9 A.
Acta Crystallogr.,Sect.D, null, 1999
|
|
377D
 
 | | 5'-R(*CP*GP*UP*AP*CP*DG)-3' | | Descriptor: | RNA-DNA (5'-R(*CP*GP*UP*AP*CP*DG)-3') | | Authors: | Biswas, R, Mitra, S.N, Sundaralingam, M. | | Deposit date: | 1998-01-26 | | Release date: | 1998-08-10 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | 1.76 A structure of a pyrimidine start alternating A-RNA hexamer r(CGUAC)dG.
Acta Crystallogr.,Sect.D, 54, 1998
|
|
335D
 
 | |
1HSW
 
 | |
5REQ
 
 | | Methylmalonyl-COA MUTASE, Y89F Mutant, substrate complex | | Descriptor: | COBALAMIN, GLYCEROL, METHYLMALONYL(CARBADETHIA)-COENZYME A, ... | | Authors: | Evans, P.R, Thomae, N.H. | | Deposit date: | 1998-08-03 | | Release date: | 1998-08-12 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Stabilization of radical intermediates by an active-site tyrosine residue in methylmalonyl-CoA mutase.
Biochemistry, 37, 1998
|
|
1IOJ
 
 | | HUMAN APOLIPOPROTEIN C-I, NMR, 18 STRUCTURES | | Descriptor: | APOC-I | | Authors: | Rozek, A, Sparrow, J.T, Weisgraber, K.H, Cushley, R.J. | | Deposit date: | 1998-05-12 | | Release date: | 1998-08-12 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Conformation of human apolipoprotein C-I in a lipid-mimetic environment determined by CD and NMR spectroscopy.
Biochemistry, 38, 1999
|
|
2BQJ
 
 | |
2BQA
 
 | |
