4YNC
 
 | | OYE1 W116A COMPLEXED WITH (Z)-METHYL-3-CYANO-3-PHENYLACRYLATE IN A NON PRODUCTIVE BINDING MODE | | Descriptor: | CHLORIDE ION, FLAVIN MONONUCLEOTIDE, NADPH dehydrogenase 1, ... | | Authors: | Santangelo, S, Brenna, E, Stewart, J.D, Powell III, R.W. | | Deposit date: | 2015-03-09 | | Release date: | 2016-01-20 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.498 Å) | | Cite: | Opposite Enantioselectivity in the Bioreduction of (Z)-beta-Aryl-beta-cyanoacrylates Mediated by the Tryptophan 116 Mutants of Old Yellow Enzyme 1: Synthetic Approach to (R)- and (S)-beta-Aryl-gamma-lactams
Adv.Synth.Catal., 357, 2015
|
|
2VTX
 
 | | ACTIVATION OF NUCLEOPLASMIN, AN OLIGOMERIC HISTONE CHAPERONE, CHALLENGES ITS STABILITY | | Descriptor: | NPM-A PROTEIN | | Authors: | Taneva, S.G, Munoz, I.G, Franco, G, Falces, J, Arregi, I, Muga, A, Montoya, G, Urbaneja, M.A, Banuelos, S. | | Deposit date: | 2008-05-16 | | Release date: | 2008-12-16 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Activation of Nucleoplasmin, an Oligomeric Histone Chaperone, Challenges its Stability.
Biochemistry, 47, 2008
|
|
7JST
 
 | | Crystal structure of SARS-CoV-2 3CL in apo form | | Descriptor: | 3C-like proteinase, PHOSPHATE ION | | Authors: | Iketani, S, Forouhar, F, Liu, H, Hong, S.J, Lin, F.-Y, Nair, M.S, Zask, A, Huang, Y, Xing, L, Stockwell, B.R, Chavez, A, Ho, D.D. | | Deposit date: | 2020-08-16 | | Release date: | 2021-03-03 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Lead compounds for the development of SARS-CoV-2 3CL protease inhibitors.
Nat Commun, 12, 2021
|
|
7JT0
 
 | | Crystal structure of SARS-CoV-2 3CL protease in complex with MAC5576 | | Descriptor: | 3C-like proteinase, PHOSPHATE ION, thiophene-2-carbaldehyde | | Authors: | Iketani, S, Forouhar, F, Liu, H, Hong, S.J, Lin, F.-Y, Nair, M.S, Zask, A, Huang, Y, Xing, L, Stockwell, B.R, Chavez, A, Ho, D.D. | | Deposit date: | 2020-08-16 | | Release date: | 2021-03-10 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.73 Å) | | Cite: | Lead compounds for the development of SARS-CoV-2 3CL protease inhibitors.
Nat Commun, 12, 2021
|
|
7JW8
 
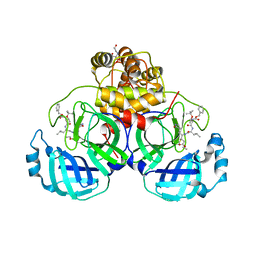 | | Crystal structure of SARS-CoV-2 3CL protease in complex with compound 4 in space group P1 | | Descriptor: | 2-[BIS-(2-HYDROXY-ETHYL)-AMINO]-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 3C-like proteinase, ethyl (4R)-4-[[(2S)-4-methyl-2-[[(2S,3R)-3-[(2-methylpropan-2-yl)oxy]-2-(phenylmethoxycarbonylamino)butanoyl]amino]pentanoyl]amino]-5-[(3S)-2-oxidanylidenepyrrolidin-3-yl]pentanoate | | Authors: | Iketani, S, Forouhar, F, Liu, H, Hong, S.J, Lin, F.-Y, Nair, M.S, Zask, A, Xing, L, Stockwell, B.R, Chavez, A, Ho, D.D. | | Deposit date: | 2020-08-25 | | Release date: | 2021-03-10 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Lead compounds for the development of SARS-CoV-2 3CL protease inhibitors.
Nat Commun, 12, 2021
|
|
7JSU
 
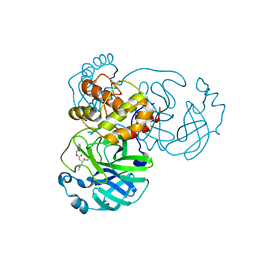 | | Crystal structure of SARS-CoV-2 3CL protease in complex with GC376 | | Descriptor: | 3C-like proteinase, N~2~-[(benzyloxy)carbonyl]-N-{(2S)-1-hydroxy-3-[(3S)-2-oxopyrrolidin-3-yl]propan-2-yl}-L-leucinamide, PHOSPHATE ION | | Authors: | Iketani, S, Forouhar, F, Liu, H, Hong, S.J, Lin, F.-Y, Nair, M.S, Zask, A, Xing, L, Stockwell, B.R, Chavez, A, Ho, D.D. | | Deposit date: | 2020-08-16 | | Release date: | 2021-03-10 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Lead compounds for the development of SARS-CoV-2 3CL protease inhibitors.
Nat Commun, 12, 2021
|
|
7JT7
 
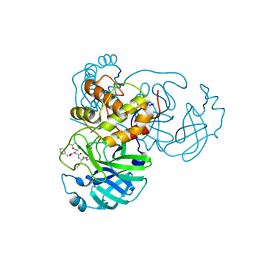 | | Crystal structure of SARS-CoV-2 3CL protease in complex with compound 4 | | Descriptor: | 3C-like proteinase, ethyl (4R)-4-[[(2S)-4-methyl-2-[[(2S,3R)-3-[(2-methylpropan-2-yl)oxy]-2-(phenylmethoxycarbonylamino)butanoyl]amino]pentanoyl]amino]-5-[(3S)-2-oxidanylidenepyrrolidin-3-yl]pentanoate | | Authors: | Iketani, S, Forouhar, F, Liu, H, Hong, S.J, Lin, F.-Y, Nair, M.S, Zask, A, Huang, Y, Xing, L, Stockwell, B.R, Chavez, A, Ho, D.D. | | Deposit date: | 2020-08-17 | | Release date: | 2021-03-17 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.94 Å) | | Cite: | Lead compounds for the development of SARS-CoV-2 3CL protease inhibitors.
Nat Commun, 12, 2021
|
|
2W27
 
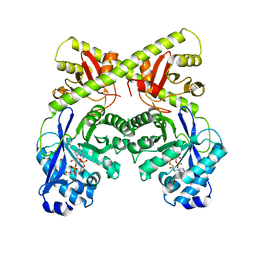 | | CRYSTAL STRUCTURE OF THE BACILLUS SUBTILIS YKUI PROTEIN, WITH AN EAL DOMAIN, IN COMPLEX WITH SUBSTRATE C-DI-GMP AND CALCIUM | | Descriptor: | CALCIUM ION, GUANOSINE-5'-MONOPHOSPHATE, YKUI PROTEIN | | Authors: | Padavattan, S, AndERSON, W.F, Schirmer, T. | | Deposit date: | 2008-10-24 | | Release date: | 2009-02-24 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal Structures of Ykui and its Complex with Second Messenger C-Di-Gmp Suggests Catalytic Mechanism of Phosphodiester Bond Cleavage by Eal Domains.
J.Biol.Chem., 284, 2009
|
|
2E1P
 
 | | Crystal structure of pro-Tk-subtilisin | | Descriptor: | CALCIUM ION, Tk-subtilisin | | Authors: | Tanaka, S, Saito, K, Chon, H, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2006-10-27 | | Release date: | 2007-01-16 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of unautoprocessed precursor of subtilisin from a hyperthermophilic archaeon: evidence for Ca2+-induced folding
J.Biol.Chem., 282, 2007
|
|
7KH8
 
 | | Human LMPTP in complex with inhibitor | | Descriptor: | 3-[(2,6-dichlorophenyl)methyl]-8-(2-methylphenyl)-3H-purin-6-amine, Low molecular weight phosphotyrosine protein phosphatase, NITRATE ION | | Authors: | Stanford, S.M, Diaz, M.A, Ardecky, R.J, Zou, J, Roosild, T, Holmes, Z.J, Hedrick, M.P, Rodiles, S, Santelli, E, Chung, T.D.Y, Jackson, M.R, Bottini, N, Pinkerton, A.B. | | Deposit date: | 2020-10-20 | | Release date: | 2021-05-19 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Discovery of Orally Bioavailable Purine-Based Inhibitors of the Low-Molecular-Weight Protein Tyrosine Phosphatase.
J.Med.Chem., 64, 2021
|
|
2D2E
 
 | | Crystal structure of atypical cytoplasmic ABC-ATPase SufC from Thermus thermophilus HB8 | | Descriptor: | CHLORIDE ION, GLYCEROL, SufC protein | | Authors: | Watanabe, S, Kita, A, Miki, K, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-09-08 | | Release date: | 2005-10-25 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal Structure of Atypical Cytoplasmic ABC-ATPase SufC from Thermus thermophilus HB8.
J.Mol.Biol., 353, 2005
|
|
2VZN
 
 | |
5GU7
 
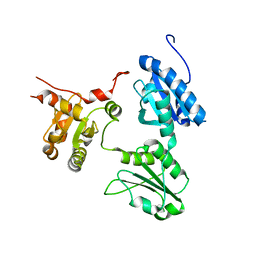 | | Crystal Structure of human ERp44 form II | | Descriptor: | Endoplasmic reticulum resident protein 44 | | Authors: | Watanabe, S, Inaba, K. | | Deposit date: | 2016-08-26 | | Release date: | 2017-04-12 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structural basis of pH-dependent client binding by ERp44, a key regulator of protein secretion at the ER-Golgi interface
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
5GU6
 
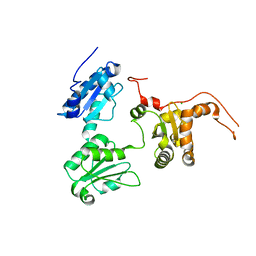 | | Crystal structure of Human ERp44 form I | | Descriptor: | CHLORIDE ION, Endoplasmic reticulum resident protein 44 | | Authors: | Watanabe, S, Inaba, K. | | Deposit date: | 2016-08-26 | | Release date: | 2017-04-12 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural basis of pH-dependent client binding by ERp44, a key regulator of protein secretion at the ER-Golgi interface
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
3AHP
 
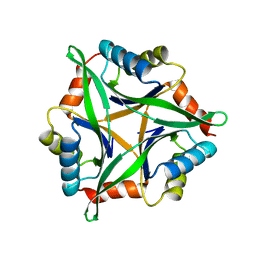 | | Crystal structure of stable protein, CutA1, from a psychrotrophic bacterium Shewanella sp. SIB1 | | Descriptor: | CutA1 | | Authors: | Tanaka, S, Angkawidjaja, C, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2010-04-26 | | Release date: | 2011-01-05 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of stable protein CutA1 from psychrotrophic bacterium Shewanella sp. SIB1
J.SYNCHROTRON RADIAT., 18, 2011
|
|
2D2F
 
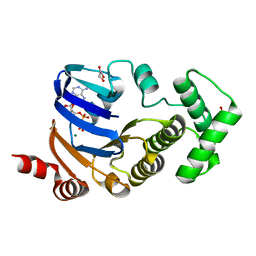 | | Crystal structure of atypical cytoplasmic ABC-ATPase SufC from Thermus thermophilus HB8 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Watanabe, S, Kita, A, Miki, K, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-09-08 | | Release date: | 2005-10-25 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Structure of Atypical Cytoplasmic ABC-ATPase SufC from Thermus thermophilus HB8.
J.Mol.Biol., 353, 2005
|
|
5AUO
 
 | | Crystal structure of the HypAB-Ni complex (AMPPCP) | | Descriptor: | ATPase involved in chromosome partitioning, ParA/MinD family, Mrp homolog, ... | | Authors: | Watanabe, S, Kawashima, T, Nishitani, Y, Miki, K. | | Deposit date: | 2015-05-27 | | Release date: | 2015-06-24 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis of a Ni acquisition cycle for [NiFe] hydrogenase by Ni-metallochaperone HypA and its enhancer
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
3A3O
 
 | | Crystal structure of complex between SA-subtilisin and Tk-propeptide with deletion of the five C-terminal residues | | Descriptor: | CALCIUM ION, Tk-subtilisin, ZINC ION | | Authors: | Tanaka, S, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2009-06-15 | | Release date: | 2009-08-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Identification of the interactions critical for propeptide-catalyzed folding of Tk-subtilisin
J.Mol.Biol., 394, 2009
|
|
3A43
 
 | | Crystal structure of HypA | | Descriptor: | Hydrogenase nickel incorporation protein hypA, ZINC ION | | Authors: | Watanabe, S, Arai, T, Matsumi, R, Aromi, H, Imanaka, T, Miki, K. | | Deposit date: | 2009-06-30 | | Release date: | 2009-10-06 | | Last modified: | 2016-12-21 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of HypA, a nickel-binding metallochaperone for [NiFe] hydrogenase maturation.
J.Mol.Biol., 394, 2009
|
|
3A3N
 
 | | Crystal structure of complex between SA-subtilisin and Tk-propeptide with deletion of the two C-terminal residues | | Descriptor: | CALCIUM ION, Tk-subtilisin, ZINC ION | | Authors: | Tanaka, S, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2009-06-15 | | Release date: | 2009-08-18 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Identification of the interactions critical for propeptide-catalyzed folding of Tk-subtilisin
J.Mol.Biol., 394, 2009
|
|
5FD9
 
 | | X-ray Crystal Structure of ESCRT-III Snf7 core domain (conformation B) | | Descriptor: | Vacuolar-sorting protein SNF7 | | Authors: | Tang, S, Henne, W.M, Borbat, P.P, Buchkovich, N.J, Freed, J.H, Mao, Y, Fromme, J.C, Emr, S.D. | | Deposit date: | 2015-12-15 | | Release date: | 2015-12-30 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural basis for activation, assembly and membrane binding of ESCRT-III Snf7 filaments.
Elife, 4, 2015
|
|
2Z2Y
 | | Crystal structure of autoprocessed form of Tk-subtilisin | | Descriptor: | CALCIUM ION, Tk-subtilisin, ZINC ION | | Authors: | Tanaka, S, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2007-05-29 | | Release date: | 2007-12-04 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Four new crystal structures of Tk-subtilisin in unautoprocessed, autoprocessed and mature forms: insight into structural changes during maturation
J.Mol.Biol., 372, 2007
|
|
2ZRQ
 
 | | Crystal structure of S324A-subtilisin | | Descriptor: | CALCIUM ION, Tk-subtilisin | | Authors: | Tanaka, S, Takeuchi, Y, Matsumura, H, Koga, Y, Takano, K, Kanaya, S. | | Deposit date: | 2008-08-28 | | Release date: | 2009-03-03 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Crystal structure of Tk-subtilisin folded without propeptide: requirement of propeptide for acceleration of folding
Febs Lett., 582, 2008
|
|
3DNC
 
 | | Carboxysome shell protein, CcmK2 C-terminal deletion mutant, with a closer spacing between hexamers | | Descriptor: | Carbon dioxide-concentrating mechanism protein ccmK homolog 2, GLYCEROL, SULFATE ION | | Authors: | Tanaka, S, Sawaya, M.R, Yeates, T.O. | | Deposit date: | 2008-07-01 | | Release date: | 2009-01-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Insights from multiple structures of the shell proteins from the beta-carboxysome.
Protein Sci., 18, 2009
|
|
3MN8
 
 | | Structure of Drosophila melanogaster carboxypeptidase D isoform 1B short | | Descriptor: | (2-GUANIDINOETHYLMERCAPTO)SUCCINIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, ... | | Authors: | Tanco, S, Arolas, J.L, Guevara, T, Lorenzo, J, Aviles, F.X, Gomis-Ruth, F.X. | | Deposit date: | 2010-04-21 | | Release date: | 2010-07-28 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure-Function Analysis of the Short Splicing Variant Carboxypeptidase Encoded by Drosophila melanogaster silver.
J.Mol.Biol., 401, 2010
|
|
