4X3A
 
 | |
4X37
 
 | |
4X38
 
 | |
4X39
 
 | |
2B9B
 
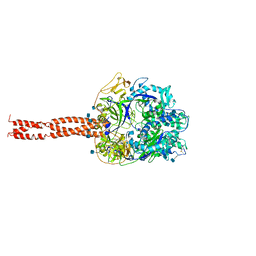 | | Structure of the Parainfluenza Virus 5 F Protein in its Metastable, Pre-fusion Conformation | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Fusion glycoprotein F0 | | Authors: | Yin, H.-S, Wen, X, Paterson, R.G, Lamb, R.A, Jardetzky, T.S. | | Deposit date: | 2005-10-11 | | Release date: | 2006-01-24 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Structure of the parainfluenza virus 5 F protein in its metastable, prefusion conformation
Nature, 439, 2006
|
|
1ZTM
 
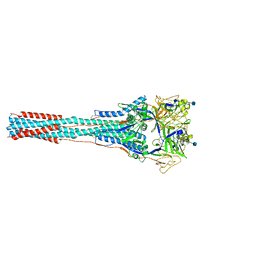 | | Structure of the Uncleaved Paramyxovirus (hPIV3) Fusion Protein | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Fusion glycoprotein | | Authors: | Yin, H.S, Paterson, R.G, Wen, X, Lamb, R.A, Jardetzky, T.S. | | Deposit date: | 2005-05-27 | | Release date: | 2005-07-19 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (3.05 Å) | | Cite: | Structure of the uncleaved ectodomain of the paramyxovirus (hPIV3) fusion protein
Proc.Natl.Acad.Sci.USA, 102, 2005
|
|
3NJ5
 
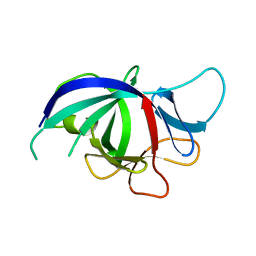 | |
3OTW
 
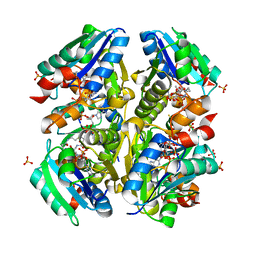 | | Structural and Functional Studies of Helicobacter pylori Wild-Type and Mutated Proteins Phosphopantetheine adenylyltransferase | | Descriptor: | COENZYME A, Phosphopantetheine adenylyltransferase, SULFATE ION | | Authors: | Yin, H.S, Cheng, C.S, Chen, C.G, Luo, Y.C, Chen, W.T, Cheng, S.Y. | | Deposit date: | 2010-09-14 | | Release date: | 2011-09-14 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural and Functional Studies of Helicobacter pylori Wild-Type and Mutated Proteins Phosphopantetheine adenylyltransferase
To be Published
|
|
8JA0
 
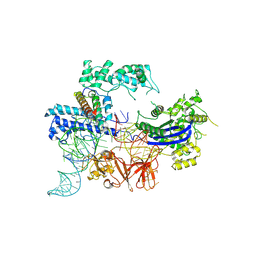 | | Cryo-EM structure of the NmeCas9-sgRNA-AcrIIC4 ternary complex | | Descriptor: | CRISPR-associated endonuclease Cas9, RNA (117-MER), Uncharacterized protein | | Authors: | Yin, H, Li, Z, Yu, G.M, Li, X.Z. | | Deposit date: | 2023-05-05 | | Release date: | 2023-08-09 | | Last modified: | 2025-02-26 | | Method: | ELECTRON MICROSCOPY (3.52 Å) | | Cite: | Inhibitory mechanism of CRISPR-Cas9 by AcrIIC4.
Nucleic Acids Res., 51, 2023
|
|
7CPR
 
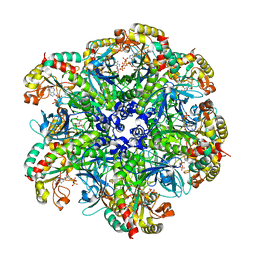 | | glutamine synthetase from Drosophila | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Glutamine synthetase 2 cytoplasmic | | Authors: | Yin, H.S, Chen, W.T. | | Deposit date: | 2020-08-07 | | Release date: | 2021-08-11 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | Structural Insight into the Contributions of the N-Terminus and Key Active-Site Residues to the Catalytic Efficiency of Glutamine Synthetase 2.
Biomolecules, 10, 2020
|
|
8XSV
 
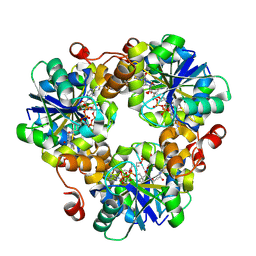 | | crystal structure of PPAT mutant P8A | | Descriptor: | DEPHOSPHO COENZYME A, Phosphopantetheine adenylyltransferase | | Authors: | Yin, H.S. | | Deposit date: | 2024-01-10 | | Release date: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | crystal structure of PPAT mutant P8A
To Be Published
|
|
8XRW
 
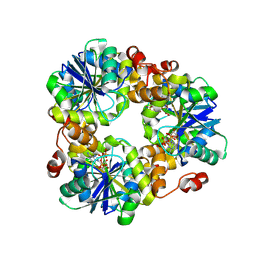 | | crystal structure of HpPPAT in complex with ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Phosphopantetheine adenylyltransferase | | Authors: | Yin, H.S. | | Deposit date: | 2024-01-08 | | Release date: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | crystal structure of HpPPAT in complex with ATP
To Be Published
|
|
8XSK
 
 | | crystal structure of PPAT | | Descriptor: | Phosphopantetheine adenylyltransferase | | Authors: | Yin, H.S. | | Deposit date: | 2024-01-09 | | Release date: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | crystal structure of PPAT
To Be Published
|
|
8XSC
 
 | | PPAT in complex with Ppant | | Descriptor: | 4'-PHOSPHOPANTETHEINE, Phosphopantetheine adenylyltransferase | | Authors: | Yin, H.S. | | Deposit date: | 2024-01-09 | | Release date: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | PPAT in complex with Ppant
To Be Published
|
|
5GZJ
 
 | | Cyclodeaminase_PA | | Descriptor: | Lysine cyclodeaminase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Ying, H, Chen, K. | | Deposit date: | 2016-09-28 | | Release date: | 2018-01-17 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.87 Å) | | Cite: | Cyclodeaminase_PA
To Be Published
|
|
5GZM
 
 | | Cyclodeaminase_PA | | Descriptor: | (2S)-piperidine-2-carboxylic acid, Lysine cyclodeaminase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Ying, H, Chen, K. | | Deposit date: | 2016-09-29 | | Release date: | 2018-01-17 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Cyclodeaminase_PA
To Be Published
|
|
5GZL
 
 | | Cyclodeaminase_PA | | Descriptor: | LYSINE, Lysine cyclodeaminase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Ying, H, Chen, K. | | Deposit date: | 2016-09-29 | | Release date: | 2018-01-17 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Cyclodeaminase_PA
To Be Published
|
|
5GZI
 
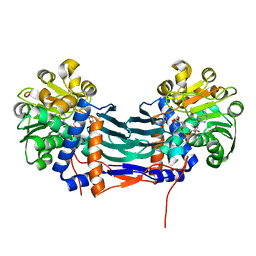 | | Cyclodeaminase_PA | | Descriptor: | (2S)-piperidine-2-carboxylic acid, Lysine cyclodeaminase, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Ying, H, Chen, K. | | Deposit date: | 2016-09-28 | | Release date: | 2018-01-17 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.17 Å) | | Cite: | Cyclodeaminase_PA
To Be Published
|
|
2WRY
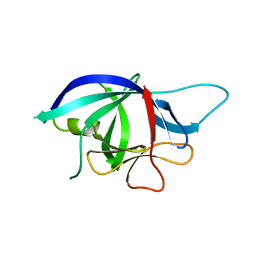 
 | | Crystal structure of chicken cytokine interleukin 1 beta | | Descriptor: | INTERLEUKIN-1BETA | | Authors: | Lu, W.S, Cheng, C.S, Lyu, P.C, Lee, L.H, Wang, W.C, Yin, H.S. | | Deposit date: | 2009-09-03 | | Release date: | 2010-09-29 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Structural and Functional Comparison of Cytokine Interleukin-1 Beta from Chicken and Human.
Mol.Immunol., 48, 2011
|
|
9JQT
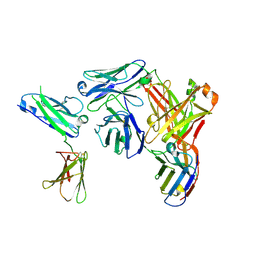 
 | | Structure of interleukin receptor common gamma chain (IL2Rgamma/CD132) in complex with 2D4 | | Descriptor: | 2D4-Heavy Chain, 2D4-Light Chain, Anti-Fab nanobody, ... | | Authors: | Lu, Q.J, Yin, H.Q. | | Deposit date: | 2024-09-28 | | Release date: | 2024-12-18 | | Method: | ELECTRON MICROSCOPY (2.7 Å) | | Cite: | 2D4, a humanized monoclonal antibody targeting CD132, is a promising treatment for systemic lupus erythematosus.
Signal Transduct Target Ther, 9, 2024
|
|
1H1V
 
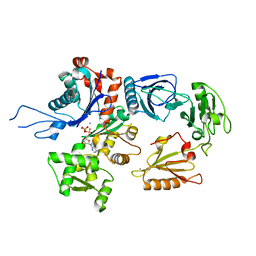 | | gelsolin G4-G6/actin complex | | Descriptor: | ACTIN, ADENOSINE-5'-TRIPHOSPHATE, CALCIUM ION, ... | | Authors: | Choe, H, Burtnick, L.D, Mejillano, M, Yin, H.L, Robinson, R.C, Choe, S. | | Deposit date: | 2002-07-23 | | Release date: | 2003-01-24 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.99 Å) | | Cite: | The Calcium Activation of Gelsolin:Insights from the 3A Structure of the G4-G6/Actin Complex
J.Mol.Biol., 324, 2002
|
|
1W63
 
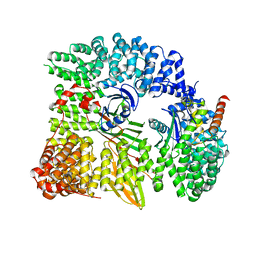 | | AP1 clathrin adaptor core | | Descriptor: | ADAPTER-RELATED PROTEIN COMPLEX 1 BETA 1 SUBUNIT, ADAPTER-RELATED PROTEIN COMPLEX 1 GAMMA 1 SUBUNIT, ADAPTER-RELATED PROTEIN COMPLEX 1 SIGMA 1A SUBUNIT, ... | | Authors: | Heldwein, E, Macia, E, Wang, J, Yin, H.L, Kirchhausen, T, Harrison, S.C. | | Deposit date: | 2004-08-12 | | Release date: | 2004-09-21 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (4 Å) | | Cite: | Crystal Structure of the Clathrin Adaptor Protein 1 Core
Proc.Natl.Acad.Sci.USA, 101, 2004
|
|
3NV7
 
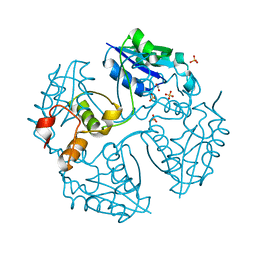 | |
3RVG
 
 | | Crystals structure of Jak2 with a 1-amino-5H-pyrido[4,3-b]indol-4-carboxamide inhibitor | | Descriptor: | 1-(cyclohexylamino)-7-(1-methyl-1H-pyrazol-4-yl)-5H-pyrido[4,3-b]indole-4-carboxamide, Tyrosine-protein kinase JAK2 | | Authors: | Lim, J, Taoka, B, Otte, R.D, Spencer, K, Dinsmore, C.J, Altman, M.D, Chan, G, Rosenstein, C, Sharma, S, Su, H.P, Szewczak, A.A, Xu, L, Yin, H, Zugay-Murphy, J, Marshall, C.G, Young, J.R. | | Deposit date: | 2011-05-06 | | Release date: | 2012-03-21 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.498 Å) | | Cite: | Discovery of 1-amino-5H-pyrido[4,3-b]indol-4-carboxamide inhibitors of Janus kinase 2 (JAK2) for the treatment of myeloproliferative disorders.
J.Med.Chem., 54, 2011
|
|
5WSZ
 
 | | Crystal structure of a lytic polysaccharide monooxygenase,BtLPMO10A, from Bacillus thuringiensis | | Descriptor: | COPPER (II) ION, LpmO10A | | Authors: | Zhao, Y, Zhang, H, Yin, H. | | Deposit date: | 2016-12-08 | | Release date: | 2017-02-22 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.565 Å) | | Cite: | Crystal structure of a lytic polysaccharide monooxygenase,BtLPMO10A, from Bacillus thuringiensis
To Be Published
|
|
