6UUG
 
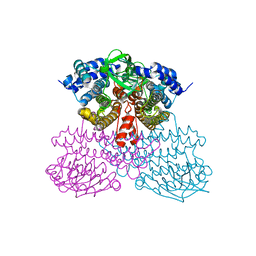 | | Structure of methanesulfinate monooxygenase MsuC from Pseudomonas fluorescens at 1.69 angstrom resolution | | Descriptor: | Putative dehydrogenase | | Authors: | Soule, J, Gnann, A.D, Gonzalez, R, Parker, M.J, McKenna, K.C, Nguyen, S.V, Phan, N.T, Wicht, D.K, Dowling, D.P. | | Deposit date: | 2019-10-30 | | Release date: | 2019-12-04 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.685 Å) | | Cite: | Structure and function of the two-component flavin-dependent methanesulfinate monooxygenase within bacterial sulfur assimilation.
Biochem.Biophys.Res.Commun., 522, 2020
|
|
1ZN7
 
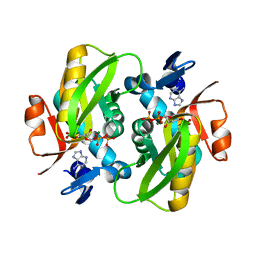 | | Human Adenine Phosphoribosyltransferase Complexed with PRPP, ADE and R5P | | Descriptor: | 1-O-pyrophosphono-5-O-phosphono-alpha-D-ribofuranose, 5-O-phosphono-alpha-D-ribofuranose, ADENINE, ... | | Authors: | Iulek, J, Silva, M, Tomich, C.H.T.P, Thiemann, O.H. | | Deposit date: | 2005-05-11 | | Release date: | 2006-04-25 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Structural Complexes of Human Adenine Phosphoribosyltransferase Reveal Novel Features of the APRT Catalytic Mechanism
J.Biomol.Struct.Dyn., 25, 2008
|
|
6CL4
 
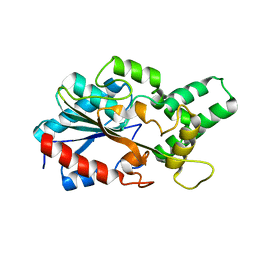 | | LipC12 - Lipase from metagenomics | | Descriptor: | Lipase C12 | | Authors: | Iulek, J, Martini, V.P, Krieger, N, Glogauer, A, Souza, E.M. | | Deposit date: | 2018-03-01 | | Release date: | 2019-03-13 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.64 Å) | | Cite: | Structure solution and analyses of the first true lipase obtained from metagenomics indicate potential for increased thermostability.
N Biotechnol, 53, 2019
|
|
1ZN9
 
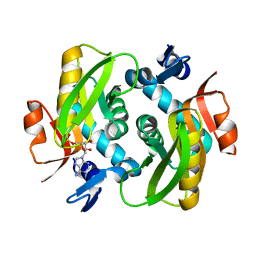 | | Human Adenine Phosphoribosyltransferase in Apo and AMP Complexed Forms | | Descriptor: | ADENOSINE MONOPHOSPHATE, Adenine phosphoribosyltransferase | | Authors: | Iulek, J, Silva, M, Tomich, C.H.T.P, Thiemann, O.H. | | Deposit date: | 2005-05-11 | | Release date: | 2006-04-25 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structural Complexes of Human Adenine Phosphoribosyltransferase Reveal Novel Features of the APRT Catalytic Mechanism
J.Biomol.Struct.Dyn., 25, 2008
|
|
1ZN8
 
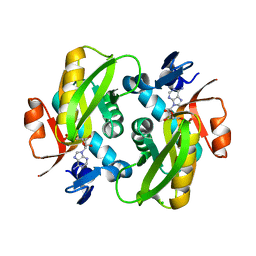 | | Human Adenine Phosphoribosyltransferase Complexed with AMP, in Space Group P1 at 1.76 A Resolution | | Descriptor: | ADENOSINE MONOPHOSPHATE, Adenine phosphoribosyltransferase, CHLORIDE ION | | Authors: | Iulek, J, Silva, M, Tomich, C.H.T.P, Thiemann, O.H. | | Deposit date: | 2005-05-11 | | Release date: | 2006-04-25 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Structural Complexes of Human Adenine Phosphoribosyltransferase Reveal Novel Features of the APRT Catalytic Mechanism
J.Biomol.Struct.Dyn., 25, 2008
|
|
6U76
 
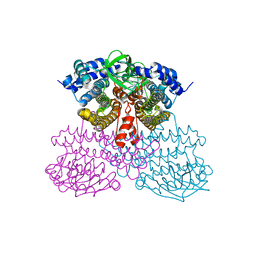 | | Structure of methanesulfinate monooxygenase MsuC from Pseudomonas fluorescens. | | Descriptor: | methanesulfinate monooxygenase | | Authors: | Soule, J, Gnann, A.D, Parker, M.J, McKenna, K.C, Nguyen, S.V, Phan, N.T, Wicht, D.K, Dowling, D.P. | | Deposit date: | 2019-08-31 | | Release date: | 2020-11-11 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | To be published
To Be Published
|
|
2F51
 
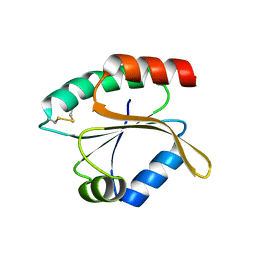 | |
3BMQ
 
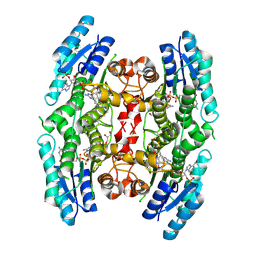 | | Structure of Pteridine Reductase 1 (PTR1) from Trypanosoma brucei in ternary complex with cofactor (NADP+) and inhibitor (Compound AX5) | | Descriptor: | 2,3-DIHYDROXY-1,4-DITHIOBUTANE, 6-(benzylsulfanyl)pyrimidine-2,4-diamine, ACETATE ION, ... | | Authors: | Martini, V.P, Iulek, J, Hunter, W.N. | | Deposit date: | 2007-12-13 | | Release date: | 2008-12-16 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure-based design of pteridine reductase inhibitors targeting african sleeping sickness and the leishmaniases.
J.Med.Chem., 53, 2010
|
|
1W1N
 
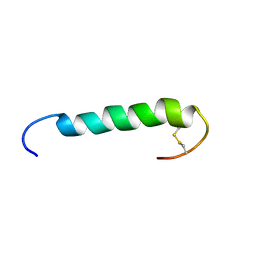 | | The solution structure of the FATC Domain of the Protein Kinase TOR1 from yeast | | Descriptor: | PHOSPHATIDYLINOSITOL 3-KINASE TOR1 | | Authors: | Dames, S.A, Mulet, J.M, Rathgeb-Szabo, K, Hall, M.N, Grzesiek, S. | | Deposit date: | 2004-06-23 | | Release date: | 2005-03-16 | | Last modified: | 2018-05-02 | | Method: | SOLUTION NMR | | Cite: | The solution structure of the FATC domain of the protein kinase target of rapamycin suggests a role for redox-dependent structural and cellular stability.
J. Biol. Chem., 280, 2005
|
|
8DE5
 
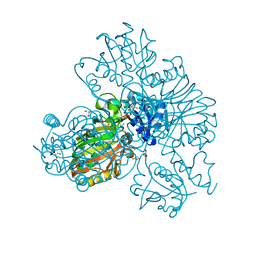 | | Structure of glyceraldehyde-3-phosphate dehydrogenase from Paracoccidioides lutzii | | Descriptor: | D-galactonic acid, GLYCEROL, Glyceraldehyde-3-phosphate dehydrogenase, ... | | Authors: | Hernandez-Prieto, J.H, Martini, V.P, Iulek, J. | | Deposit date: | 2022-06-19 | | Release date: | 2023-06-21 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | Structure of glyceraldehyde-3-phosphate dehydrogenase from Paracoccidioides lutzii in complex with an aldonic sugar acid.
Biochimie, 218, 2023
|
|
6UEK
 
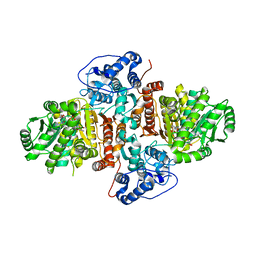 | | Structure of Urocanate Hydratase from Trypanosoma cruzi in complex with NAD+ | | Descriptor: | NICOTINAMIDE-ADENINE-DINUCLEOTIDE, Urocanate hydratase | | Authors: | Boreiko, S, Silva, M, Melo, R.F.P, Silber, A.M, Iulek, J. | | Deposit date: | 2019-09-21 | | Release date: | 2020-01-15 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Structure of Urocanate Hydratase from the protozoan Trypanosoma cruzi.
Int.J.Biol.Macromol., 146, 2019
|
|
2QVB
 
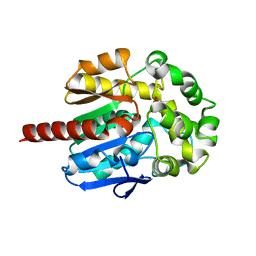 | | Crystal Structure of Haloalkane Dehalogenase Rv2579 from Mycobacterium tuberculosis | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, Haloalkane dehalogenase 3 | | Authors: | Mazumdar, P.A, Hulecki, J, Cherney, M.M, Garen, C.R, James, M.N.G, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2007-08-08 | | Release date: | 2008-02-12 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.19 Å) | | Cite: | X-ray crystal structure of Mycobacterium tuberculosis haloalkane dehalogenase Rv2579.
Biochim.Biophys.Acta, 1784, 2008
|
|
4BS2
 
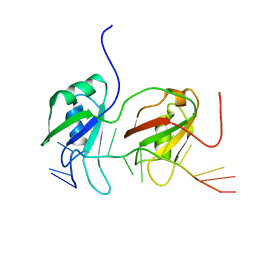 | | NMR structure of human TDP-43 tandem RRMs in complex with UG-rich RNA | | Descriptor: | 5'-R(*GP*UP*GP*UP*GP*AP*AP*UP*GP*AP*AP*UP)-3', TAR DNA-BINDING PROTEIN 43 | | Authors: | Lukavsky, P.J, Daujotyte, D, Tollervey, J.R, Ule, J, Stuani, C, Buratti, E, Baralle, F.E, Damberger, F.F, Allain, F.H.T. | | Deposit date: | 2013-06-06 | | Release date: | 2013-11-13 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Molecular Basis of Ug-Rich RNA Recognition by the Human Splicing Factor Tdp-43
Nat.Struct.Mol.Biol., 20, 2013
|
|
6V6H
 
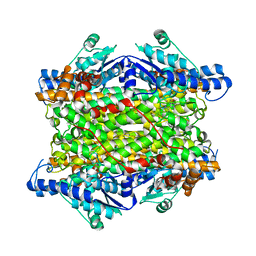 | | Crystal structure of histidine ammonia-lyase from Trypanosoma cruzi | | Descriptor: | Histidine ammonia-lyase | | Authors: | Miranda, R.R, Silva, M, Barison, M.J, Silber, A.M, Iulek, J. | | Deposit date: | 2019-12-05 | | Release date: | 2020-06-10 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Crystal structure of histidine ammonia-lyase from Trypanosoma cruzi.
Biochimie, 175, 2020
|
|
5AJM
 
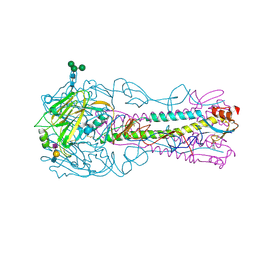 | | H5 (VN1194) Asn186Lys Mutant Haemagglutinin in Complex with Avian Receptor Analogue 3'SLN | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 3[N-MORPHOLINO]PROPANE SULFONIC ACID, ... | | Authors: | Xiong, X, Xiao, H, Martin, S.R, Coombs, P.J, Liu, J, Collins, P.J, Vachieri, S.G, Walker, P.A, Lin, Y.P, McCauley, J.W, Gamblin, S.J, Skehel, J.J. | | Deposit date: | 2015-02-25 | | Release date: | 2015-03-25 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Enhanced Human Receptor Binding by H5 Haemagglutinins.
Virology, 456, 2014
|
|
6NEZ
 
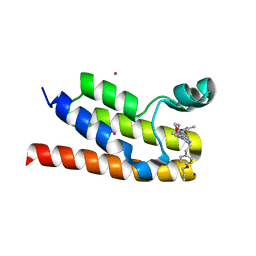 | | Trypanosoma brucei - BDF5, Tb427tmp.01.5000 A, solved with PF-CBP1 | | Descriptor: | 5-(3,5-dimethyl-1,2-oxazol-4-yl)-1-[2-(morpholin-4-yl)ethyl]-2-[2-(4-propoxyphenyl)ethyl]-1H-benzimidazole, UNKNOWN ATOM OR ION, Uncharacterized protein | | Authors: | Lin, Y.H, Dong, A, Tempel, W, McAuley, J, Loppnau, P, Bountra, C, Arrowsmith, C.H, Edwards, A.M, Hui, R, Vedadi, M, Harding, R.J, Structural Genomics Consortium (SGC) | | Deposit date: | 2018-12-18 | | Release date: | 2019-01-23 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Trypanosoma brucei - BDF5, Tb427tmp.01.5000 A, solved with PF-CBP1
to be published
|
|
4IKF
 
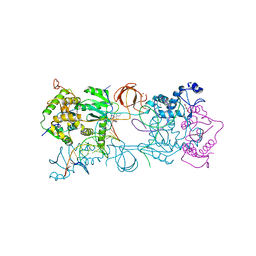 | | PFV intasome with inhibitor MB-76 | | Descriptor: | 5'-D(*AP*TP*TP*GP*TP*CP*AP*TP*GP*GP*AP*AP*TP*TP*TP*CP*GP*CP*A)-3', 5'-D(*TP*GP*CP*GP*AP*AP*AP*TP*TP*CP*CP*AP*TP*GP*AP*CP*A)-3', AMMONIUM ION, ... | | Authors: | Taltynov, O, Demeulemeester, J, Desimmie, B.A, Suchaud, V, Billamboz, M, Lion, C, Bailly, F, Debyser, Z, Cotelle, P, Christ, F, Strelkov, S.V. | | Deposit date: | 2012-12-26 | | Release date: | 2013-04-03 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | 2-Hydroxyisoquinoline-1,3(2H,4H)-diones (HIDs), novel inhibitors of HIV integrase with a high barrier to resistance.
Acs Chem.Biol., 8, 2013
|
|
5JRJ
 
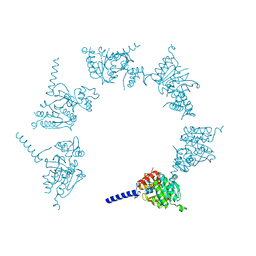 | | Crystal Structure of Herbaspirillum seropedicae RecA | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, ADENOSINE-5'-TRIPHOSPHATE, CALCIUM ION, ... | | Authors: | Leite, W.C, Galvao, C.W, Saab, S.C, Iulek, J, Etto, R.M, Steffens, M.B.R, Chitteni-Pattu, S, Stanage, T, Keck, J.L, Cox, M.M. | | Deposit date: | 2016-05-06 | | Release date: | 2016-08-03 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural and Functional Studies of H. seropedicae RecA Protein - Insights into the Polymerization of RecA Protein as Nucleoprotein Filament.
PLoS ONE, 11, 2016
|
|
3SB2
 
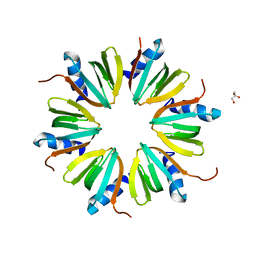 | | Crystal Structure of the RNA chaperone Hfq from Herbaspirillum seropedicae SMR1 | | Descriptor: | GLYCEROL, Protein hfq | | Authors: | Kadowaki, M.A.S, Iulek, J, Barbosa, J.A.R.G, Pedrosa, F.O, Souza, E.M, Chubatsu, L.S, Monteiro, R.A, Steffens, M.B.R. | | Deposit date: | 2011-06-03 | | Release date: | 2012-01-04 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.6301 Å) | | Cite: | Structural characterization of the RNA chaperone Hfq from the nitrogen-fixing bacterium Herbaspirillum seropedicae SmR1.
Biochim.Biophys.Acta, 1824, 2011
|
|
5JAE
 
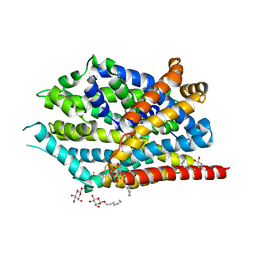 | | LeuT in the outward-oriented, Na+-free return state, P21 form at pH 6.5 | | Descriptor: | Transporter, octyl beta-D-glucopyranoside | | Authors: | Malinauskaite, L, Sahin, C, Said, S, Grouleff, J, Shahsavar, A, Bjerregaard, H, Noer, P, Severinsen, K, Boesen, T, Schiott, B, Sinning, S, Nissen, P. | | Deposit date: | 2016-04-12 | | Release date: | 2016-06-01 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | A conserved leucine occupies the empty substrate site of LeuT in the Na(+)-free return state.
Nat Commun, 7, 2016
|
|
5KSR
 
 | | Stationary phase survival protein E (SurE) from Xylella fastidiosa - XFSurE-TB (Tetramer Bigger). | | Descriptor: | 5'-nucleotidase SurE, CHLORIDE ION, IODIDE ION, ... | | Authors: | Machado, A.T.P, Fonseca, E.M.B, Dos Reis, M.A, Saraiva, A.M, Dos Santos, C.A, De Toledo, M.A, Polikarpov, I, De Souza, A.P, De Aparicio, R, Iulek, J. | | Deposit date: | 2016-07-09 | | Release date: | 2017-07-19 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.96 Å) | | Cite: | Conformational variability of the stationary phase survival protein E from Xylella fastidiosa revealed by X-ray crystallography, small-angle X-ray scattering studies, and normal mode analysis.
Proteins, 85, 2017
|
|
5KSQ
 
 | | Stationary phase survival protein E (SurE) from Xylella fastidiosa | | Descriptor: | 5'-nucleotidase SurE, IODIDE ION, MANGANESE (II) ION, ... | | Authors: | Machado, A.T.P, Fonseca, E.M.B, Dos Reis, M.A, Saraiva, A.M, Dos Santos, C.A, De Toledo, M.A, Polikarpov, I, De Souza, A.P, Aparicio, R, Iulek, J. | | Deposit date: | 2016-07-09 | | Release date: | 2017-07-19 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | Conformational variability of the stationary phase survival protein E from Xylella fastidiosa revealed by X-ray crystallography, small-angle X-ray scattering studies, and normal mode analysis.
Proteins, 85, 2017
|
|
5KSS
 
 | | Stationary phase survival protein E (SurE) from Xylella fastidiosa - XFSurE-Ds (Dimer Smaller) | | Descriptor: | 5'-nucleotidase SurE, CHLORIDE ION, IODIDE ION, ... | | Authors: | Machado, A.T.P, Fonseca, E.M.B, Dos Reis, M.A, Saraiva, A.M, Dos Santos, C.A, De Toledo, A.M.S, Polikarpov, I, De Souza, A.P, Aparicio, R, Iulek, J. | | Deposit date: | 2016-07-09 | | Release date: | 2017-07-19 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.82 Å) | | Cite: | Conformational variability of the stationary phase survival protein E from Xylella fastidiosa revealed by X-ray crystallography, small-angle X-ray scattering studies, and normal mode analysis.
Proteins, 85, 2017
|
|
5KST
 
 | | Stationary phase Survival protein E (SurE) from Xylella fastidiosa- XfSurE-TSAmp (Tetramer Smaller - crystallization with 3'AMP). | | Descriptor: | 5'-nucleotidase SurE, IODIDE ION, MANGANESE (II) ION, ... | | Authors: | Machado, A.T.P, Fonseca, E.M.B, Dos Reis, M.A, Saraiva, A.M, Dos Santos, C.A, De Toledo, M.A.S, Polikarpov, I, De Souza, A.P, Aparicio, R, Iulek, J. | | Deposit date: | 2016-07-09 | | Release date: | 2017-07-19 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.759 Å) | | Cite: | Conformational variability of the stationary phase survival protein E from Xylella fastidiosa revealed by X-ray crystallography, small-angle X-ray scattering studies, and normal mode analysis.
Proteins, 85, 2017
|
|
1ZU2
 
 | |
