4ZTY
 
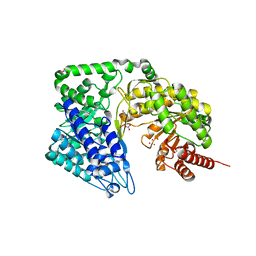 | | Neurospora crassa cobalamin-independent methionine synthase complexed with Cd2+ | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CADMIUM ION, Cobalamin-Independent Methionine synthase, ... | | Authors: | Wheatley, R.W, Ng, K.K, Kapoor, M. | | Deposit date: | 2015-05-15 | | Release date: | 2015-12-09 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Fungal cobalamin-independent methionine synthase: Insights from the model organism, Neurospora crassa.
Arch.Biochem.Biophys., 590, 2015
|
|
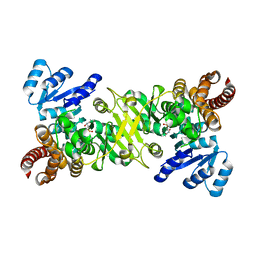
4ZQF
 
 | |
9AUQ
 
 | | Crystal structure of 4-Fluoro-tryptophan labeled Oscillatoria Agardhii agglutinin | | Descriptor: | 3-CYCLOHEXYL-1-PROPYLSULFONIC ACID, Lectin | | Authors: | Runge, B.R, Zadorozhnyi, R.R, Quinn, C.M, Russell, R.W, Lu, M, Antolinez, S, Struppe, J, Schwieters, C.D, Byeon, I.L, Hadden-Perilla, J, Gronenborn, A.M, Polenova, T. | | Deposit date: | 2024-02-29 | | Release date: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | Integrating 19 F Distance Restraints for Accurate Protein Structure Determination by Magic Angle Spinning NMR Spectroscopy.
J.Am.Chem.Soc., 2024
|
|
9B4W
 
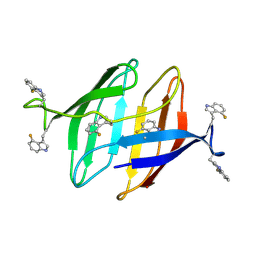 | | 4F-Trp labeled Oscillatoria Agardhii Agglutinin (OAA) | | Descriptor: | Lectin | | Authors: | Runge, B.R, Zadorozhnyi, R.R, Quinn, C.M, Russell, R.W, Lu, M, Antolinez, S, Struppe, J, Schwieters, C.D, Byeon, I.L, Hadden-Perilla, J, Gronenborn, A.M, Polenova, T. | | Deposit date: | 2024-03-21 | | Release date: | 2024-11-06 | | Method: | SOLID-STATE NMR | | Cite: | Integrating 19 F Distance Restraints for Accurate Protein Structure Determination by Magic Angle Spinning NMR Spectroscopy.
J.Am.Chem.Soc., 2024
|
|
4YNC
 
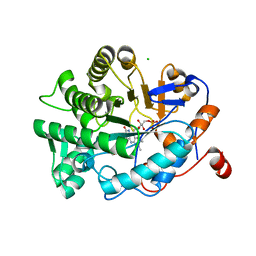 | | OYE1 W116A COMPLEXED WITH (Z)-METHYL-3-CYANO-3-PHENYLACRYLATE IN A NON PRODUCTIVE BINDING MODE | | Descriptor: | CHLORIDE ION, FLAVIN MONONUCLEOTIDE, NADPH dehydrogenase 1, ... | | Authors: | Santangelo, S, Brenna, E, Stewart, J.D, Powell III, R.W. | | Deposit date: | 2015-03-09 | | Release date: | 2016-01-20 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (1.498 Å) | | Cite: | Opposite Enantioselectivity in the Bioreduction of (Z)-beta-Aryl-beta-cyanoacrylates Mediated by the Tryptophan 116 Mutants of Old Yellow Enzyme 1: Synthetic Approach to (R)- and (S)-beta-Aryl-gamma-lactams
Adv.Synth.Catal., 357, 2015
|
|
9B4V
 
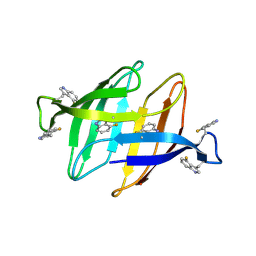 | | 4F-Trp labeled Oscillatoria Agardhii Agglutinin (OAA) | | Descriptor: | Lectin | | Authors: | Runge, B.R, Zadorozhnyi, R.R, Quinn, C.M, Russell, R.W, Lu, M, Antolinez, S, Struppe, J, Schwieters, C.D, Byeon, I.L, Hadden-Perilla, J, Gronenborn, A.M, Polenova, T. | | Deposit date: | 2024-03-21 | | Release date: | 2024-11-06 | | Method: | SOLID-STATE NMR | | Cite: | Integrating 19 F Distance Restraints for Accurate Protein Structure Determination by Magic Angle Spinning NMR Spectroscopy.
J.Am.Chem.Soc., 2024
|
|
1A5P
 
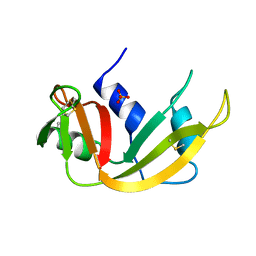 | | C[40,95]A VARIANT OF BOVINE PANCREATIC RIBONUCLEASE A | | Descriptor: | RIBONUCLEASE A, SULFATE ION | | Authors: | Pearson, M.A, Karplus, P.A, Dodge, R.W, Laity, J.H, Scheraga, H.A. | | Deposit date: | 1998-02-17 | | Release date: | 1998-05-27 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structures of two mutants that have implications for the folding of bovine pancreatic ribonuclease A.
Protein Sci., 7, 1998
|
|
1A94
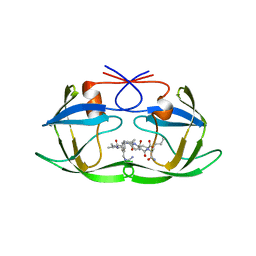 
 | | STRUCTURAL BASIS FOR SPECIFICITY OF RETROVIRAL PROTEASES | | Descriptor: | N-[(2R)-2-({N~5~-[amino(iminio)methyl]-L-ornithyl-L-valyl}amino)-4-methylpentyl]-L-phenylalanyl-L-alpha-glutamyl-L-alanyl-L-norleucinamide, PROTEASE | | Authors: | Wu, J, Adomat, J.M, Ridky, T.W, Louis, J.M, Leis, J, Harrison, R.W, Weber, I.T. | | Deposit date: | 1998-04-16 | | Release date: | 1999-01-13 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural basis for specificity of retroviral proteases.
Biochemistry, 37, 1998
|
|
4ZTX
 
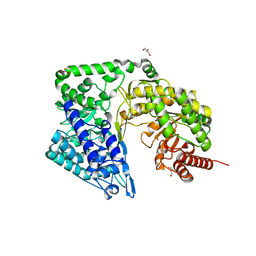 | | Neurospora crassa cobalamin-independent methionine synthase complexed with Zn2+ | | Descriptor: | Cobalamin-Independent Methionine synthase, GLYCEROL, NITRATE ION, ... | | Authors: | Wheatley, R.W, Ng, K.K, Kapoor, M. | | Deposit date: | 2015-05-15 | | Release date: | 2015-12-09 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Fungal cobalamin-independent methionine synthase: Insights from the model organism, Neurospora crassa.
Arch.Biochem.Biophys., 590, 2015
|
|
5AGF
 
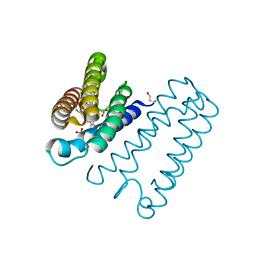 | | Nitrosyl complex of the D121Q variant of cytochrome c prime from Alcaligenes xylosoxidans | | Descriptor: | CYTOCHROME C PRIME, HEME C, NITRIC OXIDE, ... | | Authors: | Gahfoor, D.D, Kekilli, D, Abdullah, G.H, Dworkowski, F.S.N, Hassan, H.G, Wilson, M.T, Hough, M.A, Strange, R.W. | | Deposit date: | 2015-01-30 | | Release date: | 2015-09-09 | | Last modified: | 2020-03-11 | | Method: | X-RAY DIFFRACTION (1.09 Å) | | Cite: | Hydrogen Bonding of the Dissociated Histidine Ligand is not Required for Formation of a Proximal No Adduct in Cytochrome C'.
J.Biol.Inorg.Chem., 20, 2015
|
|
1A5Q
 
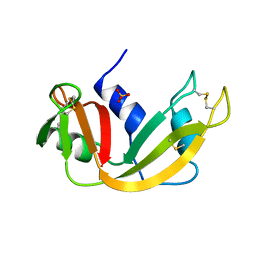 | | P93A VARIANT OF BOVINE PANCREATIC RIBONUCLEASE A | | Descriptor: | RIBONUCLEASE A, SULFATE ION | | Authors: | Pearson, M.A, Karplus, P.A, Dodge, R.W, Laity, J.H, Scheraga, H.A. | | Deposit date: | 1998-02-17 | | Release date: | 1998-05-27 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structures of two mutants that have implications for the folding of bovine pancreatic ribonuclease A.
Protein Sci., 7, 1998
|
|
5A2S
 
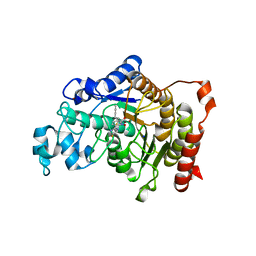 | | Potent, selective and CNS-penetrant tetrasubstituted cyclopropane class IIa histone deacetylase (HDAC) inhibitors | | Descriptor: | (1S,2S,3S)-1-fluoranyl-2-[4-(5-fluoranylpyrimidin-2-yl)phenyl]-N-oxidanyl-3-phenyl-cyclopropane-1-carboxamide, HISTONE DEACETYLASE 4, SODIUM ION, ... | | Authors: | Luckhurst, C.A, Breccia, P, Stott, A.J, Aziz, O, Birch, H, Burli, R.W, Hughes, S, Jarvis, R.E, Lamers, M, Leonard, P, Matthews, K.L, McAllister, G, Pollack, S, Saville-Stones, E, Wishart, G, Yates, D, Dominguez, C. | | Deposit date: | 2015-05-22 | | Release date: | 2016-02-10 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Potent, Selective, and Cns-Penetrant Tetrasubstituted Cyclopropane Class Iia Histone Deacetylase (Hdac) Inhibitors.
Acs Med.Chem.Lett., 7, 2016
|
|
1ANT
 
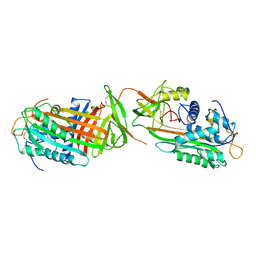 | |
1AX8
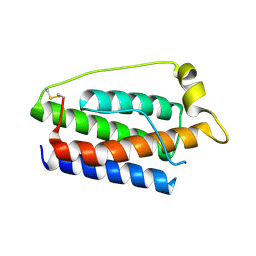 
 | | Human obesity protein, leptin | | Descriptor: | OBESITY PROTEIN | | Authors: | Zhang, F, Beals, J.M, Briggs, S.L, Clawson, D.K, Wery, J.-P, Schevitz, R.W. | | Deposit date: | 1997-10-31 | | Release date: | 1998-11-25 | | Last modified: | 2012-05-09 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of the obese protein leptin-E100.
Nature, 387, 1997
|
|
5AR2
 
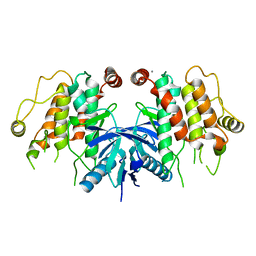 | | RIP2 Kinase Catalytic Domain (1 - 310) | | Descriptor: | CALCIUM ION, RECEPTOR-INTERACTING SERINE/THREONINE-PROTEIN KINASE 2 | | Authors: | Charnley, A.K, Convery, M.A, Lakdawala Shah, A, Jones, E, Hardwicke, P, Bridges, A, Votta, B.J, Gough, P.J, Marquis, R.W, Bertin, J, Casillas, L. | | Deposit date: | 2015-09-23 | | Release date: | 2015-10-21 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.44 Å) | | Cite: | Crystal Structures of Human Rip2 Kinase Catalytic Domain Complexed with ATP-Competitive Inhibitors: Foundations for Understanding Inhibitor Selectivity.
Bioorg.Med.Chem., 23, 2015
|
|
1BAI
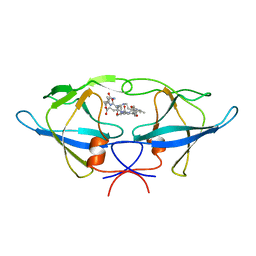 
 | | Crystal structure of Rous sarcoma virus protease in complex with inhibitor | | Descriptor: | N-[(2R)-2-({N~5~-[amino(iminio)methyl]-L-ornithyl-L-valyl}amino)-4-methylpentyl]-L-phenylalanyl-L-alpha-glutamyl-L-alanyl-L-norleucinamide, PROTEASE | | Authors: | Wu, J, Adomat, J.M, Ridky, T.W, Louis, J.M, Leis, J, Harrison, R.W, Weber, I.T. | | Deposit date: | 1998-04-17 | | Release date: | 1999-01-13 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural basis for specificity of retroviral proteases.
Biochemistry, 37, 1998
|
|
1AZX
 
 | | ANTITHROMBIN/PENTASACCHARIDE COMPLEX | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 3,4-di-O-methyl-2,6-di-O-sulfo-alpha-D-glucopyranose-(1-4)-2,3-di-O-methyl-beta-D-glucopyranuronic acid-(1-4)-2,3,6-tri-O-sulfo-alpha-D-glucopyranose-(1-4)-3-O-methyl-2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-methyl 2,3,6-tri-O-sulfo-alpha-D-glucopyranoside, ANTITHROMBIN | | Authors: | Jin, L, Abrahams, J.P, Skinner, R, Petitou, M, Pike, R.N, Carrell, R.W. | | Deposit date: | 1997-11-23 | | Release date: | 1999-01-13 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | The anticoagulant activation of antithrombin by heparin.
Proc.Natl.Acad.Sci.USA, 94, 1997
|
|
1BBC
 
 | | STRUCTURE OF RECOMBINANT HUMAN RHEUMATOID ARTHRITIC SYNOVIAL FLUID PHOSPHOLIPASE A2 AT 2.2 ANGSTROMS RESOLUTION | | Descriptor: | PHOSPHOLIPASE A2 | | Authors: | Wery, J.-P, Schevitz, R.W, Clawson, D.K, Bobbitt, J.L, Dow, E.R, Gamboa, G, Goodsonjunior, T, Hermann, R.B, Kramer, R.M, Mcclure, D.B, Mihelich, E.D, Putnam, J.E, Sharp, J.D, Stark, D.H, Teater, C, Warrick, M.W, Jones, N.D. | | Deposit date: | 1992-05-04 | | Release date: | 1993-10-31 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of recombinant human rheumatoid arthritic synovial fluid phospholipase A2 at 2.2 A resolution.
Nature, 352, 1991
|
|
1AP0
 
 | | STRUCTURE OF THE CHROMATIN BINDING (CHROMO) DOMAIN FROM MOUSE MODIFIER PROTEIN 1, NMR, 26 STRUCTURES | | Descriptor: | MODIFIER PROTEIN 1 | | Authors: | Ball, L.J, Murzina, N.V, Broadhurst, R.W, Raine, A.R.C, Archer, S.J, Stott, F.J, Murzin, A.G, Singh, P.B, Domaille, P.J, Laue, E.D. | | Deposit date: | 1997-07-22 | | Release date: | 1998-07-22 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure of the chromatin binding (chromo) domain from mouse modifier protein 1.
EMBO J., 16, 1997
|
|
5AR3
 
 | | RIP2 Kinase Catalytic Domain (1 - 310) complex with AMP-PCP | | Descriptor: | MAGNESIUM ION, PHOSPHOMETHYLPHOSPHONIC ACID ADENYLATE ESTER, RECEPTOR-INTERACTING SERINE/THREONINE-PROTEIN KINASE 2 | | Authors: | Charnley, A.K, Convery, M.A, Lakdawala Shah, A, Jones, E, Hardwicke, P, Bridges, A, Votta, B.J, Gough, P.J, Marquis, R.W, Bertin, J, Casillas, L. | | Deposit date: | 2015-09-23 | | Release date: | 2015-10-21 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3.23 Å) | | Cite: | Crystal Structures of Human Rip2 Kinase Catalytic Domain Complexed with ATP-Competitive Inhibitors: Foundations for Understanding Inhibitor Selectivity.
Bioorg.Med.Chem., 23, 2015
|
|
1CDJ
 
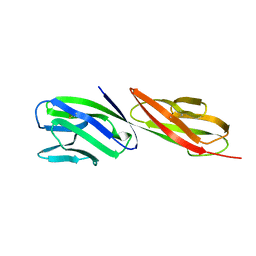 | | STRUCTURE OF T-CELL SURFACE GLYCOPROTEIN CD4 | | Descriptor: | T-CELL SURFACE GLYCOPROTEIN CD4 | | Authors: | Wu, H, Myszka, D, Tendian, S.W, Brouillette, C.G, Sweet, R.W, Chaiken, I.M, Hendrickson, W.A. | | Deposit date: | 1996-11-11 | | Release date: | 1997-04-01 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Kinetic and structural analysis of mutant CD4 receptors that are defective in HIV gp120 binding.
Proc.Natl.Acad.Sci.USA, 93, 1996
|
|
1BR8
 | | IMPLICATIONS FOR FUNCTION AND THERAPY OF A 2.9A STRUCTURE OF BINARY-COMPLEXED ANTITHROMBIN | | Descriptor: | PROTEIN (ANTITHROMBIN-III), PROTEIN (PEPTIDE) | | Authors: | Skinner, R, Chang, W.S.W, Jin, L, Pei, X.Y, Huntington, J.A, Abrahams, J.P, Carrell, R.W, Lomas, D.A. | | Deposit date: | 1998-08-26 | | Release date: | 1998-09-02 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Implications for function and therapy of a 2.9 A structure of binary-complexed antithrombin.
J.Mol.Biol., 283, 1998
|
|
1C7Z
 
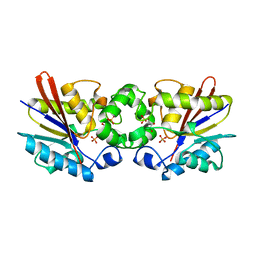 | | REGULATORY COMPLEX OF FRUCTOSE-2,6-BISPHOSPHATASE | | Descriptor: | FRUCTOSE-2,6-BISPHOSPHATASE, GLYCERALDEHYDE-3-PHOSPHATE, PHOSPHATE ION | | Authors: | Lee, Y.H, Olson, T.W, McClard, R.W, Witte, J.F, McFarlan, S.C, Banaszak, L.J, Levitt, D.G, Lange, A.J. | | Deposit date: | 2000-04-03 | | Release date: | 2003-06-10 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Reaction Mechanism of Fructose-2,6-bisphosphatase Suggested by the Crystal Structures of a pseudo-Michaelis complex and Metabolite Complexes
To be Published
|
|
1C80
 
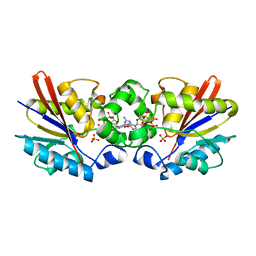 | | REGULATORY COMPLEX OF FRUCTOSE-2,6-BISPHOSPHATASE | | Descriptor: | FRUCTOSE-2,6-BISPHOSPHATASE, GUANOSINE-5'-TRIPHOSPHATE, PHOSPHATE ION | | Authors: | Lee, Y.H, Olson, T.W, McClard, R.W, Witte, J.F, McFarlan, S.C, Banaszak, L.J, Levitt, D.G, Lange, A.J. | | Deposit date: | 2000-04-03 | | Release date: | 2003-06-10 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Reaction Mechanism of Fructose-2,6-bisphosphatase Suggested by the Crystal Structures of a pseudo-Michaelis complex and Metabolite Complexes
To be Published
|
|
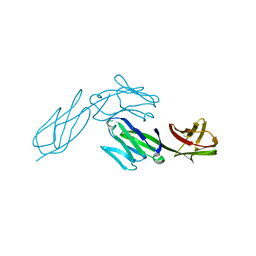
1CDH
 
 | |
