4PF4
 
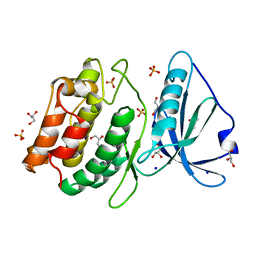 | | 1.1A X-RAY STRUCTURE OF THE APO CATALYTIC DOMAIN OF DEATH-ASSOCIATED PROTEIN KINASE 1, aa 1-277 | | Descriptor: | Death-associated protein kinase 1, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Temmerman, K, Simon, B, Wilmanns, M. | | Deposit date: | 2014-04-28 | | Release date: | 2014-05-14 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.13 Å) | | Cite: | 1.1A X-RAY STRUCTURE OF THE APO CATALYTIC DOMAIN OF DEATH-ASSOCIATED PROTEIN KINASE 1, aa 1-277
To Be Published
|
|
2XNF
 
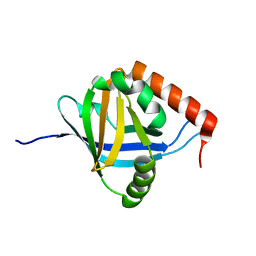 | | The Mediator Med25 activator interaction domain: Structure and cooperative binding of VP16 subdomains | | Descriptor: | MEDIATOR OF RNA POLYMERASE II TRANSCRIPTION SUBUNIT 25 | | Authors: | Vojnic, E, Mourao, A, Seizl, M, Simon, B, Wenzeck, L, Lariviere, L, Baumli, S, Meisterernst, M, Sattler, M, Cramer, P. | | Deposit date: | 2010-08-02 | | Release date: | 2011-03-09 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structure and Vp16 Binding of the Mediator Med25 Activator Interaction Domain.
Nat.Struct.Mol.Biol., 18, 2011
|
|
1R2N
 
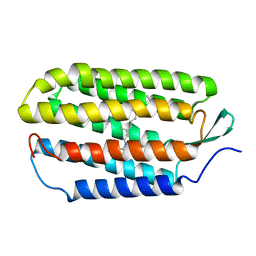 | | NMR structure of the all-trans retinal in dark-adapted Bacteriorhodopsin | | Descriptor: | Bacteriorhodopsin, RETINAL | | Authors: | Patzelt, H, Simon, B, terLaak, A, Kessler, B, Kuhne, R, Schmieder, P, Oesterhaelt, D, Oschkinat, H. | | Deposit date: | 2003-09-29 | | Release date: | 2003-10-28 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | The structures of the active center in dark-adapted bacteriorhodopsin by solution-state NMR spectroscopy
Proc.Natl.Acad.Sci.USA, 99, 2002
|
|
1R84
 
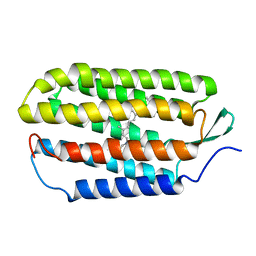 | | NMR structure of the 13-cis-15-syn retinal in dark_adapted bacteriorhodopsin | | Descriptor: | Bacteriorhodopsin, RETINAL | | Authors: | Patzelt, H, Simon, B, Ter Laak, A, Kessler, B, Kuhne, R, Schmieder, P, Oesterhaelt, D, Oschkinat, H. | | Deposit date: | 2003-10-23 | | Release date: | 2003-11-11 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | The structures of the active center in dark-adapted bacteriorhodopsin by solution-state NMR spectroscopy
Proc.Natl.Acad.Sci.USA, 99, 2002
|
|
2C5Z
 
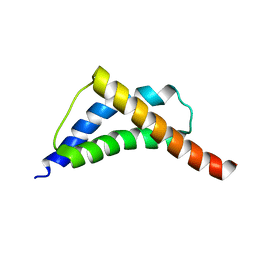 | | Structure and CTD binding of the Set2 SRI domain | | Descriptor: | SET DOMAIN PROTEIN 2 | | Authors: | Vojnic, E, Simon, B, Strahl, B.D, Sattler, M, Cramer, P. | | Deposit date: | 2005-11-03 | | Release date: | 2005-11-14 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structure and carboxyl-terminal domain (CTD) binding of the Set2 SRI domain that couples histone H3 Lys36 methylation to transcription.
J. Biol. Chem., 281, 2006
|
|
2YH1
 
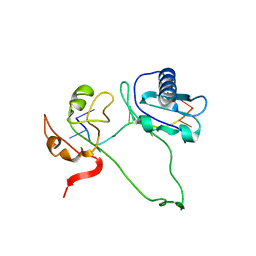 | | Model of human U2AF65 tandem RRM1 and RRM2 domains with eight-site uridine binding | | Descriptor: | 5'-R(*UP*UP*UP*UP*UP*UP*UP*UP*UP)-3', SPLICING FACTOR U2AF 65 KDA SUBUNIT | | Authors: | Mackereth, C.D, Madl, T, Simon, B, Zanier, K, Gasch, A, Sattler, M. | | Deposit date: | 2011-04-26 | | Release date: | 2011-07-20 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Multi-Domain Conformational Selection Underlies Pre-Mrna Splicing Regulation by U2Af
Nature, 475, 2011
|
|
6Y96
 
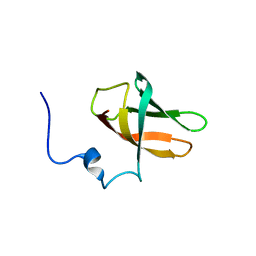 | | solution structure of cold-shock domain 9 of drosophila Upstream of N-Ras (Unr) | | Descriptor: | Upstream of N-ras, isoform A | | Authors: | Sweetapple, L.J, Hollmann, N.M, Simon, B, Hennig, J. | | Deposit date: | 2020-03-06 | | Release date: | 2020-07-29 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Pseudo-RNA-Binding Domains Mediate RNA Structure Specificity in Upstream of N-Ras.
Cell Rep, 32, 2020
|
|
6Y4H
 
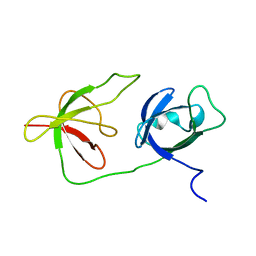 | |
2B9Z
 
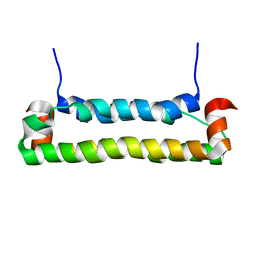 | | Solution structure of FHV B2, a viral suppressor of RNAi | | Descriptor: | B2 protein | | Authors: | Lingel, A, Simon, B, Izaurralde, E, Sattler, M. | | Deposit date: | 2005-10-13 | | Release date: | 2005-11-29 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The structure of the flock house virus B2 protein, a viral suppressor of RNA interference, shows a novel mode of double-stranded RNA recognition.
Embo Rep., 6, 2005
|
|
4BD3
 
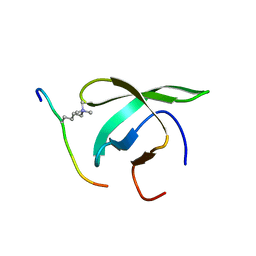 | |
2YH0
 
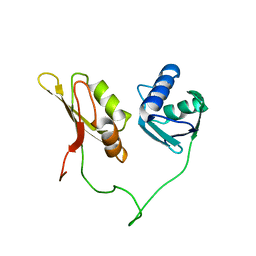 | | Solution structure of the closed conformation of human U2AF65 tandem RRM1 and RRM2 domains | | Descriptor: | SPLICING FACTOR U2AF 65 KDA SUBUNIT | | Authors: | Mackereth, C.D, Madl, T, Simon, B, Zanier, K, Gasch, A, Sattler, M. | | Deposit date: | 2011-04-26 | | Release date: | 2011-07-20 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Multi-Domain Conformational Selection Underlies Pre-Mrna Splicing Regulation by U2Af
Nature, 475, 2011
|
|
1W4M
 
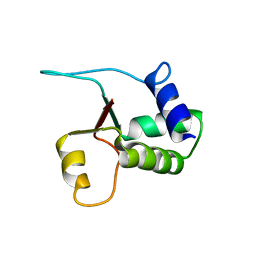 | | Structure of the human pleckstrin DEP domain by multidimensional NMR | | Descriptor: | PLECKSTRIN | | Authors: | Civera, C, Simon, B, Stier, G, Sattler, M, Macias, M.J. | | Deposit date: | 2004-07-26 | | Release date: | 2004-12-15 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structure and Dynamics of the Human Pleckstrin Dep Domain: Distinct Molecular Features of a Novel Dep Domain Subfamily
Proteins: Struct.,Funct., Genet., 58, 2005
|
|
1VYN
 
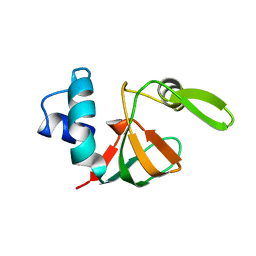 | |
2M58
 
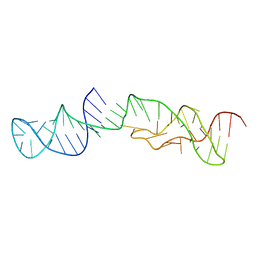 | | Structure of 2'-5' AG1 lariat forming ribozyme in its inactive state | | Descriptor: | RNA (59-MER) | | Authors: | Carlomagno, T, Amata, I, Codutti, L, Falb, M, Fohrer, J, Simon, B. | | Deposit date: | 2013-02-18 | | Release date: | 2013-04-03 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Structural principles of RNA catalysis in a 2'-5' lariat-forming ribozyme.
J.Am.Chem.Soc., 135, 2013
|
|
2N0R
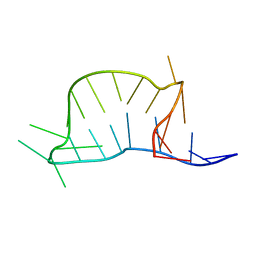 
 | |
