5MIS
 | | Crystal Structure of Lactococcus lactis Thioredoxin Reductase Exposed to Visible Light (180 min) | | Descriptor: | DI(HYDROXYETHYL)ETHER, FLAVIN-ADENINE DINUCLEOTIDE, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Skjoldager, N, Bang, M.B, Svensson, B, Hagglund, P, Harris, P. | | Deposit date: | 2016-11-29 | | Release date: | 2017-04-12 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | The structure of Lactococcus lactis thioredoxin reductase reveals molecular features of photo-oxidative damage.
Sci Rep, 7, 2017
|
|
5MIT
 | | Crystal Structure of Lactococcus lactis Thioredoxin Reductase Exposed to Visible Light (240 min) | | Descriptor: | DI(HYDROXYETHYL)ETHER, FLAVIN-ADENINE DINUCLEOTIDE, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Skjoldager, N, Bang, M.B, Svensson, B, Hagglund, P, Harris, P. | | Deposit date: | 2016-11-29 | | Release date: | 2017-04-12 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The structure of Lactococcus lactis thioredoxin reductase reveals molecular features of photo-oxidative damage.
Sci Rep, 7, 2017
|
|
5MIP
 | | Crystal Structure of Lactococcus lactis Thioredoxin Reductase Exposed to Visible Light (30 min) | | Descriptor: | DI(HYDROXYETHYL)ETHER, FLAVIN-ADENINE DINUCLEOTIDE, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Skjoldager, N, Bang, M.B, Svensson, B, Hagglund, P, Harris, P. | | Deposit date: | 2016-11-29 | | Release date: | 2017-04-12 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The structure of Lactococcus lactis thioredoxin reductase reveals molecular features of photo-oxidative damage.
Sci Rep, 7, 2017
|
|
5MIQ
 
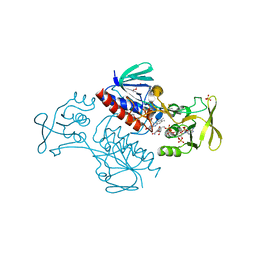 | | Crystal Structure of Lactococcus lactis Thioredoxin Reductase Exposed to Visible Light (60 min) | | Descriptor: | DI(HYDROXYETHYL)ETHER, FLAVIN-ADENINE DINUCLEOTIDE, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Skjoldager, N, Bang, M.B, Svensson, B, Hagglund, P, Harris, P. | | Deposit date: | 2016-11-29 | | Release date: | 2017-04-12 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | The structure of Lactococcus lactis thioredoxin reductase reveals molecular features of photo-oxidative damage.
Sci Rep, 7, 2017
|
|
5MJK
 
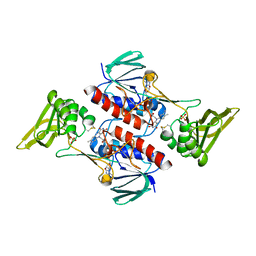 | | Crystal Structure of Lactococcus lactis Thioredoxin Reductase (FO conformation) | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, SULFATE ION, Thioredoxin reductase | | Authors: | Skjoldager, N, Bang, M.B, Svensson, B, Hagglund, P, Harris, P. | | Deposit date: | 2016-12-01 | | Release date: | 2017-04-12 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The structure of Lactococcus lactis thioredoxin reductase reveals molecular features of photo-oxidative damage.
Sci Rep, 7, 2017
|
|
5MH4
 
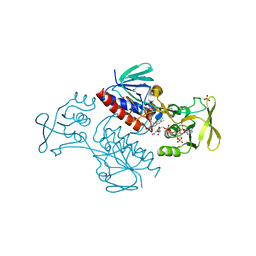 | | Crystal Structure of Lactococcus lactis Thioredoxin Reductase (FR conformation) | | Descriptor: | DI(HYDROXYETHYL)ETHER, FLAVIN-ADENINE DINUCLEOTIDE, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Skjoldager, N, Bang, M.B, Svensson, B, Hagglund, P, Harris, P. | | Deposit date: | 2016-11-23 | | Release date: | 2017-04-12 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | The structure of Lactococcus lactis thioredoxin reductase reveals molecular features of photo-oxidative damage.
Sci Rep, 7, 2017
|
|
1AVA
 
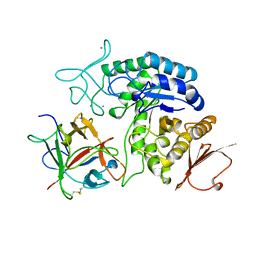 | | AMY2/BASI PROTEIN-PROTEIN COMPLEX FROM BARLEY SEED | | Descriptor: | BARLEY ALPHA-AMYLASE 2(CV MENUET), BARLEY ALPHA-AMYLASE/SUBTILISIN INHIBITOR, CALCIUM ION | | Authors: | Vallee, F, Kadziola, A, Bourne, Y, Juy, M, Svensson, B, Haser, R. | | Deposit date: | 1997-09-15 | | Release date: | 1999-03-16 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Barley alpha-amylase bound to its endogenous protein inhibitor BASI: crystal structure of the complex at 1.9 A resolution.
Structure, 6, 1998
|
|
2WV8
 
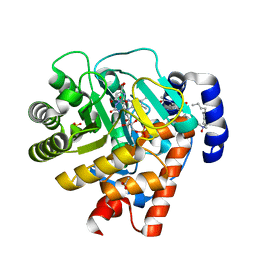 | | Complex of human dihydroorotate dehydrogenase with the inhibitor 221290 | | Descriptor: | 2-ACETAMIDO-5-(4-PHENYLPHENYL)BENZOIC ACID, ACETATE ION, DECYLAMINE-N,N-DIMETHYL-N-OXIDE, ... | | Authors: | Walse, B, Svensson, B, Fritzson, I, Dahlberg, L, Wellmar, U, Al-Karadaghi, S. | | Deposit date: | 2009-10-15 | | Release date: | 2010-08-25 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Inhibition of Human Dhodh by 4-Hydroxycoumarins, Fenamic Acids, and N-(Alkylcarbonyl)Anthranilic Acids Identified by Structure-Guided Fragment Selection.
Chemmedchem, 5, 2010
|
|
2Y5E
 
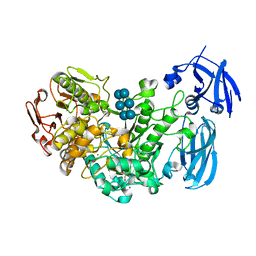 | | BARLEY LIMIT DEXTRINASE IN COMPLEX WITH ALPHA-CYCLODEXTRIN | | Descriptor: | CALCIUM ION, Cyclohexakis-(1-4)-(alpha-D-glucopyranose), GLYCEROL, ... | | Authors: | Vester-Christensen, M.B, Hachem, M.A, Svensson, B, Henriksen, A. | | Deposit date: | 2011-01-13 | | Release date: | 2011-01-26 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | Crystal Structure of an Essential Enzyme in Seed Starch Degradation: Barley Limit Dextrinase in Complex with Cyclodextrins.
J.Mol.Biol., 403, 2010
|
|
2Y4S
 
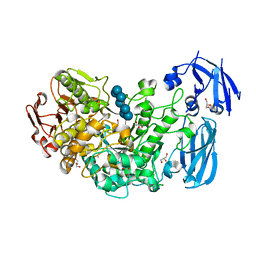 | | BARLEY LIMIT DEXTRINASE IN COMPLEX WITH BETA-CYCLODEXTRIN | | Descriptor: | CALCIUM ION, Cycloheptakis-(1-4)-(alpha-D-glucopyranose), GLYCEROL, ... | | Authors: | Vester-Christensen, M.B, Hachem, M.A, Svensson, B, Henriksen, A. | | Deposit date: | 2011-01-10 | | Release date: | 2011-01-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal Structure of an Essential Enzyme in Seed Starch Degradation: Barley Limit Dextrinase in Complex with Cyclodextrins.
J.Mol.Biol., 403, 2010
|
|
2VLT
 
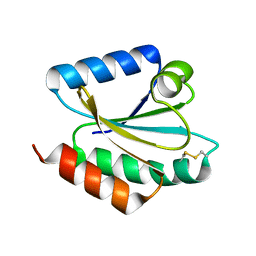 | | Crystal structure of barley thioredoxin h isoform 2 in the oxidized state | | Descriptor: | THIOREDOXIN H ISOFORM 2. | | Authors: | Maeda, K, Hagglund, P, Finnie, C, Svensson, B, Henriksen, A. | | Deposit date: | 2008-01-16 | | Release date: | 2008-04-29 | | Last modified: | 2017-07-12 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal Structures of Barley Thioredoxin H Isoforms Hvtrxh1 and Hvtrxh2 Reveal Features Involved in Protein Recognition and Possibly in Discriminating the Isoform Specificity.
Protein Sci., 17, 2008
|
|
2WHD
 | | Barley NADPH-dependent thioredoxin reductase 2 | | Descriptor: | CITRATE ANION, FLAVIN-ADENINE DINUCLEOTIDE, THIOREDOXIN REDUCTASE | | Authors: | Kirkensgaard, K.G, Hagglund, P, Finnie, C, Svensson, B, Henriksen, A. | | Deposit date: | 2009-05-04 | | Release date: | 2009-09-01 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of Hordeum Vulgare Nadph-Dependent Thioredoxin Reductase 2. Unwinding the Reaction Mechanism.
Acta Crystallogr.,Sect.D, 65, 2009
|
|
2VM1
 
 | | Crystal structure of barley thioredoxin h isoform 1 crystallized using ammonium sulfate as precipitant | | Descriptor: | SULFATE ION, THIOREDOXIN H ISOFORM 1. | | Authors: | Maeda, K, Hagglund, P, Finnie, C, Svensson, B, Henriksen, A. | | Deposit date: | 2008-01-21 | | Release date: | 2008-04-29 | | Last modified: | 2017-07-12 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal Structures of Barley Thioredoxin H Isoforms Hvtrxh1 and Hvtrxh2 Reveal Features Involved in Protein Recognition and Possibly in Discriminating the Isoform Specificity.
Protein Sci., 17, 2008
|
|
2VLU
 
 | | Crystal structure of barley thioredoxin h isoform 2 in partially radiation-reduced state | | Descriptor: | THIOREDOXIN H ISOFORM 2. | | Authors: | Maeda, K, Hagglund, P, Finnie, C, Svensson, B, Henriksen, A. | | Deposit date: | 2008-01-16 | | Release date: | 2008-04-29 | | Last modified: | 2017-07-12 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal Structures of Barley Thioredoxin H Isoforms Hvtrxh1 and Hvtrxh2 Reveal Features Involved in Protein Recognition and Possibly in Discriminating the Isoform Specificity.
Protein Sci., 17, 2008
|
|
2XN2
 
 | | Structure of alpha-galactosidase from Lactobacillus acidophilus NCFM with galactose | | Descriptor: | ALPHA-GALACTOSIDASE, GLYCEROL, IMIDAZOLE, ... | | Authors: | Fredslund, F, Abou Hachem, M, Larsen, R.J, Sorensen, P.G, Lo Leggio, L, Svensson, B. | | Deposit date: | 2010-07-30 | | Release date: | 2011-08-10 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Crystal Structure of Alpha-Galactosidase from Lactobacillus Acidophilus Ncfm: Insight Into Tetramer Formation and Substrate Binding.
J.Mol.Biol., 412, 2011
|
|
2VLV
 
 | | Crystal structure of barley thioredoxin h isoform 2 in partially radiation-reduced state | | Descriptor: | THIOREDOXIN H ISOFORM 2. | | Authors: | Maeda, K, Hagglund, P, Finnie, C, Svensson, B, Henriksen, A. | | Deposit date: | 2008-01-16 | | Release date: | 2008-04-29 | | Last modified: | 2017-07-12 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal Structures of Barley Thioredoxin H Isoforms Hvtrxh1 and Hvtrxh2 Reveal Features Involved in Protein Recognition and Possibly in Discriminating the Isoform Specificity.
Protein Sci., 17, 2008
|
|
2XN1
 
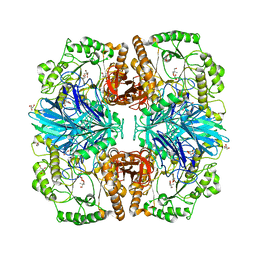 | | Structure of alpha-galactosidase from Lactobacillus acidophilus NCFM with TRIS | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, ALPHA-GALACTOSIDASE, GLYCEROL | | Authors: | Fredslund, F, Abou Hachem, M, Larsen, R.J, Sorensen, P.G, Lo Leggio, L, Svensson, B. | | Deposit date: | 2010-07-30 | | Release date: | 2011-08-10 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal Structure of Alpha-Galactosidase from Lactobacillus Acidophilus Ncfm: Insight Into Tetramer Formation and Substrate Binding.
J.Mol.Biol., 412, 2011
|
|
2VM2
 
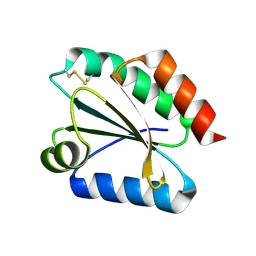 | | Crystal structure of barley thioredoxin h isoform 1 crystallized using PEG as precipitant | | Descriptor: | THIOREDOXIN H ISOFORM 1. | | Authors: | Maeda, K, Hagglund, P, Finnie, C, Svensson, B, Henriksen, A. | | Deposit date: | 2008-01-21 | | Release date: | 2008-04-29 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal Structures of Barley Thioredoxin H Isoforms Hvtrxh1 and Hvtrxh2 Reveal Features Involved in Protein Recognition and Possibly in Discriminating the Isoform Specificity.
Protein Sci., 17, 2008
|
|
2XN0
 
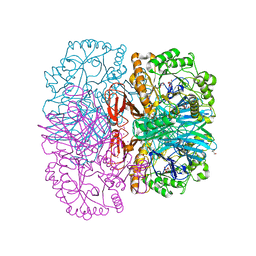 | | Structure of alpha-galactosidase from Lactobacillus acidophilus NCFM, PtCl4 derivative | | Descriptor: | ALPHA-GALACTOSIDASE, GLYCEROL, PLATINUM (II) ION | | Authors: | Fredslund, F, Abou Hachem, M, Larsen, R.J, Sorensen, P.G, Lo Leggio, L, Svensson, B. | | Deposit date: | 2010-07-30 | | Release date: | 2011-08-10 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal Structure of Alpha-Galactosidase from Lactobacillus Acidophilus Ncfm: Insight Into Tetramer Formation and Substrate Binding.
J.Mol.Biol., 412, 2011
|
|
