3AQ3
 | |
3AQ4
 
 | |
3AQ2
 
 | |
7WIT
 
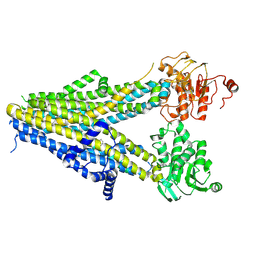 | | Structure of SUR1 in complex with mitiglinide | | Descriptor: | (2S)-4-[(3aR,7aS)-1,3,3a,4,5,6,7,7a-octahydroisoindol-2-yl]-4-oxidanylidene-2-(phenylmethyl)butanoic acid, ADENOSINE-5'-TRIPHOSPHATE, ATP-sensitive inward rectifier potassium channel 11,ATP-binding cassette sub-family C member 8 isoform X1 | | Authors: | Chen, L, Wang, M.M. | | Deposit date: | 2022-01-04 | | Release date: | 2022-11-16 | | Last modified: | 2024-06-26 | | Method: | ELECTRON MICROSCOPY (3.21 Å) | | Cite: | Structural Insights Into the High Selectivity of the Anti-Diabetic Drug Mitiglinide
Front Pharmacol, 13, 2022
|
|
