5R93
 
 | |
5R9K
 
 | |
5RA2
 
 | |
5ZEO
 
 | |
5R8V
 
 | |
5R9A
 
 | |
5R9Q
 
 | |
5RA5
 
 | |
5R8Z
 
 | |
5R9E
 
 | |
5R9U
 
 | |
5RA8
 
 | |
5R8W
 
 | |
5R9C
 
 | |
5R9R
 
 | |
5RA7
 
 | |
5R97
 
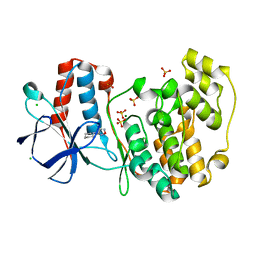 | | PanDDA analysis group deposition Form1 MAP kinase p38-alpha -- Fragment N13662a in complex with MAP kinase p38-alpha | | Descriptor: | 4-(piperidin-1-yl)-1,2,5-oxadiazol-3-amine, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | De Nicola, G.F, Nichols, C.E. | | Deposit date: | 2020-03-04 | | Release date: | 2020-07-22 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.438 Å) | | Cite: | Mining the PDB for Tractable Cases Where X-ray Crystallography Combined with Fragment Screens Can Be Used to Systematically Design Protein-Protein Inhibitors: Two Test Cases Illustrated by IL1 beta-IL1R and p38 alpha-TAB1 Complexes.
J.Med.Chem., 63, 2020
|
|
5R9L
 
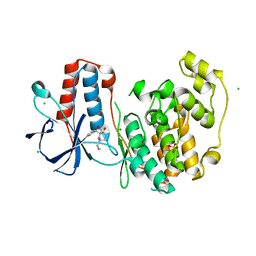 | |
5RA3
 
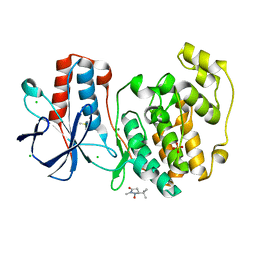 | |
5R95
 
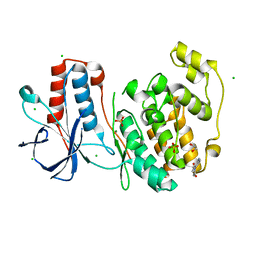 | | PanDDA analysis group deposition Form1 MAP kinase p38-alpha -- Fragment KCL093 in complex with MAP kinase p38-alpha | | Descriptor: | 1,2-ETHANEDIOL, 1-(4-aminophenyl)pyrrole-2,5-dione, CHLORIDE ION, ... | | Authors: | De Nicola, G.F, Nichols, C.E. | | Deposit date: | 2020-03-04 | | Release date: | 2020-07-22 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.59 Å) | | Cite: | Mining the PDB for Tractable Cases Where X-ray Crystallography Combined with Fragment Screens Can Be Used to Systematically Design Protein-Protein Inhibitors: Two Test Cases Illustrated by IL1 beta-IL1R and p38 alpha-TAB1 Complexes.
J.Med.Chem., 63, 2020
|
|
5R9G
 
 | | PanDDA analysis group deposition Form1 MAP kinase p38-alpha -- Fragment PC587 in complex with MAP kinase p38-alpha | | Descriptor: | (4R,4aS,7aS,9S)-3,10-dimethyl-5,6,7,7a,8,9-hexahydro-4H-4a,9-epiminopyrrolo[3',4':5,6]cyclohepta[1,2-d][1,2]oxazol-4-ol, CHLORIDE ION, DIMETHYL SULFOXIDE, ... | | Authors: | De Nicola, G.F, Nichols, C.E. | | Deposit date: | 2020-03-04 | | Release date: | 2020-07-22 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Mining the PDB for Tractable Cases Where X-ray Crystallography Combined with Fragment Screens Can Be Used to Systematically Design Protein-Protein Inhibitors: Two Test Cases Illustrated by IL1 beta-IL1R and p38 alpha-TAB1 Complexes.
J.Med.Chem., 63, 2020
|
|
5R9X
 
 | |
5R94
 
 | |
5R9H
 
 | |
5R9Y
 
 | |
