2HTN
 
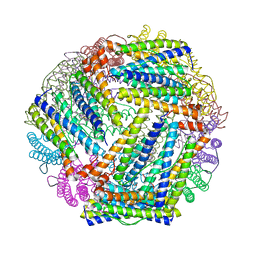 | | E. coli bacterioferritin in its as-isolated form | | Descriptor: | Bacterioferritin, FE (III) ION, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | van Eerde, A, Wolterink-Van Loo, S, Van Der Oost, J, Dijkstra, B.W. | | Deposit date: | 2006-07-26 | | Release date: | 2006-11-21 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Fortuitous structure determination of 'as-isolated' Escherichia coli bacterioferritin in a novel crystal form.
Acta Crystallogr.,Sect.F, 62, 2006
|
|
2NUX
 
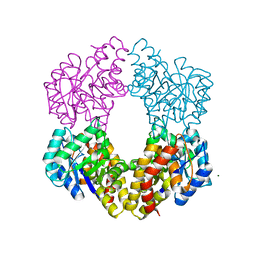 | |
2NUW
 
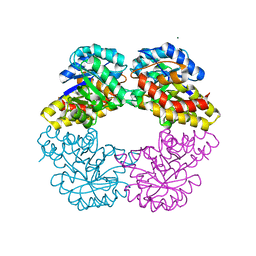 | |
2NUY
 
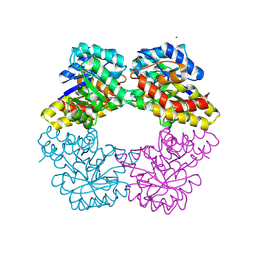 | |
1RY9
 
 | | Spa15, a Type III Secretion Chaperone from Shigella flexneri | | Descriptor: | CHLORIDE ION, Surface presentation of antigens protein spaK | | Authors: | van Eerde, A, Hamiaux, C, Perez, J, Parsot, C, Dijkstra, B.W. | | Deposit date: | 2003-12-20 | | Release date: | 2004-04-27 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Structure of Spa15, a type III secretion chaperone from Shigella flexneri with broad specificity.
Embo Rep., 5, 2004
|
|
4NDU
 
 | |
4NDV
 
 | |
4NDS
 
 | |
4NDT
 
 | |
2GDC
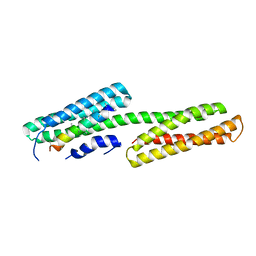 
 | | Structure of Vinculin VD1 / IpaA560-633 complex | | Descriptor: | Invasin ipaA, Vinculin | | Authors: | Hamiaux, C, van Eerde, A, Parsot, C, Broos, J, Dijkstra, B.W. | | Deposit date: | 2006-03-15 | | Release date: | 2006-08-08 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.74 Å) | | Cite: | Structural mimicry for vinculin activation by IpaA, a virulence factor of Shigella flexneri.
Embo Rep., 7, 2006
|
|
5D61
 
 | | MOA-Z-VAD-fmk complex, direct orientation | | Descriptor: | 1,2-ETHANEDIOL, Agglutinin, CALCIUM ION, ... | | Authors: | Cordara, G, van Eerde, A, Grahn, E.M, Goldstein, I.J, Krengel, U. | | Deposit date: | 2015-08-11 | | Release date: | 2016-03-02 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | An Unusual Member of the Papain Superfamily: Mapping the Catalytic Cleft of the Marasmius oreades agglutinin (MOA) with a Caspase Inhibitor.
Plos One, 11, 2016
|
|
