5HM6
 
 | |
7MTO
 | |
4PPX
 
 | | DNA Polymerase Beta E295K with Spiroiminodihydantoin in Templating Position | | Descriptor: | 5'-D(*CP*CP*GP*AP*CP*(SDH)P*GP*CP*GP*CP*AP*TP*CP*AP*GP*C)-3', 5'-D(*GP*CP*TP*GP*AP*TP*GP*CP*GP*C)-3', 5'-D(*GP*TP*CP*GP*G)-3', ... | | Authors: | Eckenroth, B.E, Doublie, S. | | Deposit date: | 2014-02-27 | | Release date: | 2014-04-02 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.08 Å) | | Cite: | Crystal Structure of DNA Polymerase beta with DNA Containing the Base Lesion Spiroiminodihydantoin in a Templating Position.
Biochemistry, 53, 2014
|
|
8GM7
 
 | |
8GM6
 
 | |
6VJI
 
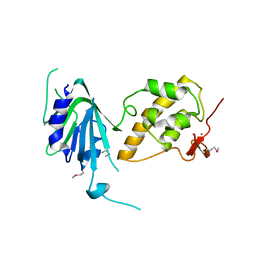 | |
8SYJ
 
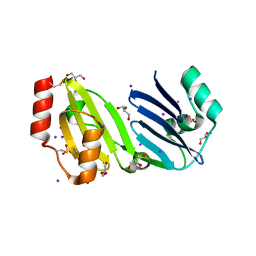 | | Structure of apurinic/apyrimidinic DNA lyase TK0353 from Thermococcus kodakarensis (Iodide crystal form) | | Descriptor: | GLYCEROL, IODIDE ION, SULFATE ION, ... | | Authors: | Eckenroth, B.E, Gardner, A.F, Doublie, S. | | Deposit date: | 2023-05-25 | | Release date: | 2023-12-06 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Thermococcus kodakarensis TK0353 is a novel AP lyase with a new fold.
J.Biol.Chem., 300, 2023
|
|
7RAG
 
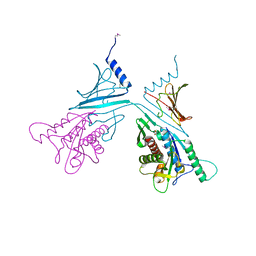 | |
6OLY
 
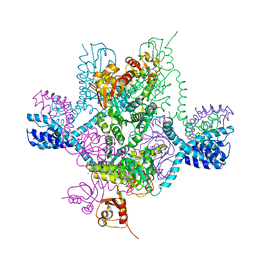 | | Full-length MthK channel at 3.1 angstrom resolution | | Descriptor: | CALCIUM ION, Calcium-gated potassium channel MthK | | Authors: | Rothberg, B.S. | | Deposit date: | 2019-04-17 | | Release date: | 2019-12-11 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (3.112 Å) | | Cite: | Molecular mechanism of a potassium channel gating through activation gate-selectivity filter coupling.
Nat Commun, 10, 2019
|
|
8TH9
 
 | |
5U8G
 
 | | DNA Polymerase Beta crystallized in PEG 400 | | Descriptor: | CHLORIDE ION, DNA (5'-D(*CP*CP*GP*AP*CP*AP*GP*CP*GP*CP*AP*TP*CP*AP*GP*C)-3'), DNA (5'-D(*GP*CP*TP*GP*AP*TP*GP*CP*GP*C)-3'), ... | | Authors: | Eckenroth, B.E, Doublie, S. | | Deposit date: | 2016-12-14 | | Release date: | 2017-04-26 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.166 Å) | | Cite: | Remote Mutations Induce Functional Changes in Active Site Residues of Human DNA Polymerase beta.
Biochemistry, 56, 2017
|
|
5U8I
 
 | | DNA Polymerase Beta S229L crystallized in PEG 400 | | Descriptor: | DNA (5'-D(*CP*CP*GP*AP*CP*AP*GP*CP*GP*CP*AP*TP*CP*AP*GP*C)-3'), DNA (5'-D(*GP*CP*TP*GP*AP*TP*GP*CP*GP*C)-3'), DNA (5'-D(*GP*TP*CP*GP*G)-3'), ... | | Authors: | Eckenroth, B.E, Doublie, S. | | Deposit date: | 2016-12-14 | | Release date: | 2017-04-26 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Remote Mutations Induce Functional Changes in Active Site Residues of Human DNA Polymerase beta.
Biochemistry, 56, 2017
|
|
5U8H
 
 | | DNA Polymerase Beta G231D crystallized in PEG 400 | | Descriptor: | DNA (5'-D(*CP*CP*GP*AP*CP*AP*GP*CP*GP*CP*AP*TP*CP*AP*GP*C)-3'), DNA (5'-D(*GP*CP*TP*GP*AP*TP*GP*CP*GP*C)-3'), DNA (5'-D(*GP*TP*CP*GP*G)-3'), ... | | Authors: | Eckenroth, B.E, Doublie, S. | | Deposit date: | 2016-12-14 | | Release date: | 2017-04-26 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.155 Å) | | Cite: | Remote Mutations Induce Functional Changes in Active Site Residues of Human DNA Polymerase beta.
Biochemistry, 56, 2017
|
|
6BTF
 
 | | DNA Polymerase Beta I260Q Ternary Complex | | Descriptor: | 2'-DEOXYURIDINE 5'-ALPHA,BETA-IMIDO-TRIPHOSPHATE, DNA Downstream Strand, DNA Primer Strand, ... | | Authors: | Eckenroth, B.E, Doublie, S. | | Deposit date: | 2017-12-06 | | Release date: | 2018-09-26 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | I260Q DNA polymerase beta highlights precatalytic conformational rearrangements critical for fidelity.
Nucleic Acids Res., 46, 2018
|
|
6BTE
 
 | | DNA Polymerase Beta I260Q Binary Complex | | Descriptor: | CHLORIDE ION, DNA Downstream Strand, DNA Primer Strand, ... | | Authors: | Eckenroth, B.E, Doublie, S. | | Deposit date: | 2017-12-06 | | Release date: | 2018-09-26 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | I260Q DNA polymerase beta highlights precatalytic conformational rearrangements critical for fidelity.
Nucleic Acids Res., 46, 2018
|
|
6MW4
 
 | |
3DGH
 
 | | Crystal Structure of Drosophila Thioredoxin Reductase, C-terminal 8-residue truncation | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, SULFATE ION, Thioredoxin reductase 1, ... | | Authors: | Eckenroth, B.E, Hondal, R.J, Everse, S.J. | | Deposit date: | 2008-06-13 | | Release date: | 2009-06-16 | | Last modified: | 2021-10-20 | | Method: | X-RAY DIFFRACTION (1.745 Å) | | Cite: | Crystal Structure of Drosophila Thioredoxin Reductase, C-terminal 8-residue truncation
To be Published
|
|
4M9H
 
 | | DNA Polymerase Beta E295K Soaked with dTTP | | Descriptor: | CHLORIDE ION, DNA Downstream Strand, DNA Primer Strand, ... | | Authors: | Eckenroth, B.E, Doublie, S. | | Deposit date: | 2013-08-14 | | Release date: | 2013-10-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.394 Å) | | Cite: | The E295K Cancer Variant of Human Polymerase beta Favors the Mismatch Conformational Pathway during Nucleotide Selection.
J.Biol.Chem., 288, 2013
|
|
4M9N
 
 | | DNA Polymerase Beta E295K Soaked with dATP | | Descriptor: | 2'-DEOXYADENOSINE 5'-TRIPHOSPHATE, DNA Downstream Strand, DNA Primer Strand, ... | | Authors: | Eckenroth, B.E, Doublie, S. | | Deposit date: | 2013-08-14 | | Release date: | 2013-10-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.275 Å) | | Cite: | The E295K Cancer Variant of Human Polymerase beta Favors the Mismatch Conformational Pathway during Nucleotide Selection.
J.Biol.Chem., 288, 2013
|
|
3DH9
 
 | |
4M9G
 
 | | DNA Polymerase Beta E295K Binary Complex | | Descriptor: | CHLORIDE ION, DNA Downstream Strand, DNA Primer Strand, ... | | Authors: | Eckenroth, B.E, Doublie, S. | | Deposit date: | 2013-08-14 | | Release date: | 2013-10-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.01 Å) | | Cite: | The E295K Cancer Variant of Human Polymerase beta Favors the Mismatch Conformational Pathway during Nucleotide Selection.
J.Biol.Chem., 288, 2013
|
|
4M9L
 
 | | DNA Polymerase Beta E295K Soaked with dCTP | | Descriptor: | 2'-DEOXYCYTIDINE-5'-TRIPHOSPHATE, DNA Downstream Strand, DNA Primer Strand, ... | | Authors: | Eckenroth, B.E, Doublie, S. | | Deposit date: | 2013-08-14 | | Release date: | 2013-10-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.091 Å) | | Cite: | The E295K Cancer Variant of Human Polymerase beta Favors the Mismatch Conformational Pathway during Nucleotide Selection.
J.Biol.Chem., 288, 2013
|
|
4M9J
 
 | | DNA Polymerase Beta E295K Soaked with dUMPNPP | | Descriptor: | 2'-DEOXYURIDINE 5'-ALPHA,BETA-IMIDO-TRIPHOSPHATE, CHLORIDE ION, DNA Downstream Strand, ... | | Authors: | Eckenroth, B.E, Doublie, S. | | Deposit date: | 2013-08-14 | | Release date: | 2013-10-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.038 Å) | | Cite: | The E295K Cancer Variant of Human Polymerase beta Favors the Mismatch Conformational Pathway during Nucleotide Selection.
J.Biol.Chem., 288, 2013
|
|
3DGZ
 
 | | Crystal Structure of Mouse Mitochondrial Thioredoxin Reductase, C-terminal 3-residue truncation | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, Thioredoxin reductase 2, [(2R,3R,4R,5R)-5-(6-AMINO-9H-PURIN-9-YL)-3-HYDROXY-4-(PHOSPHONOOXY)TETRAHYDROFURAN-2-YL]METHYL [(2R,3S,4S)-3,4-DIHYDROXYTETRAHYDROFURAN-2-YL]METHYL DIHYDROGEN DIPHOSPHATE | | Authors: | Eckenroth, B.E, Hondal, R.J, Everse, S.J. | | Deposit date: | 2008-06-16 | | Release date: | 2009-06-16 | | Last modified: | 2017-10-25 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Crystal Structure of Mouse Mitochondrial Thioredoxin Reductase, C-terminal 3-residue truncation
To be Published
|
|
2NVK
 
 | | Crystal Structure of Thioredoxin Reductase from Drosophila melanogaster | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Thioredoxin Reductase | | Authors: | Eckenroth, B.E, Rould, M.A, Hondal, R.J, Everse, S.J. | | Deposit date: | 2006-11-13 | | Release date: | 2007-04-24 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural and Biochemical Studies Reveal Differences in the Catalytic Mechanisms of Mammalian and Drosophila melanogaster Thioredoxin Reductases.
Biochemistry, 46, 2007
|
|
