8SMP
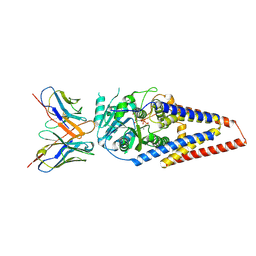 
 | | Xenopus laevis hyaluronan synthase 1, UDP-bound, gating loop inserted state | | Descriptor: | Fab15 heavy chain, Fab15 light chain, Hyaluronan synthase 1, ... | | Authors: | Gorniak, I, Zimmer, J. | | Deposit date: | 2023-04-26 | | Release date: | 2024-05-01 | | Last modified: | 2024-10-09 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Atomistic insights into hyaluronan synthesis, secretion, and length control
To Be Published
|
|
4P00
 
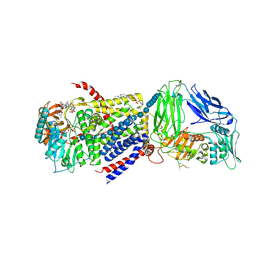 | | Bacterial Cellulose Synthase in complex with cyclic-di-GMP and UDP | | Descriptor: | 1,2-Distearoyl-sn-glycerophosphoethanolamine, 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), Cellulose Synthase A subunit, ... | | Authors: | Morgan, J.L.W, McNamara, J.T, Zimmer, J. | | Deposit date: | 2014-02-19 | | Release date: | 2014-04-09 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Mechanism of activation of bacterial cellulose synthase by cyclic di-GMP.
Nat.Struct.Mol.Biol., 21, 2014
|
|
4P02
 
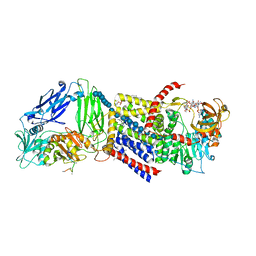 | | Structure of Bacterial Cellulose Synthase with cyclic-di-GMP bound. | | Descriptor: | 1,2-Distearoyl-sn-glycerophosphoethanolamine, 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), Cellulose Synthase subunit A, ... | | Authors: | Morgan, J.L.W, McNamara, J.T, Zimmer, J. | | Deposit date: | 2014-02-20 | | Release date: | 2014-04-09 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Mechanism of activation of bacterial cellulose synthase by cyclic di-GMP.
Nat.Struct.Mol.Biol., 21, 2014
|
|
1OOA
 
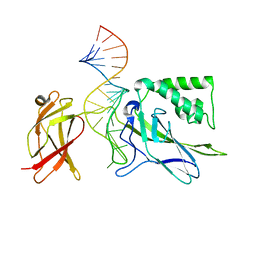 | | CRYSTAL STRUCTURE OF NF-kB(p50)2 COMPLEXED TO A HIGH-AFFINITY RNA APTAMER | | Descriptor: | Nuclear factor NF-kappa-B p105 subunit, RNA aptamer | | Authors: | Huang, D.B, Vu, D, Cassiday, L.A, Zimmerman, J.M, Maher III, L.J, Ghosh, G. | | Deposit date: | 2003-03-03 | | Release date: | 2003-07-22 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Crystal structure of NF-kappaB (p50)2 complexed to a high-affinity RNA aptamer.
Proc.Natl.Acad.Sci.USA, 100, 2003
|
|
2JP2
 
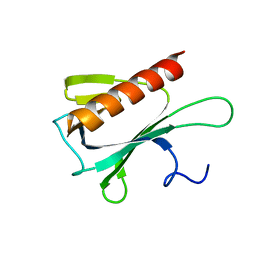 | | Solution structure and resonance assignment of the N-terminal EVH1 domain from the human Spred2 protein (Sprouty-related protein with EVH1 domain isoform 2) | | Descriptor: | Sprouty-related, EVH1 domain-containing protein 2 | | Authors: | Fossi, M, Zimmermann, J, Jarchau, T, Lemak, A, Walter, U, Wiegelt, J, Sundstrom, M, Arrowsmith, C, Edwards, A, Oschkinat, H, Ball, L.J, Structural Genomics Consortium (SGC) | | Deposit date: | 2007-04-18 | | Release date: | 2007-05-15 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | 1H, 13C and 15N resonance assignment of the human Spred2 EVH1 domain
J.Biomol.NMR, 29, 2004
|
|
2GTO
 
 | |
4NJ9
 
 | | Crystal structure of Fab 8B10 in complex with MPTS | | Descriptor: | 8-methoxypyrene-1,3,6-trisulfonic acid, 8B10 heavy chain, 8B10 light chain, ... | | Authors: | Stanfield, R.L, Romesberg, F.E, Zimmermann, J, Wilson, I.A. | | Deposit date: | 2013-11-08 | | Release date: | 2014-11-12 | | Last modified: | 2017-06-14 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Adaptive Mutations Alter Antibody Structure and Dynamics during Affinity Maturation.
Biochemistry, 54, 2015
|
|
4NJA
 
 | | Crystal structure of Fab 6C8 in complex with MPTS | | Descriptor: | 6C8 heavy chain, 6C8 light chain, 8-methoxypyrene-1,3,6-trisulfonic acid, ... | | Authors: | Stanfield, R.L, Romesberg, F.E, Zimmermann, J, Wilson, I.A. | | Deposit date: | 2013-11-08 | | Release date: | 2014-11-12 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.204 Å) | | Cite: | Adaptive Mutations Alter Antibody Structure and Dynamics during Affinity Maturation.
Biochemistry, 54, 2015
|
|
