3IS6
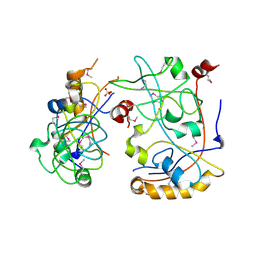 
 | | The Crystal Structure of a domain of a putative Permease protein from Porphyromonas gingivalis to 2A | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, SULFATE ION, putative permease protein, ... | | Authors: | Stein, A.J, Sather, A, Duggan, E, Moy, S, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2009-08-25 | | Release date: | 2009-09-08 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The Crystal Structure of a domain of a putative Permease protein from Porphyromonas gingivalis to 2A
To be Published
|
|
3IC8
 
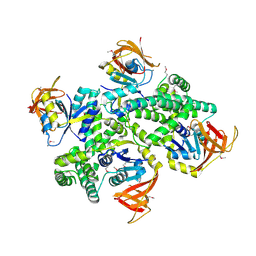 | |
3IG2
 
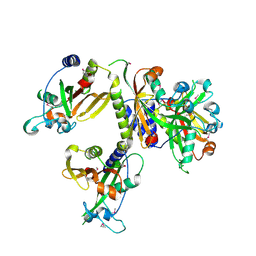 | | The Crystal Structure of a Putative Phenylalanyl-tRNA synthetase (PheRS) beta chain domain from Bacteroides fragilis to 2.1A | | Descriptor: | MAGNESIUM ION, Phenylalanyl-tRNA synthetase beta chain | | Authors: | Stein, A.J, Sather, A, Hendricks, R, Keigher, L, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2009-07-27 | | Release date: | 2009-09-01 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | The Crystal Structure of a Putative Phenylalanyl-tRNA synthetase (PheRS) beta chain domain from Bacteroides fragilis to 2.1A
To be Published
|
|
3IGR
 
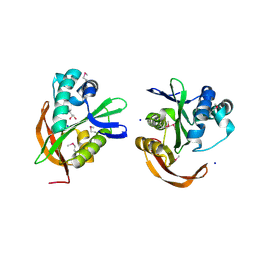 | | The Crystal Structure of Ribosomal-protein-S5-alanine Acetyltransferase from Vibrio fischeri to 2.0A | | Descriptor: | GLYCEROL, Ribosomal-protein-S5-alanine N-acetyltransferase, SODIUM ION | | Authors: | Stein, A.J, Sather, A, Shackelford, G, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2009-07-28 | | Release date: | 2009-09-01 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Crystal Structure of Ribosomal-protein-S5-alanine Acetyltransferase from Vibrio fischeri to 2.0A
To be Published
|
|
3IVL
 
 | |
3IWF
 
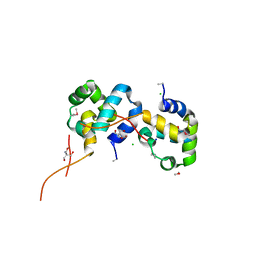 | | The Crystal Structure of the N-terminal domain of a RpiR Transcriptional Regulator from Staphylococcus epidermidis to 1.4A | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 2-METHOXYETHANOL, CHLORIDE ION, ... | | Authors: | Stein, A.J, Sather, A, Borovilos, M, Bargassa, M, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2009-09-02 | | Release date: | 2009-09-15 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | The Crystal Structure of the N-terminal domain of a RpiR Transcriptional Regulator from Staphylococcus epidermidis to 1.4A
To be Published
|
|
4J3I
 
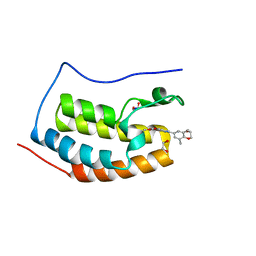 | | X-ray crystal structure of bromodomain complex to 1.24 A resolution | | Descriptor: | 1,2-ETHANEDIOL, 2-[4-(2-hydroxyethoxy)-3,5-dimethylphenyl]-5,7-dimethoxyquinazolin-4(3H)-one, Bromodomain-containing protein 4, ... | | Authors: | Stein, A.J, White, A, Suto, R.K. | | Deposit date: | 2013-02-05 | | Release date: | 2014-01-15 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.24 Å) | | Cite: | RVX-208, an Inducer of ApoA-I in Humans, Is a BET Bromodomain Antagonist.
Plos One, 8, 2013
|
|
4J1P
 
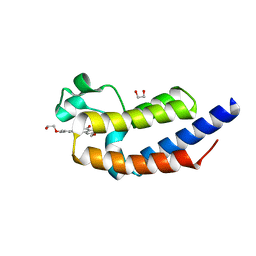 | | X-ray crystal structure of bromodomain 2 of human brd2 in complex with rvx208 to 1.08 A resolution | | Descriptor: | 1,2-ETHANEDIOL, 2-[4-(2-hydroxyethoxy)-3,5-dimethylphenyl]-5,7-dimethoxyquinazolin-4(3H)-one, Bromodomain containing 2, ... | | Authors: | Stein, A.J, White, A, Suto, R.K. | | Deposit date: | 2013-02-01 | | Release date: | 2014-01-15 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.08 Å) | | Cite: | RVX-208, an Inducer of ApoA-I in Humans, Is a BET Bromodomain Antagonist.
Plos One, 8, 2013
|
|
3LHF
 
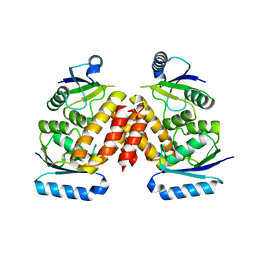 | | The Crystal Structure of a Serine Recombinase from Sulfolobus solfataricus to 2.3A | | Descriptor: | Serine Recombinase | | Authors: | Stein, A.J, Osipiuk, J, Marshall, N, Bearden, J, Davidoff, J, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-01-22 | | Release date: | 2010-03-16 | | Last modified: | 2021-10-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The Crystal Structure of a Serine Recombinase from Sulfolobus solfataricus to 2.3A
To be Published
|
|
3LK7
 
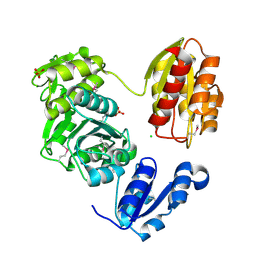 | | The Crystal Structure of UDP-N-acetylmuramoylalanine-D-glutamate (MurD) ligase from Streptococcus agalactiae to 1.5A | | Descriptor: | CHLORIDE ION, SULFATE ION, UDP-N-acetylmuramoylalanine--D-glutamate ligase | | Authors: | Stein, A.J, Sather, A, Shakelford, G, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-01-27 | | Release date: | 2010-02-09 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The Crystal Structure of UDP-N-acetylmuramoylalanine-D-glutamate (MurD) ligase from Streptococcus agalactiae to 1.5A
To be Published
|
|
3LEC
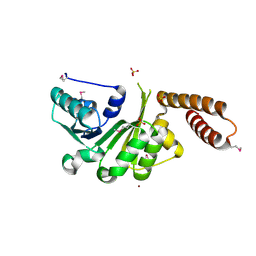 
 | | The Crystal Structure of a protein in the NADB-Rossmann Superfamily from Streptococcus agalactiae to 1.8A | | Descriptor: | NADB-Rossmann Superfamily protein, SULFATE ION, ZINC ION | | Authors: | Stein, A.J, Hatzos, C, Buck, K, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-01-14 | | Release date: | 2010-01-26 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Crystal Structure of a protein in the NADB-Rossmann Superfamily from Streptococcus agalactiae to 1.8A
To be Published
|
|
3LEQ
 
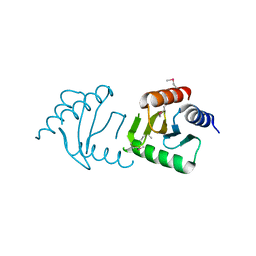 | | The Crystal Structure of the Roadblock/LC7 domain from Streptomyces avermitillis to 1.85A | | Descriptor: | uncharacterized protein cvnB5 | | Authors: | Stein, A.J, Xu, X, Cui, H, Ng, J, Edwards, A, Savchenko, A, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-01-15 | | Release date: | 2010-02-02 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | The Crystal Structure of the Roadblock/LC7 domain from Streptomyces avermitillis to 1.85A
To be Published
|
|
3LFR
 
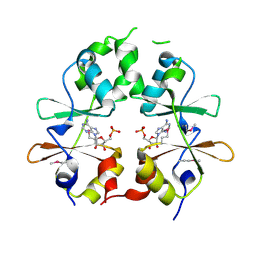 | |
3LOR
 
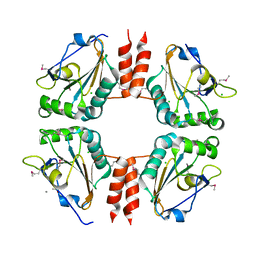 | | The Crystal Structure of a Thiol-disulfide Isomerase from Corynebacterium glutamicum to 2.2A | | Descriptor: | ACETATE ION, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Stein, A.J, Osipiuk, J, Weger, A, Cobb, G, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-02-04 | | Release date: | 2010-03-16 | | Last modified: | 2017-11-01 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The Crystal Structure of a Thiol-disulfide Isomerase from Corynebacterium glutamicum to 2.2A
To be Published
|
|
3LUQ
 
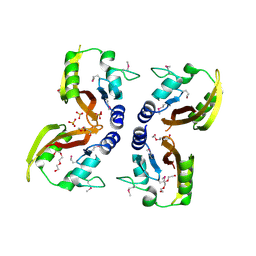 | | The Crystal Structure of a PAS Domain from a Sensory Box Histidine Kinase Regulator from Geobacter sulfurreducens to 2.5A | | Descriptor: | SULFATE ION, Sensor protein, TRIETHYLENE GLYCOL | | Authors: | Stein, A.J, Weger, A, Duggan, E, Clancy, S, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-02-18 | | Release date: | 2010-03-16 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | The Crystal Structure of a PAS Domain from a Sensory Box Histidine Kinase Regulator from Geobacter sulfurreducens to 2.5A
To be Published
|
|
3LXQ
 
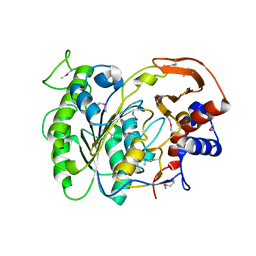 | | The Crystal Structure of a Protein in the Alkaline Phosphatase Superfamily from Vibrio parahaemolyticus to 1.95A | | Descriptor: | CHLORIDE ION, Uncharacterized protein VP1736 | | Authors: | Stein, A.J, Weger, A, Duggan, E, Clancy, S, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-02-25 | | Release date: | 2010-03-09 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The Crystal Structure of a Protein in the Alkaline Phosphatase Superfamily from Vibrio parahaemolyticus to 1.95A
To be Published
|
|
3LW7
 
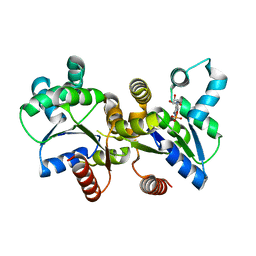 | | The Crystal Structure of an Adenylate kinase-related protein bound to AMP from sulfolobus solfataricus to 2.3A | | Descriptor: | ADENOSINE MONOPHOSPHATE, Adenylate kinase related protein (AdkA-like) | | Authors: | Stein, A.J, Sather, A, Hendricks, R, Abdullah, J, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-02-23 | | Release date: | 2010-03-09 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The Crystal Structure of an Adenylate kinase-related protein bound to AMP from sulfolobus solfataricus to 2.3A
To be Published
|
|
3LVT
 
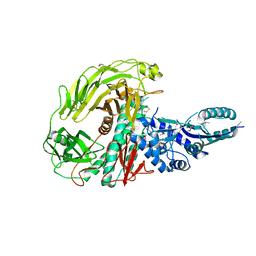 | | The Crystal Structure of a Protein in the Glycosyl Hydrolase Family 38 from Enterococcus faecalis to 2.55A | | Descriptor: | CALCIUM ION, Glycosyl hydrolase, family 38 | | Authors: | Stein, A.J, Binkowski, T.A, Weger, A, Borovilos, M, Moy, S, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-02-22 | | Release date: | 2010-03-16 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | The Crystal Structure of a Protein in the Glycosyl Hydrolase Family 38 from Enterococcus faecalis to 2.55A
To be Published
|
|
3LWA
 
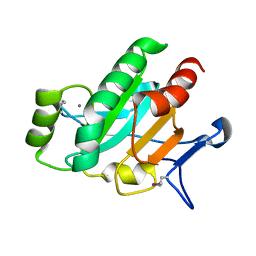 | | The Crystal Structure of a Secreted Thiol-disulfide Isomerase from Corynebacterium glutamicum to 1.75A | | Descriptor: | CALCIUM ION, Secreted thiol-disulfide isomerase | | Authors: | Stein, A.J, Weger, A, Hendricks, R, Cobb, G, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-02-23 | | Release date: | 2010-03-02 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The Crystal Structure of a Secreted Thiol-disulfide Isomerase from Corynebacterium glutamicum to 1.75A
To be Published
|
|
3M1A
 
 | | The Crystal Structure of a Short-chain Dehydrogenase from Streptomyces avermitilis to 2A | | Descriptor: | ACETATE ION, Putative dehydrogenase, SODIUM ION | | Authors: | Stein, A.J, Evdokimova, E, Egorova, O, Savchenko, A, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-03-04 | | Release date: | 2010-03-23 | | Last modified: | 2021-10-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The Crystal Structure of a Short-chain Dehydrogenase from Streptomyces avermitilis to 2A
To be Published
|
|
3O5Y
 
 | |
3O60
 
 | |
3O5V
 
 | | The Crystal Structure of the Creatinase/Prolidase N-terminal domain of an X-PRO dipeptidase from Streptococcus pyogenes to 1.85A | | Descriptor: | CHLORIDE ION, GLYCEROL, X-PRO dipeptidase | | Authors: | Stein, A.J, Wu, R, Clancy, S, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-07-28 | | Release date: | 2010-08-11 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | The Crystal Structure of the Creatinase/Prolidase N-terminal domain of an X-PRO dipeptidase from Streptococcus pyogenes to 1.85A
To be Published
|
|
3MY7
 
 | | The Crystal Structure of the ACDH domain of an Alcohol Dehydrogenase from Vibrio parahaemolyticus to 2.25A | | Descriptor: | Alcohol dehydrogenase/acetaldehyde dehydrogenase, CHLORIDE ION | | Authors: | Stein, A.J, Weger, A, Volkart, L, Gu, M, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-05-10 | | Release date: | 2010-06-02 | | Last modified: | 2017-11-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The Crystal Structure of the ACDH domain of an Alcohol Dehydrogenase from Vibrio parahaemolyticus to 2.25A
To be Published
|
|
3MZY
 
 | |
