7GT8
 
 | |
7GT7
 
 | |
7GS8
 
 | | PanDDA Analysis group deposition -- Crystal structure of PTP1B in complex with FMOPL000466a | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Tyrosine-protein phosphatase non-receptor type 1, ~{N},~{N},5,6-tetramethylthieno[2,3-d]pyrimidin-4-amine | | Authors: | Mehlman, T, Ginn, H.M, Keedy, D.A. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.67 Å) | | Cite: | An expanded view of ligandability in the allosteric enzyme PTP1B from computational reanalysis of large-scale crystallographic data.
Biorxiv, 2024
|
|
7GSI
 
 | |
7GTU
 
 | |
7GS9
 
 | |
7GSE
 
 | |
7GSG
 
 | |
4Q8H
 
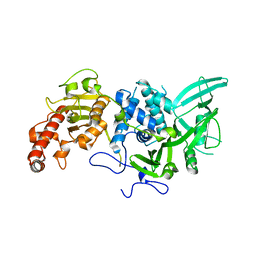 | |
4Q8G
 
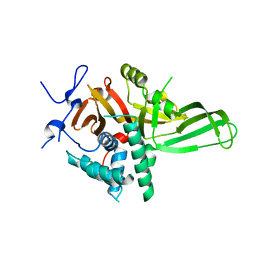 | |
7GSA
 
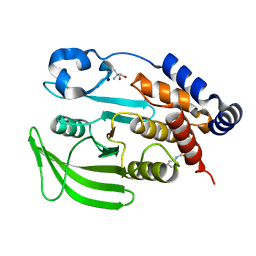 | |
7GS7
 
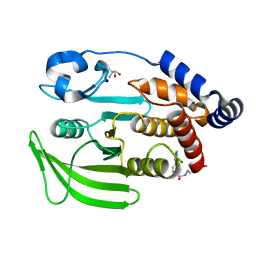 | | PanDDA Analysis group deposition -- Crystal structure of PTP1B in complex with FMOPL000621a | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 4-(1,2,3-thiadiazol-4-yl)phenyl ethylcarbamate, Tyrosine-protein phosphatase non-receptor type 1 | | Authors: | Mehlman, T, Ginn, H.M, Keedy, D.A. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | An expanded view of ligandability in the allosteric enzyme PTP1B from computational reanalysis of large-scale crystallographic data.
Biorxiv, 2024
|
|
7GSQ
 
 | |
7GTD
 
 | |
7GSU
 
 | |
7GTS
 
 | |
7GSY
 
 | |
7GT5
 
 | | PanDDA Analysis group deposition -- Crystal structure of PTP1B in complex with FMOOA000529a | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Tyrosine-protein phosphatase non-receptor type 1, methyl [(3R,4S)-3-ethyl-4-hydroxy-1,1-dioxo-3,4-dihydro-1lambda~6~,2-benzothiazin-2(1H)-yl]acetate | | Authors: | Mehlman, T, Ginn, H.M, Keedy, D.A. | | Deposit date: | 2024-01-03 | | Release date: | 2024-01-24 | | Last modified: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.61 Å) | | Cite: | An expanded view of ligandability in the allosteric enzyme PTP1B from computational reanalysis of large-scale crystallographic data.
Biorxiv, 2024
|
|
3X1K
 
 | |
5AZI
 
 | | Crystal structure of glycerol kinase from Trypanosoma brucei gambiense complexed with 4NP | | Descriptor: | 4-NITROPHENYL PHOSPHATE, GLYCEROL, Glycerol kinase | | Authors: | Balogun, E.O, Inaoka, D.K, Shiba, T, Tokuoka, S.M, Tokumasu, F, Sakamoto, K, Michels, P.A.M, Harada, S, Kita, K. | | Deposit date: | 2015-10-08 | | Release date: | 2016-10-19 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Glycerol kinase of African trypanosomes possesses an intrinsic phosphatase activity.
Biochim Biophys Acta Gen Subj, 1861, 2017
|
|
5AZJ
 
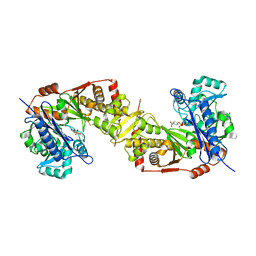 | | Crystal structure of glycerol kinase from Trypanosoma brucei gambiense complexed with 4NP (with disulfide bridge) | | Descriptor: | 4-NITROPHENYL PHOSPHATE, GLYCEROL, Glycerol kinase | | Authors: | Balogun, E.O, Inaoka, D.K, Shiba, T, Tokuoka, S.M, Tokumasu, F, Sakamoto, K, Michels, P.A.M, Harada, S, Kita, K. | | Deposit date: | 2015-10-08 | | Release date: | 2016-10-19 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.61 Å) | | Cite: | Glycerol kinase of African trypanosomes possesses an intrinsic phosphatase activity.
Biochim Biophys Acta Gen Subj, 1861, 2017
|
|
3X1J
 
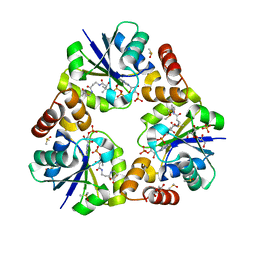 | |
4E1A
 
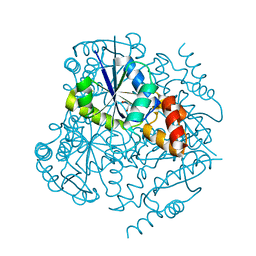 | | Phosphopantetheine adenylyltransferase from Mycobacterium tuberculosis at 1.62A resolution | | Descriptor: | GLYCEROL, Phosphopantetheine adenylyltransferase | | Authors: | Timofeev, V.I, Smirnova, E.A, Chupova, L.A, Esipov, R.S, Kuranova, I.P. | | Deposit date: | 2012-03-06 | | Release date: | 2012-11-21 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | X-ray study of the conformational changes in the molecule of phosphopantetheine adenylyltransferase from Mycobacterium tuberculosis during the catalyzed reaction.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
4FAY
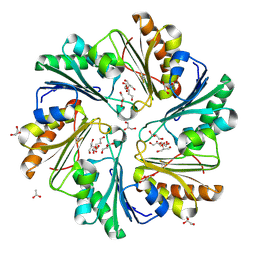 
 | | Crystal structure of a trimeric bacterial microcompartment shell protein PduB with glycerol metabolites | | Descriptor: | ACETATE ION, GLYCEROL, Microcompartments protein | | Authors: | Pang, A.H, Prentice, M.B, Pickersgill, R.W. | | Deposit date: | 2012-05-22 | | Release date: | 2012-11-21 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Substrate channels revealed in the trimeric Lactobacillus reuteri bacterial microcompartment shell protein PduB.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
5NG1
 
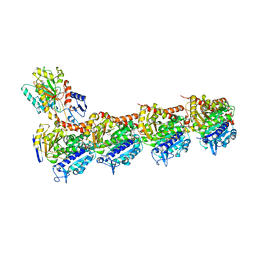 | | TUBULIN-MTC-zampanolide complex | | Descriptor: | (2Z,4E)-N-[(S)-[(1S,2E,5S,8E,10Z,17S)-3,11-dimethyl-19-methylidene-7,13-dioxo-6,21-dioxabicyclo[15.3.1]henicosa-2,8,10-trien-5-yl](hydroxy)methyl]hexa-2,4-dienamide, (2~{Z},4~{E})-~{N}-[(~{S})-oxidanyl-[(1~{S},2~{E},5~{S},11~{R},17~{S},19~{R})-3,11,19-trimethyl-7,13-bis(oxidanylidene)-6,21-dioxabicyclo[15.3.1]henicos-2-en-5-yl]methyl]hexa-2,4-dienamide, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, ... | | Authors: | Field, J.J, Pera, B, Estevez Gallego, J, Calvo, E, Rodriguez-Salarichs, J, Saez-Calvo, G, Zuwerra, D, Jordi, M, Prota, A.E, Menchon, G, Miller, J.H, Altmann, K.-H, Diaz, J.F. | | Deposit date: | 2017-03-16 | | Release date: | 2017-10-25 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Zampanolide Binding to Tubulin Indicates Cross-Talk of Taxane Site with Colchicine and Nucleotide Sites.
J. Nat. Prod., 81, 2018
|
|
