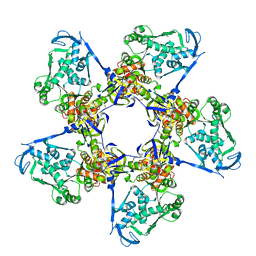
3EZK
 
 | |
3C6H
 
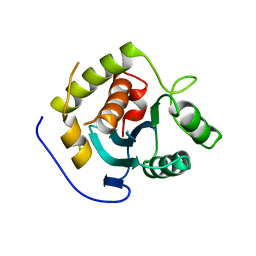 | | Crystal Structure of the RB49 gp17 nuclease domain | | Descriptor: | MAGNESIUM ION, Terminase large subunit | | Authors: | Sun, S, Rossmann, M.G. | | Deposit date: | 2008-02-04 | | Release date: | 2009-01-13 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | The structure of the phage T4 DNA packaging motor suggests a mechanism dependent on electrostatic forces.
Cell(Cambridge,Mass.), 135, 2008
|
|
3C6A
 
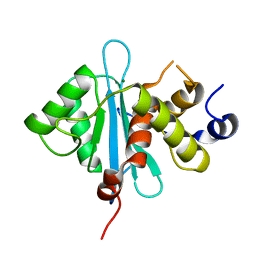 | | Crystal Structure of the RB49 gp17 nuclease domain | | Descriptor: | MAGNESIUM ION, Terminase large subunit | | Authors: | Sun, S, Rossmann, M.G. | | Deposit date: | 2008-02-04 | | Release date: | 2009-01-13 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.16 Å) | | Cite: | The structure of the phage T4 DNA packaging motor suggests a mechanism dependent on electrostatic forces.
Cell(Cambridge,Mass.), 135, 2008
|
|
3CPE
 
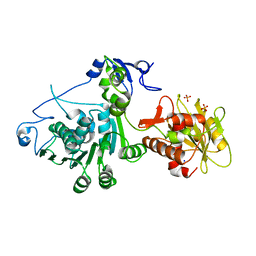 | | Crystal Structure of T4 gp17 | | Descriptor: | DNA packaging protein Gp17, PHOSPHATE ION, SODIUM ION | | Authors: | Sun, S, Rossmann, M.G. | | Deposit date: | 2008-03-31 | | Release date: | 2009-01-13 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | The structure of the phage T4 DNA packaging motor suggests a mechanism dependent on electrostatic forces
Cell(Cambridge,Mass.), 135, 2008
|
|
2A1K
 
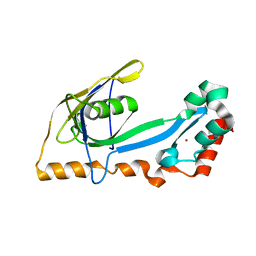 | | RB69 single-stranded DNA binding protein core domain | | Descriptor: | ZINC ION, gp32 single stranded DNA binding protein | | Authors: | Sun, S, Geng, L, Shamoo, Y. | | Deposit date: | 2005-06-20 | | Release date: | 2006-05-09 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure and enzymatic properties of a chimeric bacteriophage RB69 DNA polymerase and single-stranded DNA binding protein with increased processivity.
Proteins, 65, 2006
|
|
3J2X
 
 | |
3TXS
 
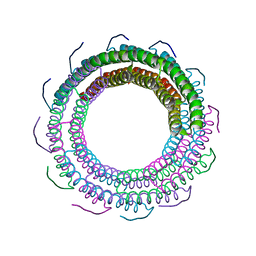 | | Crystal Structure of phage 44RR small terminase gp16 | | Descriptor: | Terminase DNA packaging enzyme small subunit | | Authors: | Sun, S, Xiang, Y, Rossmann, M.G. | | Deposit date: | 2011-09-23 | | Release date: | 2011-12-28 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Structure and function of the small terminase component of the DNA packaging machine in T4-like bacteriophages.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
3J2W
 
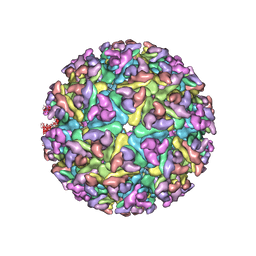 | | Electron cryo-microscopy of Chikungunya virus | | Descriptor: | Capsid protein, Glycoprotein E1, Glycoprotein E2 | | Authors: | Sun, S, Xiang, Y, Rossmann, M.G. | | Deposit date: | 2013-01-28 | | Release date: | 2013-04-24 | | Last modified: | 2018-07-18 | | Method: | ELECTRON MICROSCOPY (5 Å) | | Cite: | Structural analyses at pseudo atomic resolution of Chikungunya virus and antibodies show mechanisms of neutralization.
Elife, 2, 2013
|
|
3J30
 
 | |
3KK5
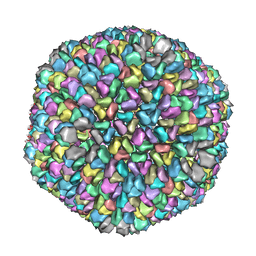 
 | |
3J2Y
 
 | |
3J2Z
 
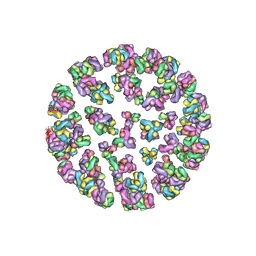 | |
3TXQ
 
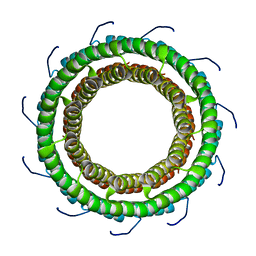 | |
4GQ9
 
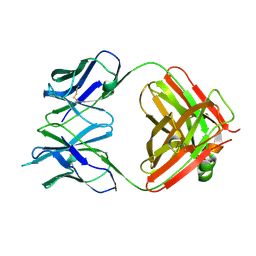 | | Chikungunya virus neutralizing antibody 9.8B Fab fragment | | Descriptor: | Chikungunya virus neutralizing antibody 9.8B Fab fragment heavy chain, Chikungunya virus neutralizing antibody 9.8B Fab fragment light chain | | Authors: | Sun, S, Rossmann, M.G. | | Deposit date: | 2012-08-22 | | Release date: | 2013-07-10 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.995 Å) | | Cite: | Structural analyses at pseudo atomic resolution of Chikungunya virus and antibodies show mechanisms of neutralization.
Elife, 2, 2013
|
|
5XMI
 
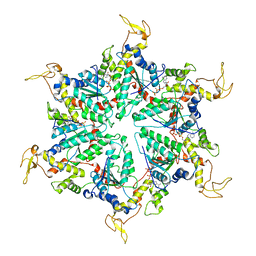 | | Cryo-EM Structure of the ATP-bound VPS4 mutant-E233Q hexamer (masked) | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Vacuolar protein sorting-associated protein 4 | | Authors: | Sun, S, Li, L, Yang, F, Wang, X, Fan, F, Li, X, Wang, H, Sui, S. | | Deposit date: | 2017-05-15 | | Release date: | 2017-08-09 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Cryo-EM structures of the ATP-bound Vps4(E233Q) hexamer and its complex with Vta1 at near-atomic resolution
Nat Commun, 8, 2017
|
|
5XMK
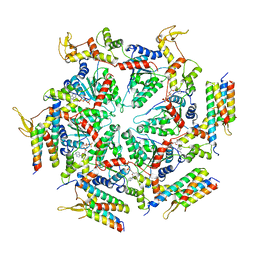 
 | | Cryo-EM structure of the ATP-bound Vps4 mutant-E233Q complex with Vta1 (masked) | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Vacuolar protein sorting-associated protein 4, Vacuolar protein sorting-associated protein VTA1 | | Authors: | Sun, S, Li, L, Yang, F, Wang, X, Fan, F, Li, X, Wang, H, Sui, S. | | Deposit date: | 2017-05-15 | | Release date: | 2017-08-09 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (4.18 Å) | | Cite: | Cryo-EM structures of the ATP-bound Vps4(E233Q) hexamer and its complex with Vta1 at near-atomic resolution
Nat Commun, 8, 2017
|
|
2O0K
 
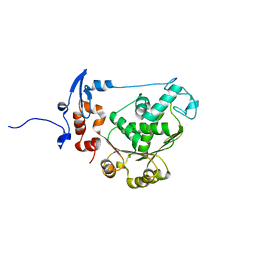 | | T4 gp17 ATPase domain mutant | | Descriptor: | DNA packaging protein Gp17 | | Authors: | Sun, S, Rossmann, M.G. | | Deposit date: | 2006-11-27 | | Release date: | 2007-04-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | The Structure of the ATPase that Powers DNA Packaging into Bacteriophage T4 Procapsids
MOL.CELL, 25, 2007
|
|
2O0H
 
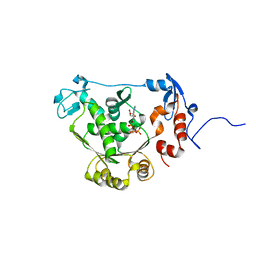 | | T4 gp17 ATPase domain mutant complexed with ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, DNA packaging protein Gp17 | | Authors: | Sun, S, Rossmann, M.G. | | Deposit date: | 2006-11-27 | | Release date: | 2007-04-03 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | The Structure of the ATPase that Powers DNA Packaging into Bacteriophage T4 Procapsids
MOL.CELL, 25, 2007
|
|
2ATQ
 
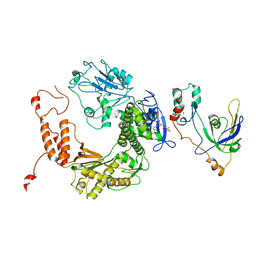 | | RB69 single-stranded DNA binding protein-DNA polymerase fusion | | Descriptor: | DNA polymerase, GUANOSINE-5'-DIPHOSPHATE, ZINC ION, ... | | Authors: | Sun, S, Geng, L, Shamoo, Y. | | Deposit date: | 2005-08-25 | | Release date: | 2006-05-09 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structure and enzymatic properties of a chimeric bacteriophage RB69 DNA polymerase and single-stranded DNA binding protein with increased processivity.
Proteins, 65, 2006
|
|
2O0J
 
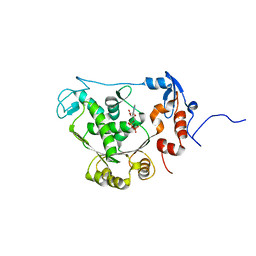 | | T4 gp17 ATPase domain mutant complexed with ADP | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, DNA packaging protein Gp17 | | Authors: | Sun, S, Rossmann, M.G. | | Deposit date: | 2006-11-27 | | Release date: | 2007-04-03 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Structure of the ATPase that Powers DNA Packaging into Bacteriophage T4 Procapsids
MOL.CELL, 25, 2007
|
|
2RAO
 
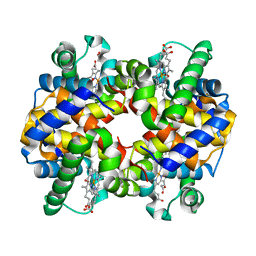 | | X ray crystal structure of rabbit hemoglobin (oxy form) at 2.0 angstrom resolution | | Descriptor: | Hemoglobin subunit alpha-1/2, Hemoglobin subunit beta-1/2, OXYGEN MOLECULE, ... | | Authors: | Sundaresan, S, Packianathan, C, Neelagandan, K, Ponnuswamy, M.N. | | Deposit date: | 2007-09-17 | | Release date: | 2008-09-09 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | X-ray crystal structure determination of hemoglobin from rabbit at 2.0 angstrom resloution
To be Published
|
|
7JM4
 
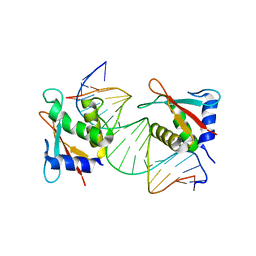 | | IRF Transcription Factor | | Descriptor: | Interferon regulatory factor 4, Interferon-Stimulated Response Elements | | Authors: | Sundararaj, S, Williams, S.J, Casarotto, M.G. | | Deposit date: | 2020-07-31 | | Release date: | 2021-03-24 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structural determinants of the IRF4/DNA homodimeric complex.
Nucleic Acids Res., 49, 2021
|
|
8BRF
 
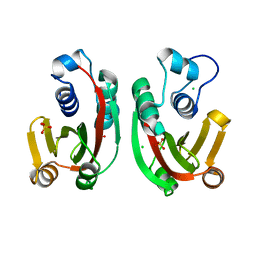 | |
4TUG
 
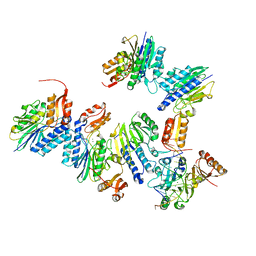 | | Crystal structure of MjMre11-DNA2 complex | | Descriptor: | DNA (5'-D(P*CP*TP*GP*TP*CP*CP*TP*AP*CP*GP*TP*GP*CP*CP*A)-3'), DNA (5'-D(P*GP*CP*AP*CP*GP*TP*AP*GP*GP*AP*CP*AP*GP*C)-3'), DNA double-strand break repair protein Mre11, ... | | Authors: | Sung, S, Cho, Y. | | Deposit date: | 2014-06-24 | | Release date: | 2014-10-15 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.55 Å) | | Cite: | DNA end recognition by the Mre11 nuclease dimer: insights into resection and repair of damaged DNA.
Embo J., 33, 2014
|
|
4TUI
 
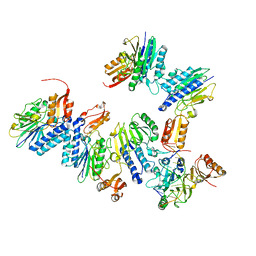 | | Crystal structure of MjMre11-DNA1 complex | | Descriptor: | DNA (5'-D(P*TP*CP*CP*TP*AP*CP*GP*TP*GP*CP*CP*AP*G)-3'), DNA (5'-D(P*TP*GP*GP*CP*AP*CP*GP*TP*AP*GP*GP*AP*C)-3'), DNA double-strand break repair protein Mre11 | | Authors: | Sung, S, Cho, Y. | | Deposit date: | 2014-06-24 | | Release date: | 2014-10-15 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.59 Å) | | Cite: | DNA end recognition by the Mre11 nuclease dimer: insights into resection and repair of damaged DNA.
Embo J., 33, 2014
|
|
