6SZC
 
 | |
5HUZ
 
 | | Solution structure of coiled coil domain of myosin binding subunit of myosin light chain phosphatase | | Descriptor: | Protein phosphatase 1 regulatory subunit 12A | | Authors: | Sharma, A.K, Birrane, G, Anklin, C, Rigby, A.C, Pollak, M, Alper, S.L. | | Deposit date: | 2016-01-27 | | Release date: | 2016-03-02 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | To be published
To Be Published
|
|
5NMY
 
 | |
5LXL
 
 | | NMR structure of the N-terminal domain of the Bacteriophage T5 decoration protein pb10 | | Descriptor: | Decoration protein | | Authors: | Vernhes, E, Gilquin, B, Cuniasse, P, Boulanger, P, Zinn-Justin, S. | | Deposit date: | 2016-09-22 | | Release date: | 2017-04-19 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | High affinity anchoring of the decoration protein pb10 onto the bacteriophage T5 capsid.
Sci Rep, 7, 2017
|
|
5LW8
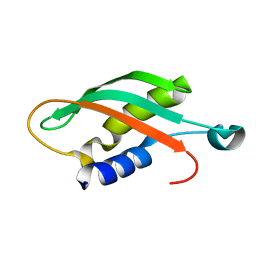 
 | |
6RC7
 
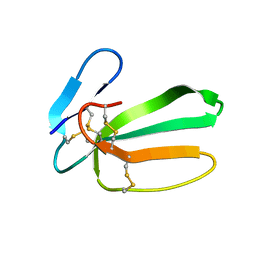 | | NMR structure of cytotoxin 3 from Naja kaouthia in solution, major form | | Descriptor: | Cytotoxin 3 | | Authors: | Dubinnyi, M.A, Dubovskii, P.V, Utkin, Y.N, Arseniev, A.S. | | Deposit date: | 2019-04-11 | | Release date: | 2020-05-13 | | Last modified: | 2021-05-26 | | Method: | SOLUTION NMR | | Cite: | The omega-loop of cobra cytotoxins tolerates multiple amino acid substitutions.
Biochem.Biophys.Res.Commun., 558, 2021
|
|
6S10
 
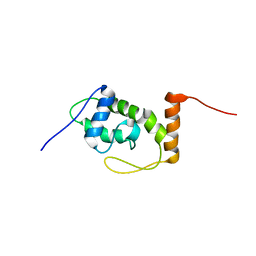 | |
5Y70
 
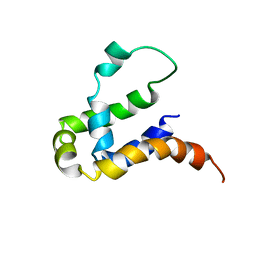 | | NMR structure of KMP11 in DPC micelle | | Descriptor: | Kinetoplastid membrane protein 11 | | Authors: | Lu, Y, Lim, L.Z, Song, J. | | Deposit date: | 2017-08-16 | | Release date: | 2018-02-07 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Kinetoplastid membrane protein-11 adopts a four-helix bundle fold in DPC micelle.
FEBS Lett., 591, 2017
|
|
6BA3
 
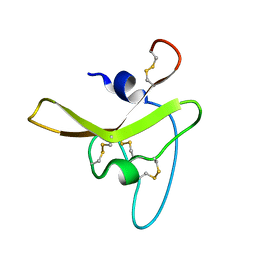 | |
5KI0
 | | NMR structure of human antimicrobial peptide KAMP-19 | | Descriptor: | Antimicrobial peptide KAMP-19 | | Authors: | Wang, G. | | Deposit date: | 2016-06-16 | | Release date: | 2016-11-23 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Membrane-Active Epithelial Keratin 6A Fragments (KAMPs) Are Unique Human Antimicrobial Peptides with a Non-alpha beta Structure.
Front Microbiol, 7, 2016
|
|
5L82
 
 | |
5KWX
 
 | |
5KWO
 
 | |
5KX2
 | |
5KWZ
 
 | |
5KX0
 
 | |
5KWP
 
 | |
5KX1
 | |
5KVN
 | |
5LCS
 
 | |
6E3C
 
 | |
7KPD
 
 | |
7LHC
 
 | |
5J7J
 
 | |
6PIP
 
 | |
